Find the nearest DNA-flagged relatives in a GEDCOM family tree
Project description
GEDCOM DNA Finder
This tool provides useful ways to explore a GEDCOM file exported from services like Ancestry, MyHeritage, Geni, and Family Tree Maker:
- Find the closest DNA-flagged person to any other person in a family tree
- Show multiple paths between any two people in your tree
- Search your tree for variations on names and filter on other information like geographical locations
- Rapidly explore names and connections in your tree
Available as a graphical tool as well as a command-line version.
Downloads:
- Windows (see security note)
- Mac (see security note)
- Linux
This is an alpha release. Only one person has tested it so far--me. If you are interested in experimenting with a "dummy" GEDCOM file rather than your own, several are available at https://github.com/findmypast/gedcom-samples
The problem this solves
Many genealogists working with autosomal DNA add unfamiliar people to their family tree based on DNA matches and then build out those people's lines, hoping to find the most recent common ancestor between the match and themselves. After accumulating thousands of these speculative additions, you often end up looking at a person in your tree and thinking: why is this person here? which DNA match did this branch come from?
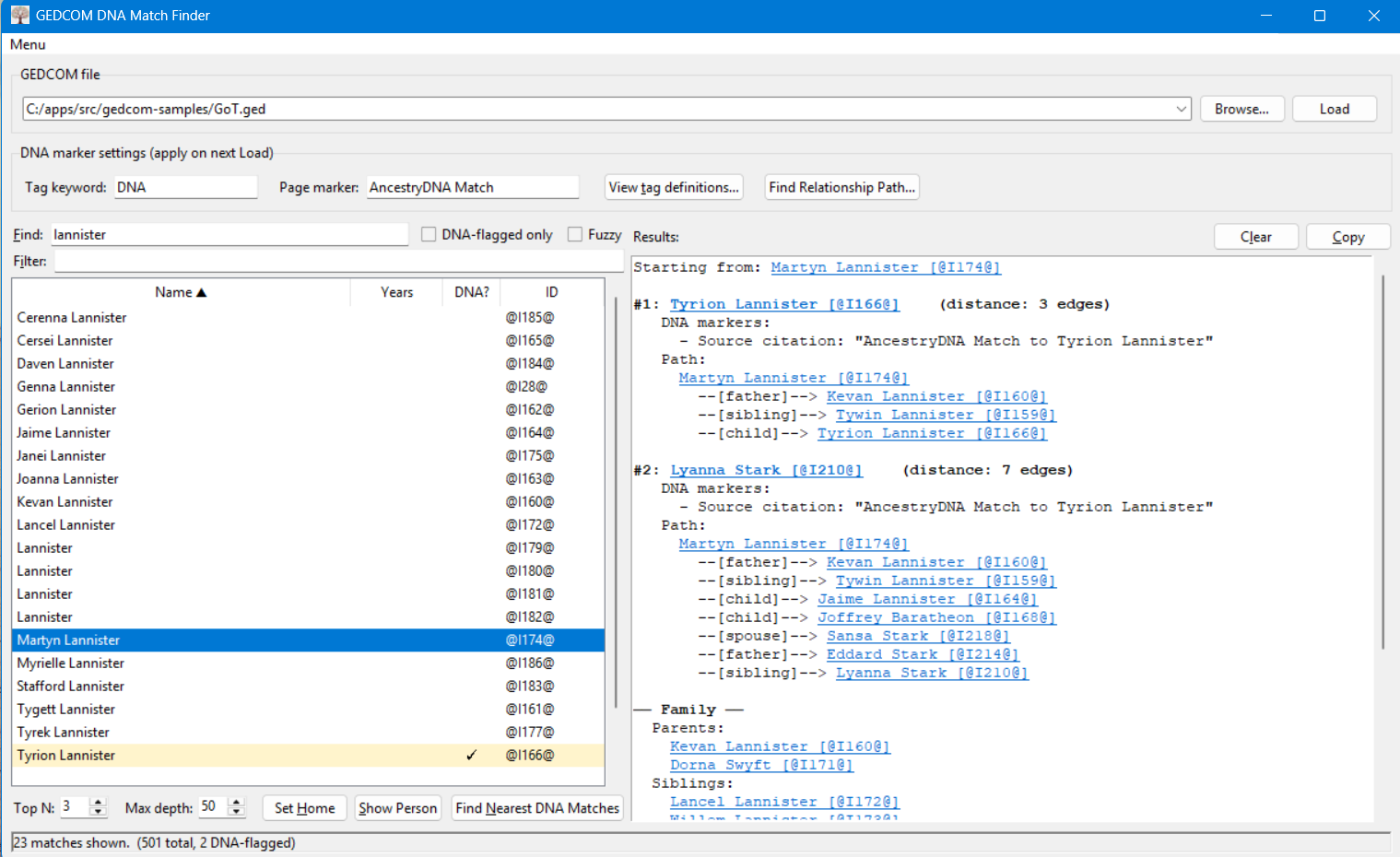
Ancestry, Family Tree Maker, and standard GEDCOM viewers can show you a flat list of everyone you've tagged as a DNA match, but none of them will, given an arbitrary person in the tree, walk outward through the relationship graph and tell you the nearest tagged relative. That is the main purpose of this tool.
As an added bonus, you can use this tool to find multiple paths between any two people in your tree and also view individual records from your tree. If you set a person as the "Home Person" using the "Set Home" button, the results will include the path from the selected person to the Home Person in addition to the closest people with DNA match markers.
Finally, if you have a large tree, you may find it difficult to search for specific individuals in tools like Ancestry or Family Tree Maker. Ancestry only searches on the person's "main name" and not any of the alternate names, and neither service allows fuzzy matching searches. Ancestry also does not allow you to easily search on multiple fields, like name and location. With this tool, you can search for a name with fuzzy matching (e.g. "John Smith" in the "Find:" box) and then further limit the results by a term that appears anywhere in the person's record (e.g. "Chicago" in the "Filter" box).
What it does
Given a GEDCOM file and a target individual, the tool performs a breadth-first search through the tree's relationship graph (parents, children, siblings, spouses) and returns the closest individuals flagged as DNA matches, along with the relationship path connecting each match to the target.
Two flag formats are recognized out of the box:
- AncestryDNA citations. When an Ancestry-managed tree marks a
person as a DNA match, the exported GEDCOM contains a source
citation with a
PAGEline of the form:2 PAGE AncestryDNA Match to Jane Q. Doe - MyTreeTags / Family Tree Maker custom tags. Tags applied via
Ancestry MyTreeTags or as a custom fact in Family Tree Maker show
up as a pointer to a tag-definition record:
with the corresponding definition elsewhere in the file:1 _MTTAG @T182059@0 @T182059@ _MTTAG 1 NAME DNA Match
Both substrings are configurable, so you can adapt the tool to other genealogy software's conventions.
Although this software was developed for this DNA use case, you could use it to find the closest path to any tag or page marker by entering that string into "tag keyword" or "page marker" rather than a DNA-specific term. For example, if your paternal relatives are tagged with a "paternal" tag, you could use this tool to find the path between anyone in your tree and anyone tagged as a paternal relative.
Requirements
The pre-built executables have no requirements.
If you want to run these scripts from the source code, you will need:
- Python 3.8 or newer
- Tkinter (only for the GUI). It ships with the official Python
installers on Windows and macOS. On most Linux distributions it is
in a separate package, typically
python3-tk.
No third-party libraries; the entire tool uses only the Python standard library.
Installation
Pre-built executables
Download the latest release for your operating system. No Python installation required.
pip (PyPI)
This application is also available on PyPI via pip. Use this command to install:
pip install gedcom-dna-finder
After installation two commands are available:
| Command | Description |
|---|---|
gedcom-dna-finder |
Command-line interface |
gedcom-dna-finder-gui |
Graphical interface |
The GUI requires Tkinter, which ships with the official Python installers on Windows and macOS. On most Linux distributions it is in a separate package:
# Debian / Ubuntu
sudo apt install python3-tk
# Fedora
sudo dnf install python3-tkinter
# Arch
sudo pacman -S tk
Run from source
git clone https://github.com/ajkessel/gedcom-dna-finder.git
cd gedcom-dna-finder
python src/gedcom-dna-finder-gui.py # GUI
python src/gedcom-dna-finder-cli.py --help # CLI
No third-party libraries are needed to run from source.
Build executables yourself
Use build.sh to compile for Linux or Mac and build.ps1 for Windows. The build script automatically creates a Python virtual environment and installs the required dependencies. These dependencies are only needed for building, not for running from source.
A build_and_release.sh script is also available that builds for all three platforms under WSL.
Usage
For pre-built binaries, just run the executable.
Relationship finder
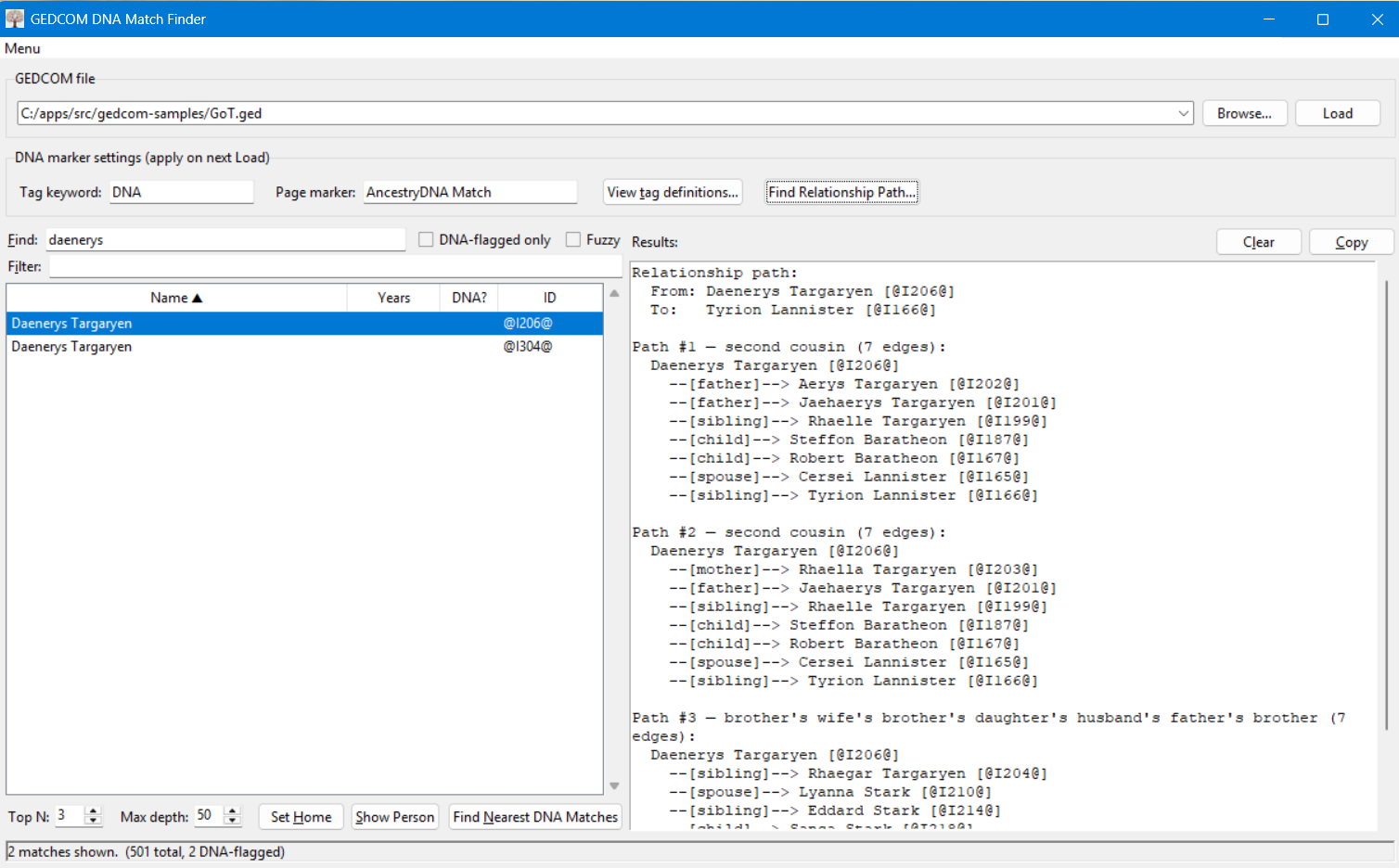
This is an alternate use of this tool. Select a person in the search panel, then click on "Find Relationship Path..." and select a second person. This tool will then show you the top three paths (if they exist) between those two people. You can change the number of paths to an arbitrary number by editing the "Top N" value in the bottom right. If the two people are very distantly related, you may need to increase the "Max Depth" setting to find the connection. The default max depth of 50 should find connections at least up to 4th cousins.
GUI
python gedcom-dna-finder-gui.py # opens with no file loaded
python gedcom-dna-finder-gui.py /path/to/tree.ged # auto-loads on startup
- Click Browse and select your
.gedfile (or pass it on the command line as shown above). The tool will also find a.gedfile inside a.zipfile, since a tree downloaded from Ancestry will be zipped. - Optionally adjust the tag keyword (default
DNA) or page marker (defaultAncestryDNA Match). The defaults work for files exported from Ancestry and Family Tree Maker. - Click Load. The status bar will show how many individuals, families, and DNA-flagged people were found.
- Type a name or INDI ID into the search box to filter the people
list. Names are matched by whitespace-separated tokens, in any
order, each as a case-insensitive substring — so
John Smithwill findJohn Adam Smith. The "DNA-flagged only" checkbox hides everyone else. - Select a person and click Find Nearest DNA Matches (or just double-click the row).
- The right pane shows the closest flagged relative(s) and the relationship path from the selected person to each one.
The View tag definitions... button opens a window listing every
_MTTAG record in the file with its name, which is useful for
deciding what tag-keyword filter to use.
Command line
# List all _MTTAG definitions in the file (use "_" as a placeholder for the target)
python gedcom-dna-finder-cli.py tree.ged --list-tags _
# List every flagged individual
python gedcom-dna-finder-cli.py tree.ged --list-flagged _
# Find the three nearest DNA-flagged relatives by name
python gedcom-dna-finder-cli.py tree.ged "Jane Doe"
# Names are matched by whitespace-separated tokens, in any order, each as
# a case-insensitive substring. The middle name is not required:
# this matches "John Adam Smith".
python gedcom-dna-finder-cli.py tree.ged "John Smith"
# Fuzzy matching tolerates typos and spelling variants. The default
# similarity threshold is 0.6; raise it for stricter matches.
python gedcom-dna-finder-cli.py tree.ged "John Smth" --fuzzy
python gedcom-dna-finder-cli.py tree.ged "John Smth" --fuzzy --fuzzy-threshold 0.75
# Find by exact INDI ID
python gedcom-dna-finder-cli.py tree.ged @I1234@
# Restrict the tag filter to actual DNA matches only (excludes
# "DNA Connection" or "Common DNA Ancestor" if you use those tags)
python gedcom-dna-finder-cli.py tree.ged "Jane Doe" --tag-keyword "DNA Match"
# Return the top 5 nearest matches with a deeper search
python gedcom-dna-finder-cli.py tree.ged "Jane Doe" --top 5 --max-depth 80
Full CLI options
| Flag | Default | Description |
|---|---|---|
--top |
3 | Number of nearest matches to return. |
--max-depth |
50 | Maximum BFS depth, in edges. |
--page-marker |
AncestryDNA Match |
Substring to look for in source-citation PAGE text. Case-insensitive. |
--tag-keyword |
DNA |
Substring to look for in _MTTAG NAME values. Case-insensitive. |
--fuzzy |
off | Enable fuzzy name matching for typos and spelling variants. |
--fuzzy-threshold |
0.6 | Similarity cutoff for --fuzzy, between 0.0 and 1.0. Lower = more matches. |
--list-tags |
Print all _MTTAG definitions in the file and exit. |
|
--list-flagged |
Print every individual currently flagged as a DNA match and exit. |
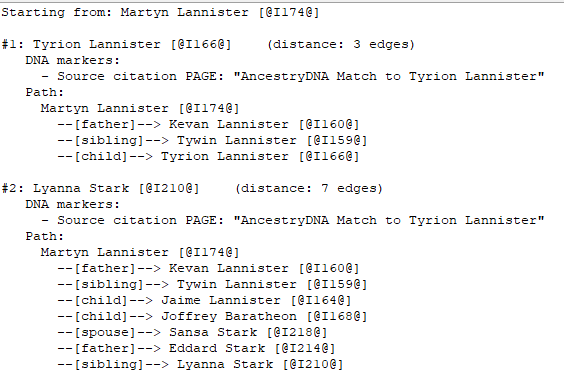
Example output
Starting from: John A. Smith (1850-1920) [@I1234@]
#1: Mary E. Doe (1965-) [@I9876@] (distance: 5 edges)
DNA markers:
- Source citation PAGE: "AncestryDNA Match to Mary E. Doe"
Path:
John A. Smith (1850-1920) [@I1234@]
--[child]--> Robert Smith (1880-1950) [@I1240@]
--[child]--> Helen Smith (1910-1985) [@I1245@]
--[child]--> Janet Smith (1942-) [@I1250@]
--[child]--> Mary E. Doe (1965-) [@I9876@]
Important caveat for Ancestry users
Ancestry's GEDCOM export is well known to be lossy and its handling
of MyTreeTags has varied across versions. If this tool reports far
fewer flagged individuals than you expected, load the file and click
View tag definitions... (or run --list-tags _ from the command
line) to confirm whether your tag records actually made it into the
export. If they did not, the workarounds are:
- Sync the Ancestry tree to Family Tree Maker, add a custom fact (for
example, named
DNA Match) on those individuals in FTM, then export the GEDCOM from FTM. Custom facts in FTM survive the GEDCOM export reliably. - Or rely on the
2 PAGE AncestryDNA Match to ...citation, which is generated automatically by Ancestry when you tag a person as a DNA match while building their tree from a match's profile.
Run the tool with --list-flagged _ (CLI) or use the
"DNA-flagged only" checkbox (GUI) right after loading to confirm the
flagged set looks complete before drawing conclusions from any
individual query.
How "closest" is defined
The tool measures distance as the number of edges traversed in the GEDCOM relationship graph. Each of the following counts as one edge:
- parent ↔ child
- sibling ↔ sibling (within the same
FAMrecord) - spouse ↔ spouse
This means a sibling and a parent are treated as equidistant from ego (both are one edge away), which fits the practical question "how many hops do I need to figure out why this person is in my tree" but is not the same as a genealogical relationship coefficient.
Limitations and notes
- Edge weighting is uniform, as described above. If you would prefer
to weight blood relationships and marriages differently, the
neighbors()function is the place to change it. - The BFS stops as soon as it has accumulated
--topmatches at shortest distances. It does not guarantee a globally optimal cover of all equally close matches if there is a tie at the boundary — raise--topif you suspect ties. - Custom tags whose names happen to contain the substring
DNAwill also be picked up under the default--tag-keyword. Use a more specific keyword, such as"DNA Match", if you want to exclude Ancestry'sDNA ConnectionandCommon DNA Ancestortags. - The tool does not write back to your GEDCOM and does not phone home; it reads the file and prints results.
- Tested with GEDCOM 5.5.1 files exported from Ancestry and Family
Tree Maker. Files from other software should work as long as they
use standard
INDI/FAM/HUSB/WIFE/CHILstructures and one of the two recognized flag formats (or a substitute that you configure via--tag-keywordand--page-marker).
Privacy
The tool runs entirely locally on your machine. Nothing is uploaded.
Be aware, however, that your .ged file likely contains personal
information about living people; do not commit your real GEDCOM to a
public repository or create issues or provide feedback to this repository with any personal data.
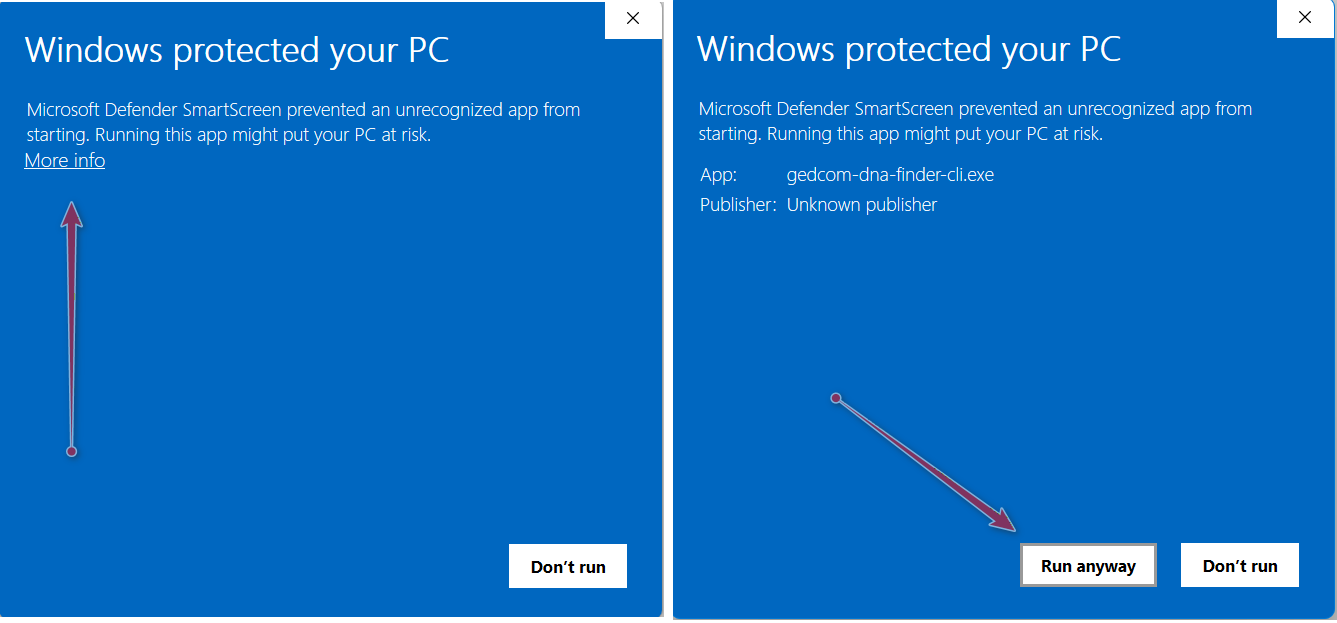
Windows security
You may get a warning from Windows Defender that this is an unrecognized app from an unknown publisher. You can run the application by clicking first on "more info" and then "run anyway." It should only ask the first time you execute the software.
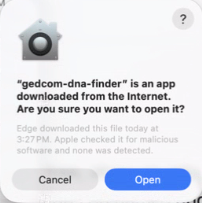
MacOS security
If you are on Mac and not running from the source code, you will have to tell the operating system manually to trust the program. As of 29 April 2026, I am testing a new build process that should only require one click to approve after downloading--just select "open" when presented with the box below the first time you run a new version of the application.
Old instructions
The instructions below apply to older versions of this application and hopefully are no longer necessary.
- Attempt to open the app (it will fail).
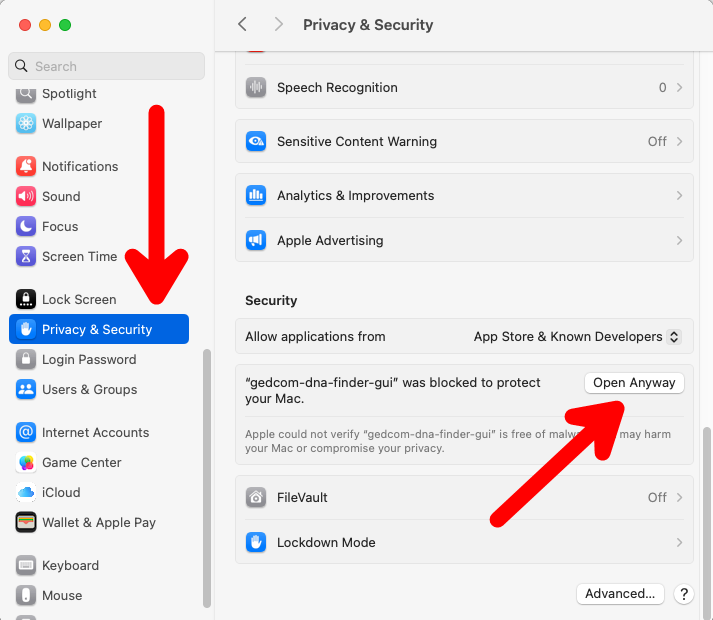
- Open System Settings > Privacy & Security.
- Scroll down to the "Security" section.
- Click "Open Anyway" next to the notification about the blocked app. You will likely need to enter the username and password of an administrator user on the device to approve the application.
If you follow these steps and are seeing an error along the lines of "This file is damaged and can't be opened" it is typically because a false positive from your security settings. This can be fixed by opening the Terminal application (via Applications->Utilities or Spotlight search), typing xattr -cr (with a space after cr) and then dragging and dropping the application ito the Terminal window and hitting enter. This will remove the "quarantine" setting on the application and allow you to run it again.
License
This project is released under the BSD 2-Clause License. See the
LICENSE file for the full text.
Recent changes
See CHANGELOG.
Contributing
Bug reports and pull requests are welcome. If you encounter a GEDCOM file whose tag format is not recognized, please open an issue and include the relevant excerpt (with personal names redacted) so the parser can be extended.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file gedcom_dna_finder-0.2.4.tar.gz.
File metadata
- Download URL: gedcom_dna_finder-0.2.4.tar.gz
- Upload date:
- Size: 5.4 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.13.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
21c37f530bcc824085c2d179da367d60ccecdc6f010609d2498a1f8eba7435ea
|
|
| MD5 |
76dbc1e3c3c83bd09e75c61e87cc1a81
|
|
| BLAKE2b-256 |
33ed4e525d5590f09874e5279b5503183c7e89e21c36581b67f0818d8848a3e1
|
File details
Details for the file gedcom_dna_finder-0.2.4-py3-none-any.whl.
File metadata
- Download URL: gedcom_dna_finder-0.2.4-py3-none-any.whl
- Upload date:
- Size: 5.4 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.13.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
674545f7f6badcf5dee0b60f01961616d9a4b9a742ee049fec082ba71f1ddd17
|
|
| MD5 |
7b4451449fdb685c619fc1183f2d2efd
|
|
| BLAKE2b-256 |
6b547d40a0c86504f49cfbe737a23f74980c6e0730e09e4daeb053737619435c
|