A self-supervised deep learning method for reference-free deconvolution.
Project description
SURF
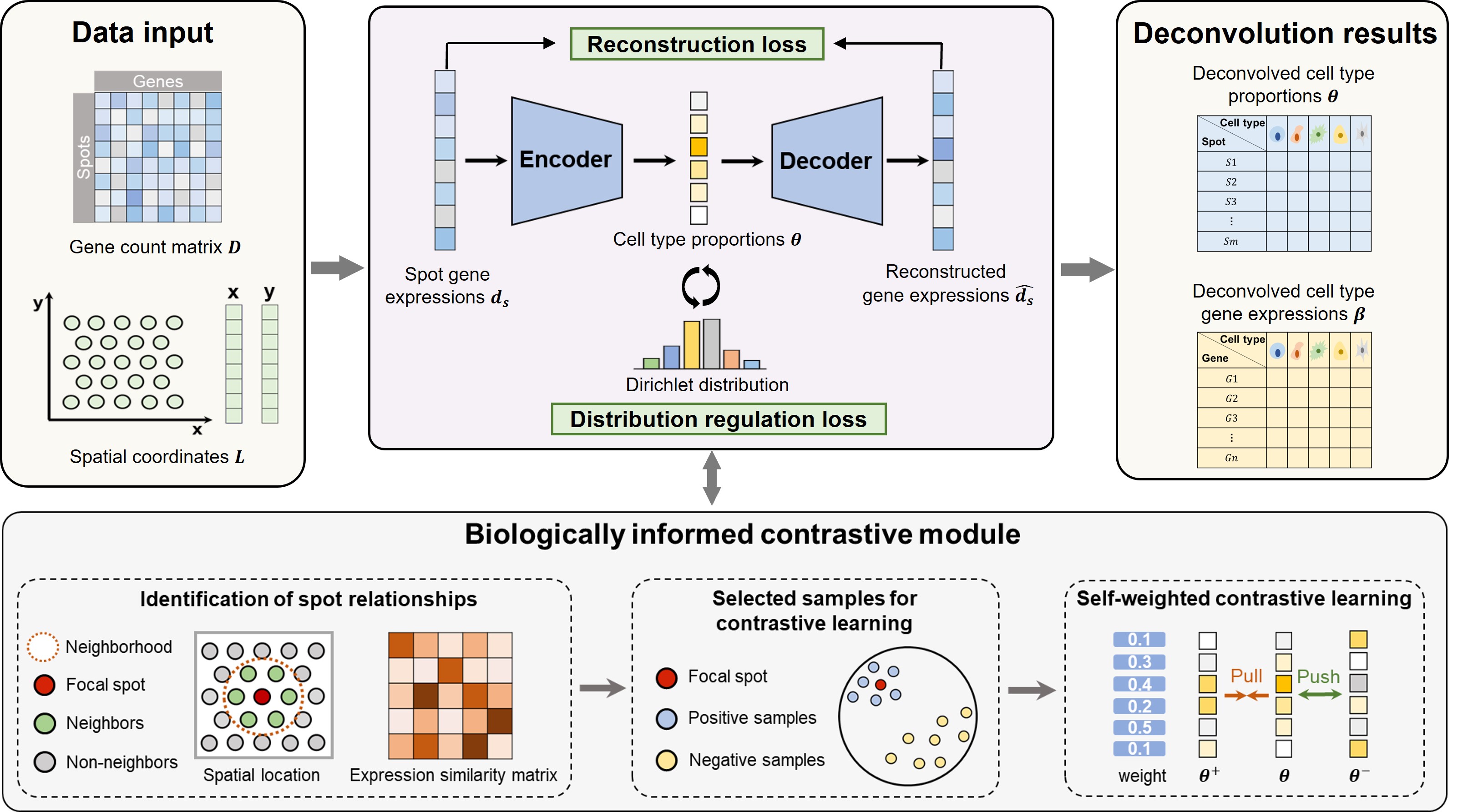
A self-supervised deep learning method for reference-free deconvolution. The overall approach is detailed in the official paper out in xxx.
Data input
df_expr: (n_spots * n_genes), dataframe, with column names (gene names)
df_pos: (n_spots * 2), dataframe, with column names [‘x’, ‘y’]
barcodes: (n_spots,), a numpy array
Data output
‘pred.csv’: The predicted cell types proportions in each spot.
‘beta.csv’: The deconvolved gene expressions of each cell type.
‘last.pkl’: The saved trained model.
Installation
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file genesurf-0.1.tar.gz.
File metadata
- Download URL: genesurf-0.1.tar.gz
- Upload date:
- Size: 8.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.9.20
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
2e1127ba7aa61f197f10938b64dd7706c7808612c9cbb8606819fc4cb47e3de5
|
|
| MD5 |
1896c4e1a126a16d1e1535721cba57bc
|
|
| BLAKE2b-256 |
8c40f52162f9aadee993a1d07e1b2447a2d36597ec8be817e3cada70d64adba1
|
File details
Details for the file genesurf-0.1-py3-none-any.whl.
File metadata
- Download URL: genesurf-0.1-py3-none-any.whl
- Upload date:
- Size: 20.9 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.9.20
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cff8debd88dfc341e8cf2802cc77cbe76a68bffae840eb5ebe863e05d6486e6e
|
|
| MD5 |
e910b11eb3cb88754ce5322e8bb2145d
|
|
| BLAKE2b-256 |
81cc5b13b49d658451c13c52d726a77f1385b2aff28d2d29ec1a848dd1746810
|