Genomes in Python
Project description
Easily install and use genomes in Python and elsewhere!
The goal is to have a simple and straightforward way to download and use genomic sequences. Currently, genomepy supports UCSC, Ensembl and NCBI.
Pssst, hey there! Is genomepy not doing what you want? Does it fail? Is it clunky? Is the documentation unclear? Have any other ideas on how to improve it? Don’t be shy and let us know!
Installation
Genomepy works with Python 3.4+. You can install it via bioconda:
$ conda install genomepy
Or via pip:
$ pip install genomepy
If you install via pip, you will have to install some dependencies, only if you want to use the annotation download feature. You will have to install the following utilities and make sure they are in your PATH:
genePredToBed
genePredToGtf
bedToGenePred
gtfToGenePred
gff3ToGenePred
You can find the binaries here.
Plugins and indexing
By default genomepy generates a file with chromosome sizes and a BED file with gap locations (Ns in the sequence).
For some genomes genomepy can download blacklist files (generated by the Kundaje lab). This will only work when installing these genomes from UCSC. Enable this plugin to use it.
$ genomepy plugin enable blacklist
You can also create indices for some widely using aligners. Currently, genomepy supports:
Note 1: these programs are not installed by genomepy and need to be installed seperately for the indexing to work.
Note 2: the index is based on the genome, annotation (splice sites) is currently not taken into account.
You can configure the index creation using the genomepy plugin command (see below)
Configuration
To change the default configuration, generate a personal config file:
$ genomepy config generate Created config file /home/simon/.config/genomepy/genomepy.yaml
Genome location
By default genomes will be saved in ~/.local/share/genomes.
To set the default genome directory to /data/genomes for instance, edit ~/.config/genomepy/genomepy.yaml and change the following line:
genome_dir: ~/.local/share/genomes/
to:
genome_dir: /data/genomes
The genome directory can also be explicitly specified in both the Python API as well as on the command-line.
Compression
Optionally genome FASTA files can be saved using bgzip compression. This means that the FASTA files will take up less space on disk. To enable this, add the following line to your config file:
bgzip: True
Most tools are able to use bgzip-compressed genome files. One notable exception is bedtools getfasta. As an alternative, you can use the faidx command-line script from pyfaidx which comes installed with genomepy.
Usage
Command line
Usage: genomepy [OPTIONS] COMMAND [ARGS]... Options: --version Show the version and exit. -h, --help Show this message and exit. Commands: config manage configuration genomes list available genomes install install genome plugin manage plugins providers list available providers search search for genomes
Install a genome.
The most important command. The most simple form:
$ genomepy install hg38 UCSC downloading... done... name: hg38 fasta: /data/genomes/hg38/hg38.fa
Here, genomes are downloaded to the directory specified in the config file. To choose a different directory, use the -g option.
$ genomepy install sacCer3 UCSC -g ~/genomes/ downloading from http://hgdownload.soe.ucsc.edu/goldenPath/sacCer3/bigZips/chromFa.tar.gz... done... name: sacCer3 local name: sacCer3 fasta: /home/simon/genomes/sacCer3/sacCer3.fa
You can use a regular expression to filter for matching sequences (or non-matching sequences by using the --no-match option). For instance, the following command downloads hg38 and saves only the major chromosomes:
$ genomepy install hg38 UCSC -r 'chr[0-9XY]+$' downloading from http://hgdownload.soe.ucsc.edu/goldenPath/hg38/bigZips/hg38.fa.gz... done... name: hg38 local name: hg38 fasta: /data/genomes/hg38/hg38.fa $ grep ">" /data/genomes/hg38/hg38.fa >chr1 >chr10 >chr11 >chr12 >chr13 >chr14 >chr15 >chr16 >chr17 >chr18 >chr19 >chr2 >chr20 >chr21 >chr22 >chr3 >chr4 >chr5 >chr6 >chr7 >chr8 >chr9 >chrX >chrY
By default, sequences are soft-masked. Use -m hard for hard masking.
The chromosome sizes are saved in file called <genome_name>.fa.sizes.
For genomes from UCSC and Ensembl, you can choose to download gene annotation files with the --annotation option. These will be saved in BED and GTF format.
$ genomepy install hg38 UCSC --annotation
Finally, in the spirit of reproducibility all selected options are stored in a README.txt. This includes the original name and download location.
Manage plugins.
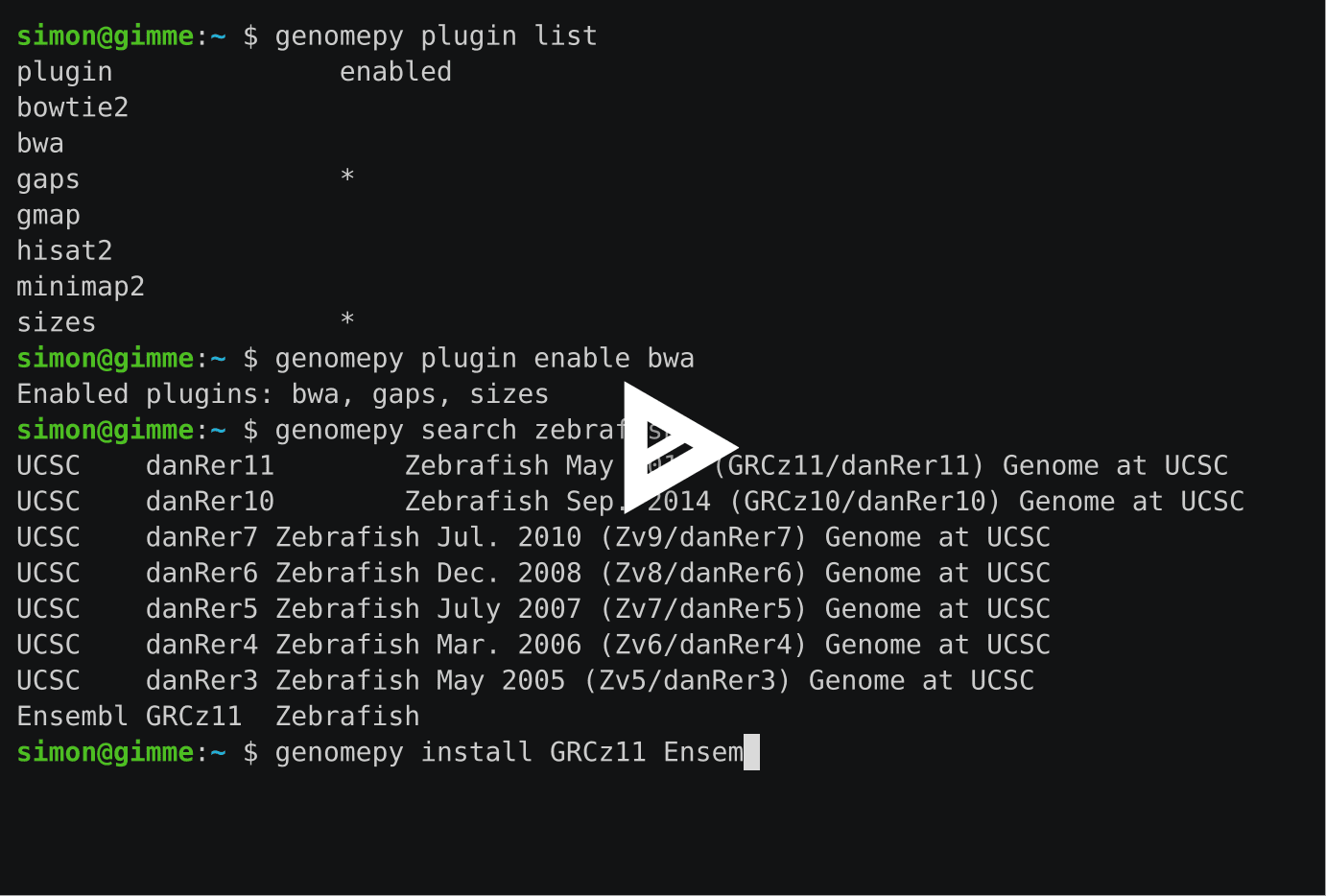
Use genomepy plugin list to view the available plugins.
$ genomepy plugin list plugin enabled bowtie2 bwa gaps * gmap hisat2 minimap2 sizes *
Enable plugins as follows:
$ genomepy plugin enable bwa hisat2 Enabled plugins: bwa, gaps, hisat2, sizes
And disable like this:
$ genomepy plugin disable bwa Enabled plugins: gaps, hisat2, sizes
Search for a genome.
$ genomepy search Xenopus NCBI Xenopus_tropicalis_v9.1 Xenopus tropicalis; DOE Joint Genome Institute NCBI ViralProj30173 Xenopus laevis endogenous retrovirus Xen1; NCBI Xenopus_laevis_v2 Xenopus laevis; International Xenopus Sequencing Consortium NCBI v4.2 Xenopus tropicalis; DOE Joint Genome Institute NCBI Xtropicalis_v7 Xenopus tropicalis; DOE Joint Genome Institute Ensembl JGI 4.2 Xenopus
Only search a specific provider:
$ genomepy search tropicalis -p UCSC UCSC xenTro7 X. tropicalis Sep. 2012 (JGI 7.0/xenTro7) Genome at UCSC UCSC xenTro3 X. tropicalis Nov. 2009 (JGI 4.2/xenTro3) Genome at UCSC UCSC xenTro2 X. tropicalis Aug. 2005 (JGI 4.1/xenTro2) Genome at UCSC UCSC xenTro1 X. tropicalis Oct. 2004 (JGI 3.0/xenTro1) Genome at UCSC
Note that searching doesn’t work flawlessly, so try a few variations if you don’t get any results. Search is case-insensitive.
List available providers
$ genomepy providers Ensembl UCSC NCBI
List available genomes
You can constrain the genome list by using the -p option to search only a specific provider.
$ genomepy genomes -p UCSC UCSC hg38 Human Dec. 2013 (GRCh38/hg38) Genome at UCSC UCSC hg19 Human Feb. 2009 (GRCh37/hg19) Genome at UCSC UCSC hg18 Human Mar. 2006 (NCBI36/hg18) Genome at UCSC ... UCSC danRer4 Zebrafish Mar. 2006 (Zv6/danRer4) Genome at UCSC UCSC danRer3 Zebrafish May 2005 (Zv5/danRer3) Genome at UCSC
Manage configuration
List the current configuration file that genomepy uses:
$ genomepy config file /home/simon/.config/genomepy/genomepy.yaml
To show the contents of the config file:
$ genomepy config show # Directory were downloaded genomes will be stored genome_dir: ~/.local/share/genomes/ plugin: - gaps - sizes
To generate a personal configuration file (existing file will be overwritten):
$ genomepy config generate Created config file /home/simon/.config/genomepy/genomepy.yaml
Local cache.
Note that the first time you run genomepy search or list the command will take a long time as the genome lists have to be downloaded. The lists are cached locally, which will save time later. The cached files are stored in ~/.cache/genomepy and expire after 7 days. You can also delete this directory to clean the cache.
From Python
>>> import genomepy
>>> for row in genomepy.search("GRCh38"):
... print("\t".join(row))
...
UCSC hg38 Human Dec. 2013 (GRCh38/hg38) Genome at UCSC
NCBI GRCh38.p10 Homo sapiens; Genome Reference Consortium
NCBI GRCh38 Homo sapiens; Genome Reference Consortium
NCBI GRCh38.p1 Homo sapiens; Genome Reference Consortium
NCBI GRCh38.p2 Homo sapiens; Genome Reference Consortium
NCBI GRCh38.p3 Homo sapiens; Genome Reference Consortium
NCBI GRCh38.p4 Homo sapiens; Genome Reference Consortium
NCBI GRCh38.p5 Homo sapiens; Genome Reference Consortium
NCBI GRCh38.p6 Homo sapiens; Genome Reference Consortium
NCBI GRCh38.p7 Homo sapiens; Genome Reference Consortium
NCBI GRCh38.p8 Homo sapiens; Genome Reference Consortium
NCBI GRCh38.p9 Homo sapiens; Genome Reference Consortium
Ensembl GRCh38.p10 Human
>>> genomepy.install_genome("hg38", "UCSC", genome_dir="/data/genomes")
downloading...
done...
name: hg38
fasta: /data/genomes/hg38/hg38.fa
>>> g = genomepy.Genome("hg38", genome_dir="/data/genomes")
>>> g["chr6"][166502000:166503000]
tgtatggtccctagaggggccagagtcacagagatggaaagtggatggcgggtgccgggggctggggagctactgtgcagggggacagagctttagttctgcaagatgaaacagttctggagatggacggtggggatgggggcccagcaatgggaacgtgcttaatgccactgaactgggcacttaaacgtggtgaaaactgtaaaagtcatgtgtatttttctacaattaaaaaaaATCTGCCACAGAGTTAAAAAAATAACCACTATTTTCTGGAAATGGGAAGGAAAAGTTACAGCATGTAATTAAGATGACAATTTATAATGAACAAGGCAAATCTTTTCATCTTTGCCTTTTGGGCATATTCAATCTTTGCCCAGAATTAAGCACCTTTCAAGATTAATTCTCTAATAATTCTAGTTGAACAACACAACCTTTTCCTTCAAGCTTGCAATTAAATAAGGCTATTTTTAGCTGTAAGGATCACGCTGACCTTCAGGAGCAATGAGAACCGGCACTCCCGGCCTGAGTGGATGCACGGGGAGTGTGTCTAACACACAGGCGTCAACAGCCAGGGCCGCACGAGGAGGAGGAGTGGCAACGTCCACACAGACTCACAACACGGCACTCCGACTTGGAGGGTAATTAATACCAGGTTAACTTCTGGGATGACCTTGGCAACGACCCAAGGTGACAGGCCAGGCTCTGCAATCACCTCCCAATTAAGGAGAGGCGAAAGGGGACTCCCAGGGCTCAGAGCACCACGGGGTTCTAGGTCAGACCCACTTTGAAATGGAAATCTGGCCTTGTGCTGCTGCTCTTGTGGGGAGACAGCAGCTGCGGAGGCTGCTCTCTTCATGGGATTACTCTGGATAAAGTCTTTTTTGATTCTACgttgagcatcccttatctgaaatgcctgaaaccggaagtgtttaggatttggggattttgcaatatttacttatatataatgagatatcttggagatgggccacaaThe genomepy.Genome() method returns a Genome object. This has all the functionality of a pyfaidx.Fasta object, see the documentation for more examples on how to use this.
Known issues
There might be issues with specific genome sequences. Sadly, not everything (naming, structure, filenames) is always consistent on the provider end. Let me know if you encounter issues with certain downloads.
Todo
Linking genomes to NCBI taxonomy ID
Optionally: Ensembl bacteria (although there might be better options specifically for bacterial sequences)
Citation
If you use genomepy in your research, please cite it: 10.21105/joss.00320.
Getting help
If you want to report a bug or issue, or have problems with installing or running the software please create a new issue. This is the preferred way of getting support. Alternatively, you can mail me.
Contributing
Contributions welcome! Send me a pull request or get in touch.
License
This module is licensed under the terms of the MIT license.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
File details
Details for the file genomepy-0.6.1.tar.gz.
File metadata
- Download URL: genomepy-0.6.1.tar.gz
- Upload date:
- Size: 24.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/1.13.0 pkginfo/1.5.0.1 requests/2.22.0 setuptools/41.0.1 requests-toolbelt/0.9.1 tqdm/4.32.2 CPython/3.6.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
2caafeb2848cc129b68fa8a76bfed027a585ec9d3fd96b29f22b48caafc1b09c
|
|
| MD5 |
7d174e014f48369b6e035c704f91cb2f
|
|
| BLAKE2b-256 |
28ab7394f19245f797cf6fb383963b6dd692db42780804a84c811b4abc0160d8
|