Module for kinetic RNA-seq analysis.
Project description

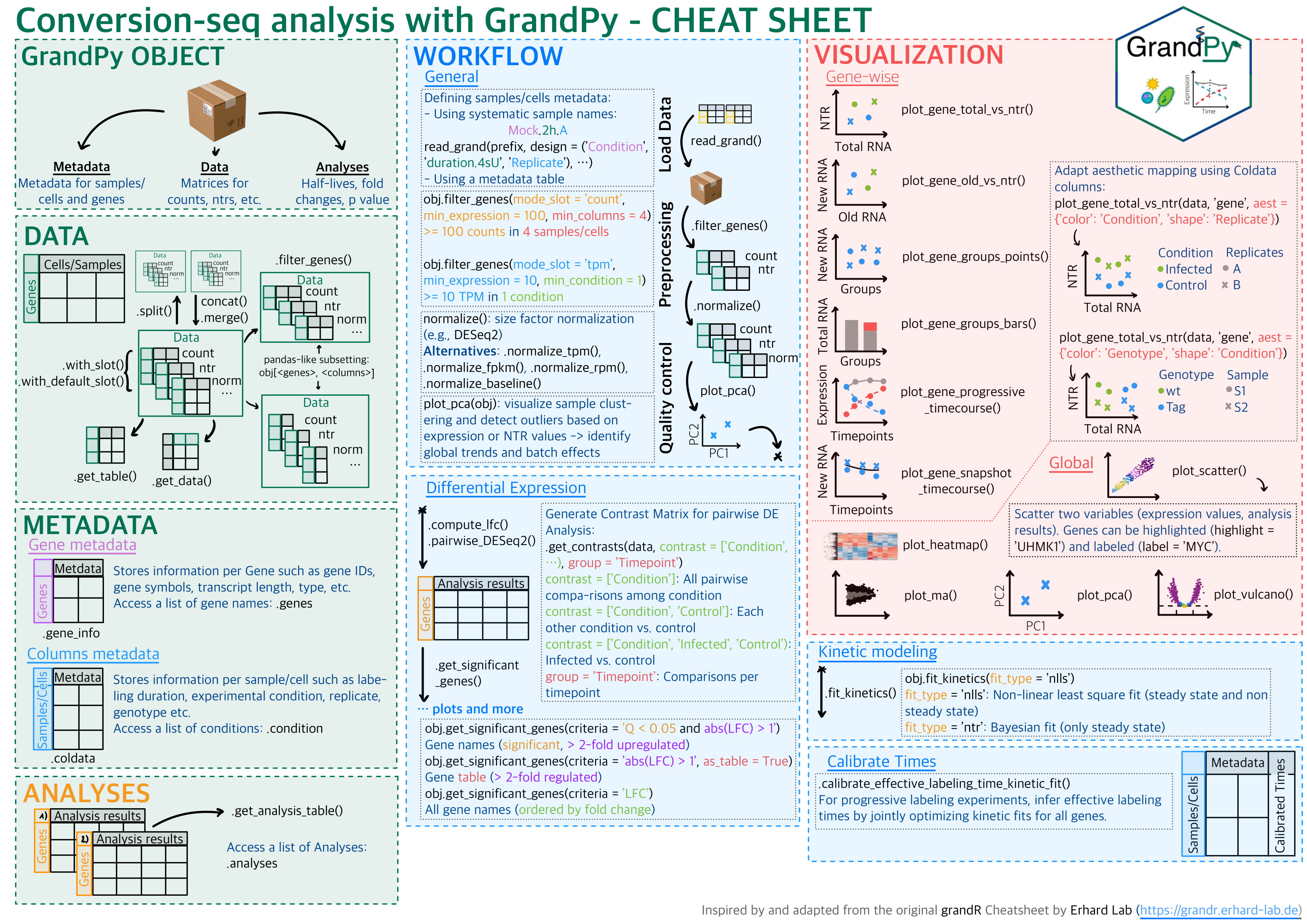
GrandPy
Nucleotide conversion sequencing experiments have been developed to add a temporal dimension to RNA-seq and single-cell RNA seq. Such experiments require specialized tools for primary processing such as GRAND-SLAM, and specialized tools for downstream analyses. GrandPy provides a comprehensive toolbox for quality control, kinetic modeling, differential gene expression analysis and visualization of such data. It mimics the core functionality of the original grandR package, by which it is inspired.
Installation
GrandPy can be installed using the following command:
pip install grandpy-lib
You can also install the development version directly from GitHub:
pip install git+https://github.com/maus2310/GrandPy
System Requirements
GrandPy has mostly been tested on Windows but should also run on Linux and macOS. The package runs on standard laptops (multicore CPUs are recommended; memory requirements depend on the size of your datasets).
Requires python 3.12.0
Installing it via pip will make sure that the following (standard) packages are available:
numpy, pandas, scipy, anndata, tqdm, matplotlib, seaborn, scanpy, pydeseq2
Additional packages are optional and important for particular functions:
scikit-learn, mygene, numdifftools
Cheatsheet
How to get started
First, have a look at the getting started notebook.
Next, explore differential expression or kinetic modeling, which provide an overview of the two primary settings for nucleotide conversion experiments.
There are also additional notebooks:
- Loading data and working with GrandPy objects: Learn more about programming with GrandPy
- Working with data matrices and analysis results: Learn more about how to retrieve data from GrandPy objects
- Plotting: Learn how to create and store plots with GrandPy
- Pulse-chase: Learn how to fit pulse-chase data with GrandPy
Acknowledgements
GrandPy is heavily inspired by the grandR R package by Prof. Dr. Florian Erhard,
Teresa Rummel, Lygeri Sakellaridi and Kevin Berg. We gratefully acknowledge their work,
as well as the efforts of the team behind it, especially Julian
Selke and Rahaf Issa, whose contributions made GrandPy possible.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file grandpy_lib-0.1.1.tar.gz.
File metadata
- Download URL: grandpy_lib-0.1.1.tar.gz
- Upload date:
- Size: 103.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.13.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
0f53b6c7e293299f6fcd7b3a34608b466cdd9e220baf4af61ec3423356d6abbf
|
|
| MD5 |
d2ce4e57c21281f4a42a9f77ff68a2be
|
|
| BLAKE2b-256 |
25ab1adf3a1cd29ee0193952c8edd837baf4eb48f1b361311c50c1fe7444cd2b
|
File details
Details for the file grandpy_lib-0.1.1-py3-none-any.whl.
File metadata
- Download URL: grandpy_lib-0.1.1-py3-none-any.whl
- Upload date:
- Size: 107.1 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.13.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
193e34b2def5a967c52699c9d3e3d7326cac000740d71c29dc3e10c64567db32
|
|
| MD5 |
04855f9bec67ea9399a730b87a56caa3
|
|
| BLAKE2b-256 |
a4dd9f1110973b1c29224d37057648f0329dfdd1916045f5ed313a5a5f1d7089
|