Estimate growth curves using an Extended Kalman Filter
Project description
grpredict
The study of the growth of bacterial cultures does not constitute a specialized subject or branch of research: it is the basic method of microbiology.” Monitoring and controlling the specific growth rate should not constitute a specialized subject or branch of research: it should be the basic method of advanced microbial bioprocessing.
- Jacques Monod
The first step to controlling a culture's growth is to measure it. To do this, we need to model the rate of growth. We've seen other attempts at modelling the growth rate, but they all assumed too much:
- Exponential growth model: perhaps truly early on, but certainly not the case near stationary.
- Logistic growth model: requires an assumption about the max optical density, and assumes symmetric growth around the inflection point.
- Gompertz models and other models: more parameters, less interpretable, and still make strong assumptions about the trajectory of growth.
Further more, none of these can model a culture in a turbidostat or chemostat mode, where the optical density is dropping quickly during a dilution, but the growth rate should remain constant.
Re-visiting a growth model
We are often introduced to a growth model by the simple exponential growth model:
$\text{OD}(t) = \exp{ \text{gr} t }$
Plainly put, the culture grows exponentially at rate $\text{gr}$. Like we mentioned above, this might be true for small time-scales, but certainly over the entire lag, log, and then stationary phases, this is not the case.
There's a hint of an interesting idea in the last paragraph though: "over small time scales". What if we cut up the growth curve into many small time intervals, and computed a growth rate for each interval? Then our growth rate can be changing: starts near 0 in the lag phase, can increase to a peak in the log phase, and then drop to 0 again in the stationary phase. We also don't need to assume any parametric form for the growth rate, we can just measure it directly from the data.
Our new formula might look like:
$$\text{OD}(t) = \exp{ \left( \text{gr}_0 \Delta t + \text{gr}_1 \Delta t + ... \right)}$$
If we think more about this, and we keep shrinking our time interval towards zero, this is just an integral:
$$ \text{nOD}(t) = \exp{ \left( \int_0^t \text{gr}(s)ds \right)} $$
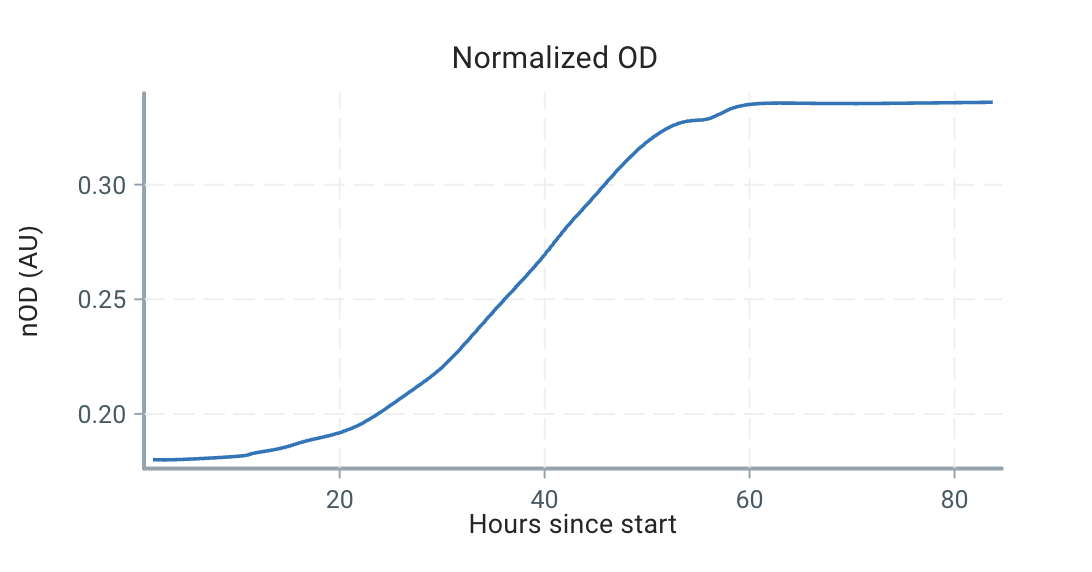
There's still no particular assumption about the shape of the growth rate function, $gr(s)$. For example, consider the following ODs from a batch experiment:
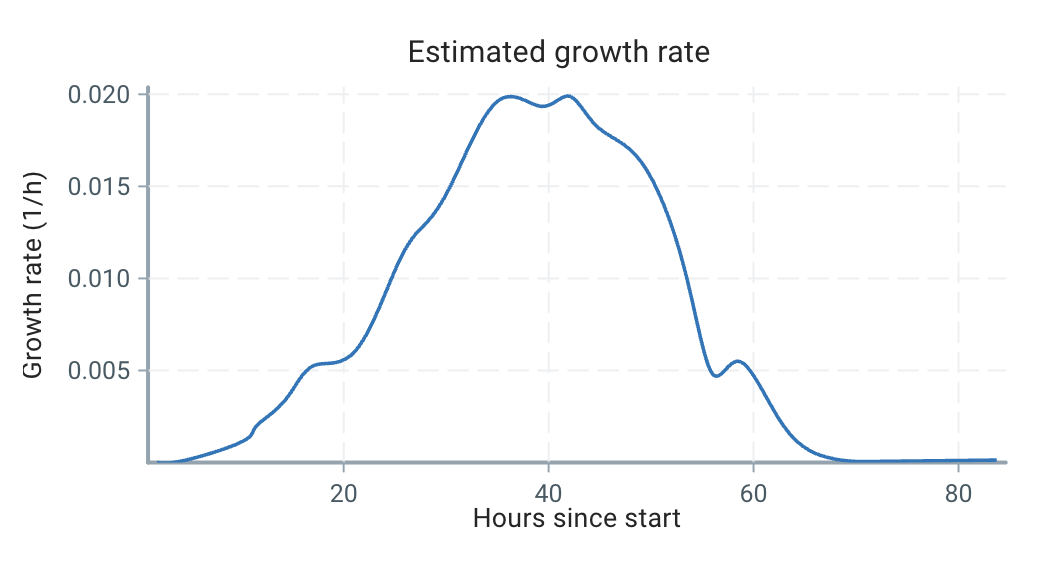
Using our estimation technique outlined below, we can estimate the growth curve as:
We can see that the growth is very dynamic, and certainly not a single number or a constrained form!
Estimating a non-parametric growth rate
A non-parametric, dynamic growth rate sounds great, but we've replaced a estimation of a single value (or handle of values) to an entire function! This seems expensive!
Luckily, we do the work for you. We will compute $gr(t)$ using a statistical algorithm, the Kalman filter. The Kalman filter is an algorithm that estimates the state of a dynamic system from noisy measurements. In our case, the state is the growth rate, and the measurements are the optical density readings. We input optical density observations one at a time, and the Kalman filter updates its estimate of the growth rate based on the new observation. This allows us to track the growth rate in real-time, as the culture grows.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file grpredict-26.2.10.tar.gz.
File metadata
- Download URL: grpredict-26.2.10.tar.gz
- Upload date:
- Size: 13.1 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
3738e41f5657badd4ff7bceb685d2769c12bb4ca3292167799b8a21dc67b1c63
|
|
| MD5 |
e840a0ba62343a797b0ad671809f0df5
|
|
| BLAKE2b-256 |
dc119d81630adc96e0d19f4d7150a66210c22763637e86f25d1cdd9d884fbc9b
|
File details
Details for the file grpredict-26.2.10-py3-none-any.whl.
File metadata
- Download URL: grpredict-26.2.10-py3-none-any.whl
- Upload date:
- Size: 9.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
853a2b9fd789d282820df6d7d38923c0ddbcc602708b11eb61f872eed296d440
|
|
| MD5 |
120174cdd7f3377f5848f4e1b74f1c9e
|
|
| BLAKE2b-256 |
9702fde1e94bde44137c21f90b735931f1e4cb21cd4ce9ed2a0c5cbc06012a72
|