gwaspeek: interactive terminal viewer for GWAS summary statistics
Project description
gwaspeek
- Interactive terminal Manhattan plots for GWAS summary statistics: move the view along the genome and zoom in a real TTY, inspect regions, and optionally show a protein-coding gene track when the view is a single chromosome and spans ≤ 1 Mb.
- Static one-shot renders (
-s) for logs or CI. - Column names are auto-detected from bundled formatbook-style aliases.
- Chromosome layout uses full canonical human chromosome lengths for GRCh37 or GRCh38.
- Drawing uses Unicode or
--ascii.
Note: Intended for quick checks of summary statistics. Plotted positions are not exact on screen because the terminal has limited pixels (resolution).
Interactive mode (default)
Interactive mode (default) opens the live TTY viewer when you pass one input path—gwaspeek FILE or gwaspeek -i FILE.
In the viewer, a / d pan the plot in smaller steps (and A / D pan in larger steps). w / s zoom out and in gently; W / S use the original coarser zoom. Arrow keys and + / - mirror the fine controls, the mouse wheel zooms at the hovered cursor position, and g jumps directly to a typed region such as chr3:45000000-46000000.
Try it
After installation, from a directory that contains the file:
gwaspeek t2d_bbj_p1e-5.txt.gz
Keys
| Key | Action |
|---|---|
a / d |
Pan view left/right (fine) |
A / D |
Pan view left/right (coarse) |
w / s |
Zoom out / in (fine) |
W / S |
Zoom out / in (coarse) |
| Arrow keys | Fine pan / zoom |
+ / - |
Zoom in / out (fine) |
| Mouse wheel | Zoom at cursor position |
g |
Jump to region (chr:start-end) |
l |
Toggle lead-variant labels |
t |
Cycle gene track: 37 → 38 → off |
v |
Toggle variants-in-view list |
h |
Toggle help |
r |
Reset to full genome |
q |
Quit |
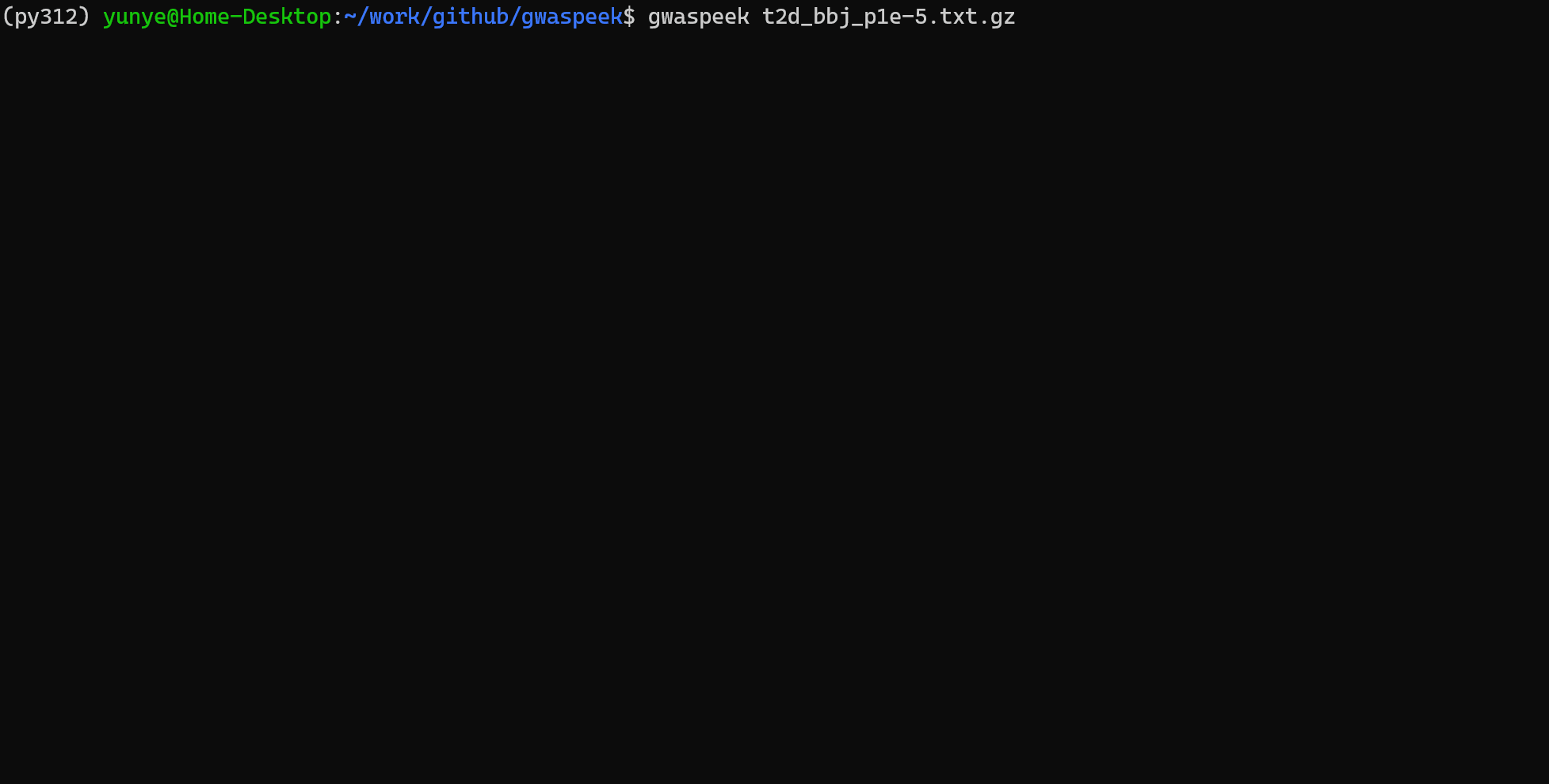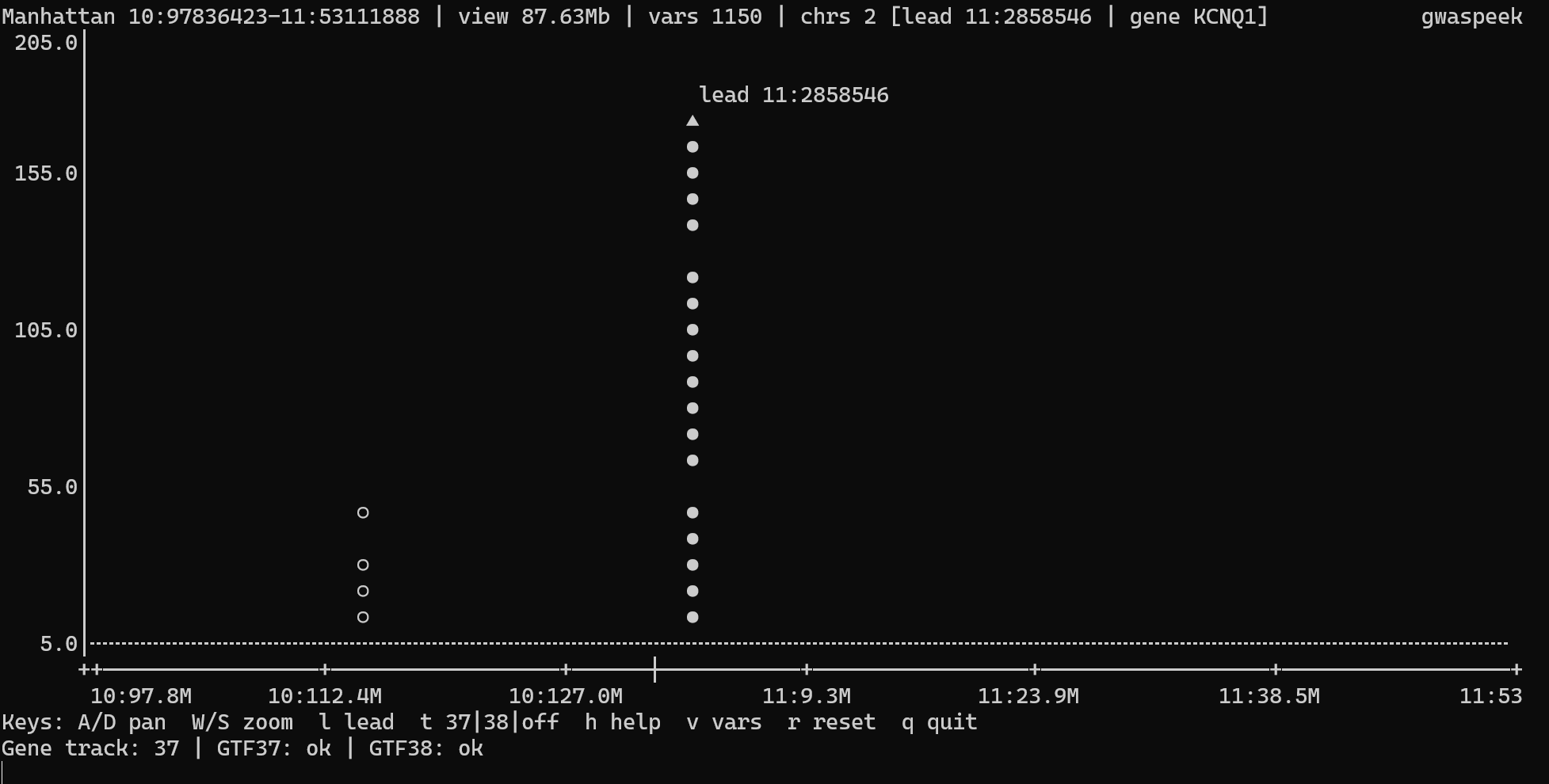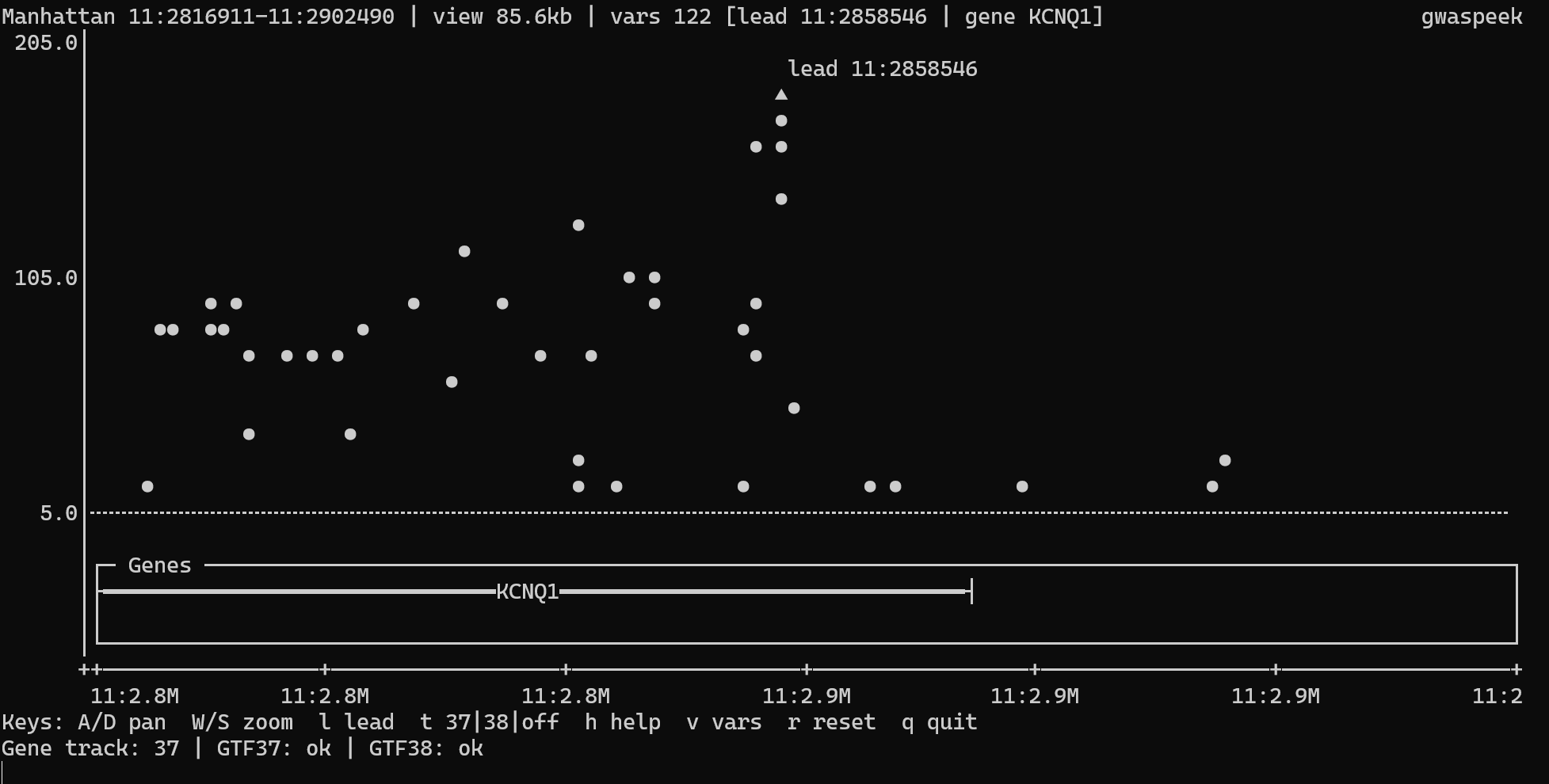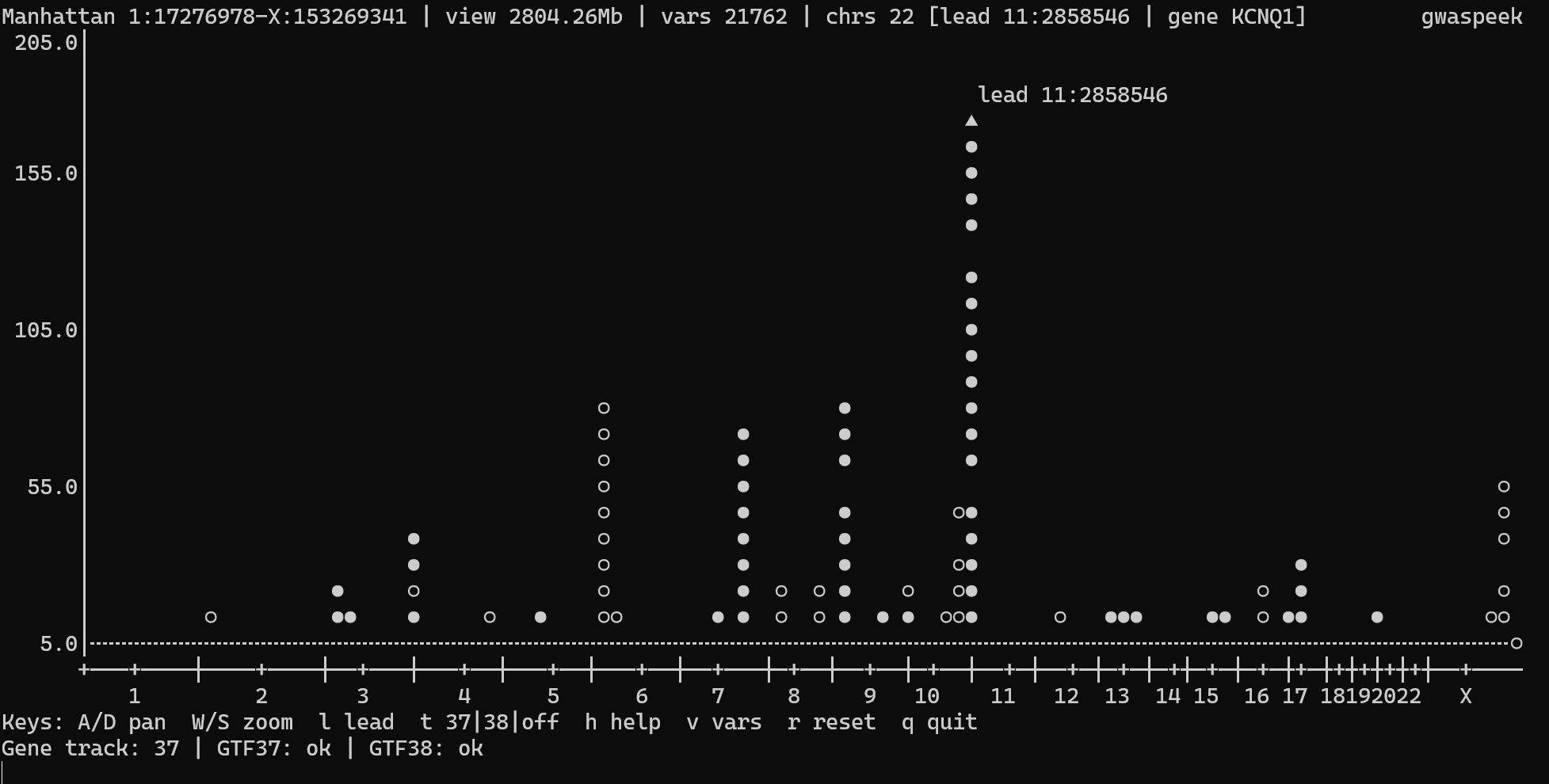
The gene track appears only on a single-chromosome view with span ≤ 1 Mb. --gtf and --gtf38 (see CLI reference) apply only in interactive mode; static -s output ignores them. --build sets the canonical chromosome-length layout, and interactive t build switching keeps the layout aligned with the active gene-track build.
The interactive viewer uses an alternate terminal screen, lazy-loads gene annotations by genome build, and renders dense cells with heavier glyphs instead of simply dropping overlapping points.
Common sizing and column flags work in interactive mode too (for example --width, --height, --skip, --sig-level):
gwaspeek sumstats.tsv \
--sep "\t" \
--width 120 \
--height 32 \
--skip 2 \
--sig-level 5e-8
Installation
pip install gwaspeek
gwaspeek --help
Static mode (-s)
For a one-shot text snapshot (non-interactive), add -s before the path:
gwaspeek -s t2d_bbj_p1e-5.txt.gz --width 120 --ascii
Sample output:
Manhattan 1:17276978-X:153269341 gwaspeek
205.0│
│
│
│
│ ●
│ ●
155.0│ ●
│ ●
│
│
│ ●
│
105.0│ ●
│ ●
│ ● ●
│ ○ ● ●
│ ○ ● ● ●
55.0│ ○ ● ○
│ ○ ● ● ○ ○
│ ● ○ ● ○● ○
│ ● ○ ○ ● ○ ● ○○ ●
│ ○ ●● ● ○ ● ○○ ○ ● ● ○ ○ ● ● ● ○○○ ○○ ●● ● ●● ○ ●● ○ ● ○○
5.0│╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌○
+───+─────────+───────+───────+──────+──────+─────+─────+─────+─────+────+────+───+───+──+───+──+──+─+─+─+───+────
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18192022 X
Invocation at a glance
| Form | Mode |
|---|---|
gwaspeek FILE |
Interactive (same as -i) |
gwaspeek -i FILE |
Interactive |
gwaspeek -s FILE |
Static |
Input files
Tabular GWAS summary stats (TSV/CSV or other delimiter via --sep).
Per row you need: chromosome, base-pair position, and either P or -log10(P) (names are auto-detected; override with --chr, --pos, --p, --mlog10p). Do not pass both --p and --mlog10p. Internally rows are normalized to CHR, POS, P, and derived mlog10p.
Chromosome tokens understood by the parser include numeric 1–22, X, Y, MT, and chr-prefixed forms (e.g. chr1, chrX, chrM).
CLI reference
| Option | Type | Default | Description |
|---|---|---|---|
FILE (positional) |
path | — | Same as -i FILE (interactive). |
-i, --interactive FILE |
path | — | Interactive viewer. |
-s, --static FILE |
path | — | One-shot static plot. |
--sep STR |
string | tab (\t) |
Delimiter. |
--chr NAME |
string | auto | Chromosome column. |
--pos NAME |
string | auto | Position column. |
--p NAME |
string | auto | P-value column (not with --mlog10p). |
--mlog10p NAME |
string | auto | -log10(P) column (not with --p). |
--skip FLOAT |
float | 5.0 |
Drop variants below this -log10(P); also y-axis floor. |
--width INT |
int | 100 |
Width (chars): static size; interactive initial/fallback. |
--height INT |
int | 28 |
Height (lines): static size; interactive initial/fallback. |
--ascii |
flag | off | ASCII drawing instead of Unicode. |
--build {37,38} |
choice | 37 |
Canonical chromosome-length layout / default genome build. |
--no-color |
flag | off | Disable ANSI color in interactive status/footer lines. |
--sig-level FLOAT |
float | 5e-8 |
Genome-wide line in P space. |
--ymax FLOAT |
float | auto | Y-axis max in -log10(P). |
--gtf PATH |
path | bundled GRCh37 | GTF (.gz ok); interactive gene track. |
--gtf38 PATH |
path | — | GRCh38 GTF (.gz ok); interactive mode uses bundled gene-only GTF when omitted. |
-h, --help |
flag | off | Help. |
Examples
# Interactive (default): positional or -i
gwaspeek tests/fixtures/sumstats_small.tsv
gwaspeek -i tests/fixtures/sumstats_small.tsv
# Static
gwaspeek -s tests/fixtures/sumstats_small.tsv
gwaspeek -s tests/fixtures/sumstats_small.tsv --build 38
# Explicit columns (flags are the same in interactive mode)
gwaspeek -s sumstats.tsv --chr CHROM --pos BP --p PVAL
# Precomputed -log10(P)
gwaspeek -s sumstats_mlog10p.tsv --chr CHR --pos POS --mlog10p LOG10P
# Narrow terminals
gwaspeek -s sumstats.tsv --ascii
Tips
- Dense sumstats: raise
--skipto thin points and reduce clutter. - Unicode is usually clearer;
--asciiwhen the terminal cannot draw box drawing / braille well.
License
MIT.
Repository
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file gwaspeek-0.2.0.tar.gz.
File metadata
- Download URL: gwaspeek-0.2.0.tar.gz
- Upload date:
- Size: 5.7 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
50c5227b43e14ac29b41c3f6cd4b40a85091d868de8ae75e2a9b5ad7da6b8693
|
|
| MD5 |
e719598c68a425299139a9adb74f9d9b
|
|
| BLAKE2b-256 |
5e43b52778248374a38a62ffa0d6fc364738494173aae3488168796fe952526e
|
File details
Details for the file gwaspeek-0.2.0-py3-none-any.whl.
File metadata
- Download URL: gwaspeek-0.2.0-py3-none-any.whl
- Upload date:
- Size: 5.7 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
482e8c1abaf55e36f39922c321d3b8d793948bf85d36bcae24601ef0d4e25490
|
|
| MD5 |
8680895e77e1e808cf958b4b44549ccc
|
|
| BLAKE2b-256 |
e63fba641488a55a7436455c217f4cb34d50b68992099737ad42c1d927af2f97
|











