Hydrogen Bond Analysis Tool for PDB structures
Project description
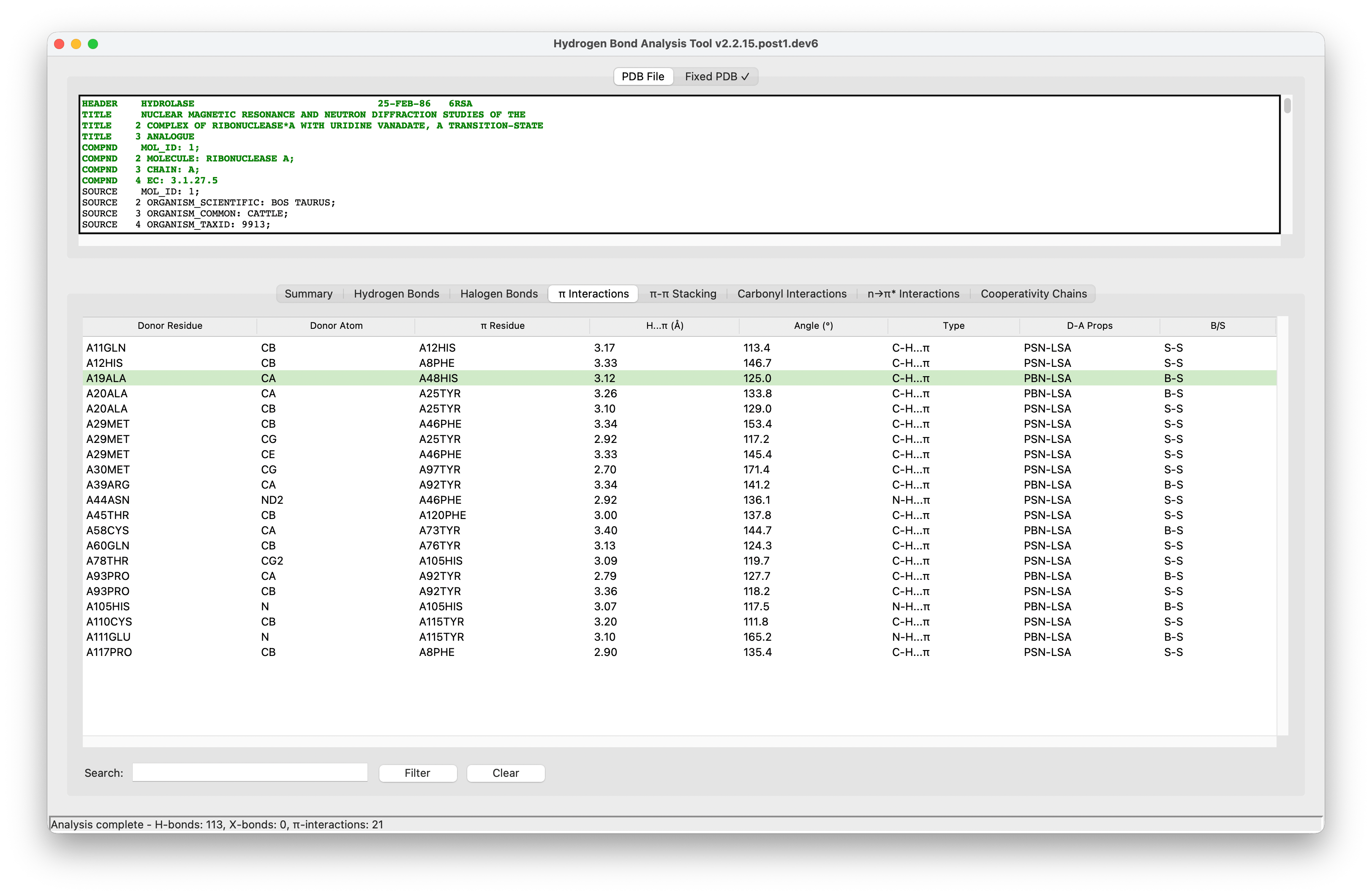
Hydrogen Bond Analysis Tool (HBAT) v2
A Python package to automate the analysis of potential hydrogen bonds and similar type of weak interactions in macromolecular structures, available in Protein Data Bank (PDB) file format. HBAT uses a geometric approach to identify molecular interactions by analyzing distance and angular criteria.
Supported Interaction Types:
- Hydrogen Bonds: Classical
N-H···O,O-H···O, and weakC-H···Ointeractions - Halogen Bonds:
C-X···Ainteractions (X = Cl, Br, I) - π Interactions: X-H···π and
C-X···πinteractions with aromatic rings (Phe,Tyr,Trp,His, etc.) - π-π Stacking: Aromatic ring-ring interactions (parallel, T-shaped, offset)
- Carbonyl Interactions:
n→π*interactions between carbonyl groups - n-π Interactions: Lone pair interactions with aromatic
πsystems
Background
HBAT v2 is a modern Python re-implementation of the original Perl-based tool developed by Abhishek Tiwari and Sunil Kumar Panigrahi. HBAT v1 can still be downloaded from SourceForge however Perl version is not maintained anymore.
Highlights of HBAT v2
- Detect and analyze potential hydrogen bonds, halogen bonds, π interactions, π-π stacking, carbonyl interactions, and n-π interactions
- Automated PDB fixing with OpenBabel and PDBFixer integration
- Support graphical (tkinter), command-line, and programming API interfaces
- Use graphical interfaces for interactive analysis, CLI/API for batch processing and automation
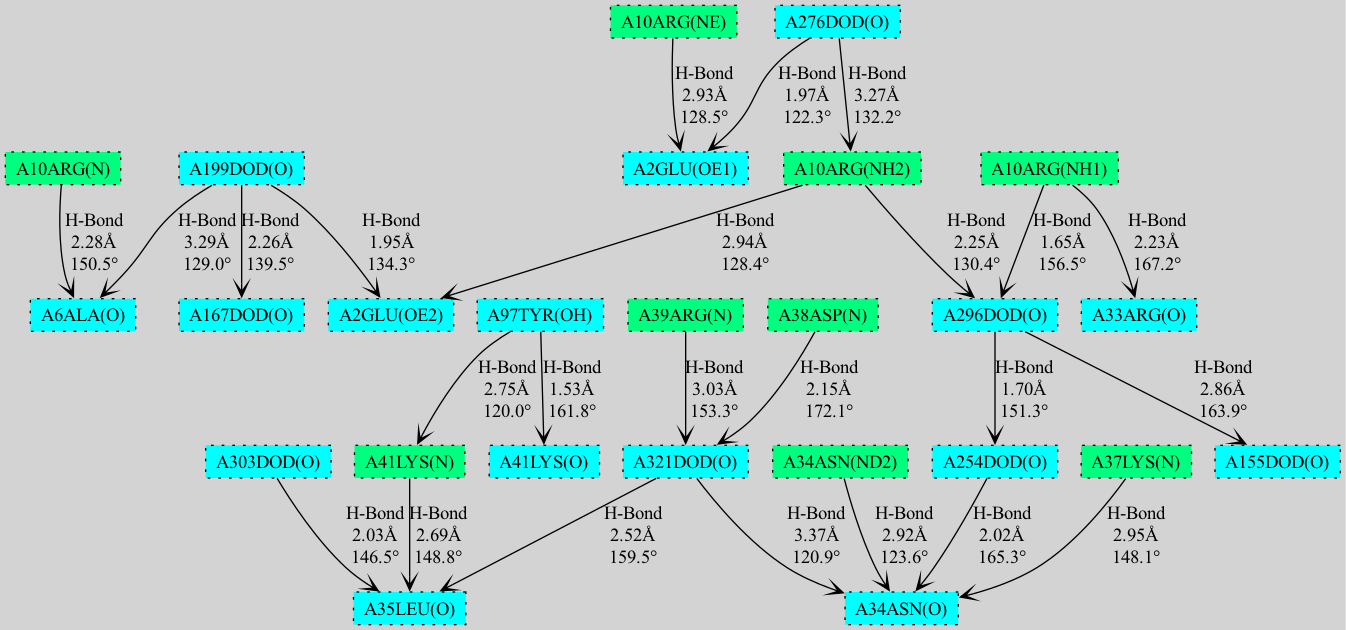
- Cooperativity chain visualization using NetworkX/matplotlib and GraphViz
- Export cooperativity chain visualizations to PNG, SVG, PDF formats
- Built-in presets for different structure types (high-resolution, NMR, membrane proteins, etc.)
- Customizable distance cutoffs, angle thresholds, and analysis modes.
- Multiple Output Formats: Text, CSV, and JSON export options
- Optimized algorithms for efficient analysis of large structures
- Cross-Platform: Works on Windows, macOS, and Linux.
Please review HBAT documentation for more details.
Installation
Option 1: Install from PyPI (Recommended)
pip install hbat
Run HBAT Command-Line Interface (CLI) using hbat or launch HBAT GUI using hbat-gui.
Option 2: Install from Source
git clone https://github.com/abhishektiwari/hbat.git
cd hbat
pip install -e .
Alternatively,
pip install git+https://github.com/abhishektiwari/hbat.git
Run HBAT Command-Line Interface (CLI) using hbat or launch HBAT GUI using hbat-gui.
Option 3: Install from Conda
conda install -c hbat hbat
Requirements
System Requirements
- Python: 3.9 or higher
- tkinter: tkinter is included with Python standard library on most systems. However, on Mac install Python and tkinter using
brew.
brew install python python3-tk
- GraphViz (Optional): Required for advanced cooperativity chain visualization with high-quality graph rendering. HBAT will automatically fall back to NetworkX/matplotlib visualization if GraphViz is not available.
Install GraphViz:
On Ubuntu/Debian:
sudo apt-get update
sudo apt-get install graphviz
On macOS (using Homebrew):
brew install graphviz
On Windows:
- Download and install from GraphViz official website
- Or using Chocolatey:
choco install graphviz - Or using conda:
conda install -c conda-forge graphviz
Note: After installing GraphViz, restart your terminal/command prompt before running HBAT to ensure the GraphViz executables are available in your PATH.
Usage
Graphical Interface
Launch the GUI application:
hbat-gui
The GUI provides,
- File browser for loading PDB files
- Parameter configuration panels
- Tabbed results display
- Export and visualization options
Command-Line Interface
Basic usage:
hbat input.pdb
Output Format Options
HBAT supports multiple output formats:
# No output flag - displays results to console
hbat input.pdb # Display results to console
# Single file outputs (format auto-detected from extension)
hbat input.pdb -o results.txt # Text format (human-readable summary + details)
hbat input.pdb -o results.json # JSON format (single file with all interactions)
# Multiple file outputs (separate files per interaction type)
hbat input.pdb --csv results # Creates results_h_bonds.csv, results_x_bonds.csv, etc.
hbat input.pdb --json results # Creates results_h_bonds.json, results_x_bonds.json, etc.
With custom parameters:
hbat input.pdb -o results.txt --hb-distance 3.0 --mode local
List Available Presets
hbat --list-presets
Use a specific preset
hbat protein.pdb --preset high_resolution
hbat membrane_protein.pdb --preset membrane_proteins
Use preset with custom overrides
hbat protein.pdb --preset drug_design_strict --hb-distance 3.0 --verbose
CLI Options
positional arguments:
input Input PDB file
optional arguments:
-h, --help show this help message and exit
-o OUTPUT, --output OUTPUT
Output file (format auto-detected from extension: .txt, .json)
--json JSON Export to multiple JSON files (base name for files)
--csv CSV Export to multiple CSV files (base name for files)
Preset Options:
--preset PRESET Load parameters from preset file (.hbat or .json)
--list-presets List available example presets and exit
Analysis Parameters:
--hb-distance HB_DISTANCE
Hydrogen bond H...A distance cutoff in Å (default: 3.5)
--hb-angle HB_ANGLE Hydrogen bond D-H...A angle cutoff in degrees (default: 120)
--da-distance DA_DISTANCE
Donor-acceptor distance cutoff in Å (default: 4.0)
--xb-distance XB_DISTANCE
Halogen bond X...A distance cutoff in Å (default: 4.0)
--xb-angle XB_ANGLE Halogen bond C-X...A angle cutoff in degrees (default: 120)
--pi-distance PI_DISTANCE
π interaction H...π distance cutoff in Å (default: 4.5)
--pi-angle PI_ANGLE π interaction D-H...π angle cutoff in degrees (default: 90)
--covalent-factor COVALENT_FACTOR
Covalent bond detection factor (default: 1.2)
--mode {complete,local}
Analysis mode: complete (all interactions) or local (intra-residue only)
Output Control:
--verbose, -v Verbose output with detailed progress
--quiet, -q Quiet mode with minimal output
--summary-only Output summary statistics only
Analysis Filters:
--no-hydrogen-bonds Skip hydrogen bond analysis
--no-halogen-bonds Skip halogen bond analysis
--no-pi-interactions Skip π interaction analysis
License
This project is licensed under the MIT License - see the LICENSE file for details.
Citation
If you use HBAT in your research, please cite:
@article{tiwari2007hbat,
author = {Tiwari, Abhishek and Panigrahi, Sunil Kumar},
doi = {10.3233/ISI-2007-00337},
journal = {In Silico Biology},
month = dec,
number = {6},
title = {{HBAT: A Complete Package for Analysing Strong and Weak Hydrogen Bonds in Macromolecular Crystal Structures}},
volume = {7},
year = {2007}
}
Contributing
See our contributing guide and development guide. At a high-level,
- Fork the repository
- Create a feature branch
- Make your changes
- Add tests if applicable
- Submit a pull request
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file hbat-2.2.15rc33177633022.tar.gz.
File metadata
- Download URL: hbat-2.2.15rc33177633022.tar.gz
- Upload date:
- Size: 2.5 MB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
2f24e657f6870a1f86920f39b4b9204d3068092531c87d2a4e63be388d763159
|
|
| MD5 |
66d679a97d0802e8c0599ca282a98246
|
|
| BLAKE2b-256 |
650cfc2cf7b2210b858b21f980e2e1003a1f2e46fb8d22b4ed070918f1f04785
|
Provenance
The following attestation bundles were made for hbat-2.2.15rc33177633022.tar.gz:
Publisher:
release.yml on abhishektiwari/hbat
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
hbat-2.2.15rc33177633022.tar.gz -
Subject digest:
2f24e657f6870a1f86920f39b4b9204d3068092531c87d2a4e63be388d763159 - Sigstore transparency entry: 707092309
- Sigstore integration time:
-
Permalink:
abhishektiwari/hbat@a9676fe7c2ac383bf351773c2d6014cadbd7a607 -
Branch / Tag:
refs/pull/33/merge - Owner: https://github.com/abhishektiwari
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yml@a9676fe7c2ac383bf351773c2d6014cadbd7a607 -
Trigger Event:
pull_request
-
Statement type:
File details
Details for the file hbat-2.2.15rc33177633022-py3-none-any.whl.
File metadata
- Download URL: hbat-2.2.15rc33177633022-py3-none-any.whl
- Upload date:
- Size: 178.1 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
2560eacfeadc7d31d268f377d34e1134a8665c3fc3374ce0c91bd7b3a0f6f54a
|
|
| MD5 |
a44d852bf695f96663d770fe6601370d
|
|
| BLAKE2b-256 |
1e11d3abf13be75f7f20b1b3b3507f9af1811c0c4ad584dae06c4b4832c0836d
|
Provenance
The following attestation bundles were made for hbat-2.2.15rc33177633022-py3-none-any.whl:
Publisher:
release.yml on abhishektiwari/hbat
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
hbat-2.2.15rc33177633022-py3-none-any.whl -
Subject digest:
2560eacfeadc7d31d268f377d34e1134a8665c3fc3374ce0c91bd7b3a0f6f54a - Sigstore transparency entry: 707092322
- Sigstore integration time:
-
Permalink:
abhishektiwari/hbat@a9676fe7c2ac383bf351773c2d6014cadbd7a607 -
Branch / Tag:
refs/pull/33/merge - Owner: https://github.com/abhishektiwari
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yml@a9676fe7c2ac383bf351773c2d6014cadbd7a607 -
Trigger Event:
pull_request
-
Statement type: