A professional evolutionary discovery tool for Humanin-like peptides (sORFs) using a Hybrid AI approach.
Project description
HumaninFinder v1.0.9 🧬🤖

HumaninFinder is a professional, high-performance Python framework designed for the discovery and classification of Humanin-like peptides (sORFs) within mitochondrial genomes. It employs a Hybrid AI Engine that integrates deep structural embeddings from the ESM-2 Protein Language Model with explicit biophysical analysis to identify functional, non-canonical, and pseudogenic sequences across any taxonomic group.
📂 Repository Structure
.
├── conda/ # Bioconda recipe and metadata
├── deploy/ # Containerization (Dockerfile, Singularity.def)
├── docs/ # Technical documentation and user guides
├── examples/ # Quick-start samples (FASTA genomes)
├── galaxy/ # Galaxy Tool wrapper and integration
├── paper/ # Publication manuscripts
│ ├── joss/ # Software description for JOSS
│ └── primate_study/ # Scientific case study on 61 primate genomes
├── src/
│ └── humaninfinder/ # Main Python Package
│ ├── cli.py # Subcommand-based Command-line interface
│ ├── core.py # Locus localization and ORF finding logic
│ ├── classifier.py # Hybrid AI Engine (ESM-2 + Biophysical)
│ ├── agent.py # AI Research Agent (Ollama integration)
│ ├── data/ # HMM models and 16S probes
│ └── models/ # Pre-trained hybrid classifier weights
├── tests/ # Unit and biological validation tests
├── pyproject.toml # Build system and PyPI definitions
└── pixi.toml # Modern environment management
🧩 Core Components Detail
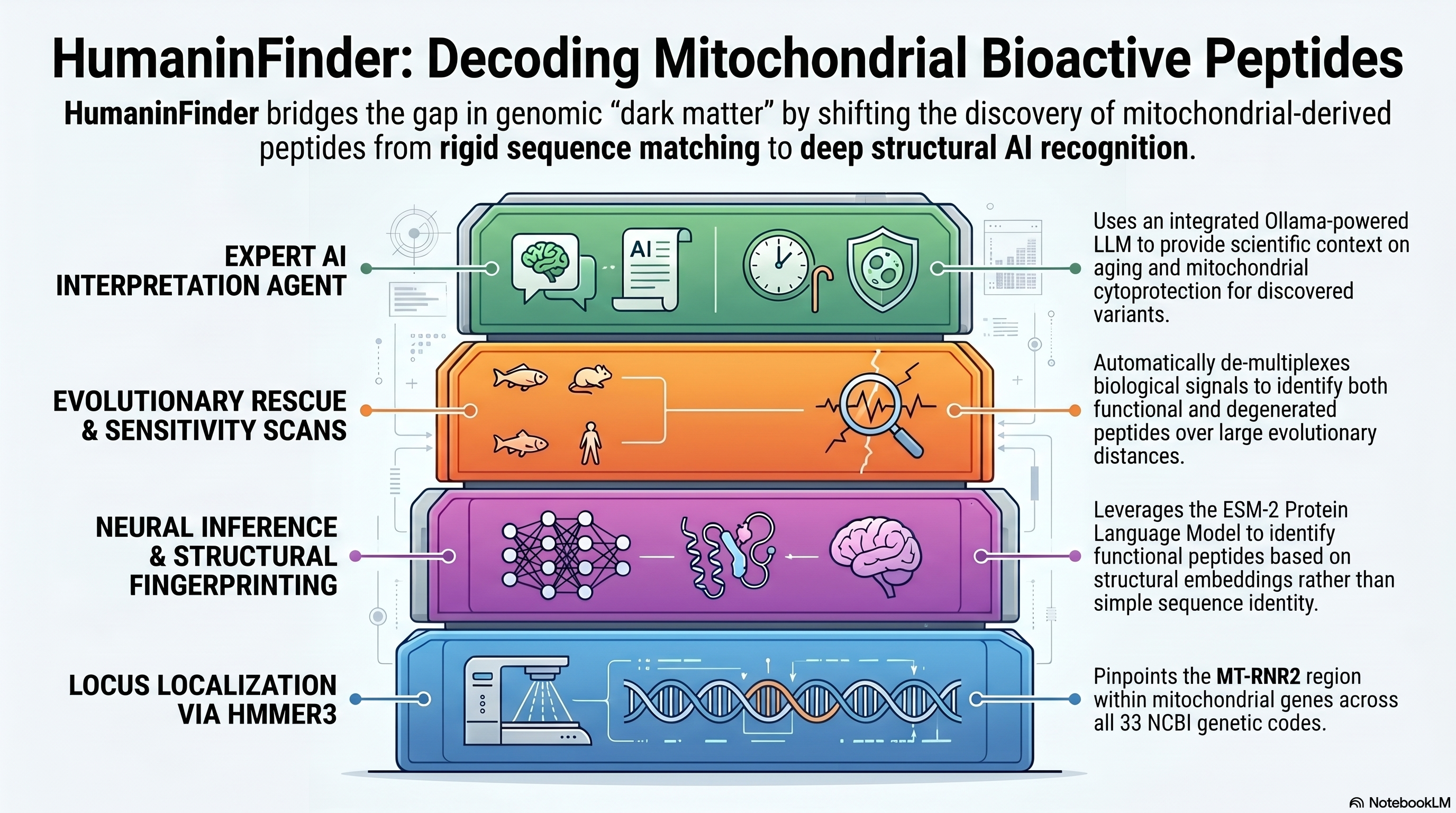
Hybrid AI Engine: Combines mean-pooled embeddings from the ESM-2 transformer model (esm2_t6_8M_UR50D) with charge, pI, and hydrophobicity metrics.Evolutionary Rescue: A high-sensitivity sliding-window scanner that "rescues" non-canonical and pseudogenic relics in diverged lineages.AI Research Agent: An integrated specialist assistant powered by local LLMs (via Ollama) to provide biological interpretation of results in the context of mitochondrial aging and cytoprotection.Biological Deduplication: A specialized filter that ensures independent evolutionary signals by removing technical windowing artifacts.
🚀 Key Features
- Organism Agnostic: Supports all 33 NCBI genetic codes, enabling MDP discovery in any mitochondria-bearing taxon.
- Expert AI Agent: Built-in specialist in Humanin, mitochondrial signaling, and aging biology to interpret your findings.
- High-Throughput Ready: Parallelized processing and optimized inference for large genomic collections.
- Validated Science: Built-in reproduction of the 61-primate evolutionary case study.
🛠️ Quick Start
1. Installation
Option A: via pip (Fastest)
pip install "humaninfinder[agent]"
humanin-finder setup
Note: Ensure HMMER3 is installed on your system.
Option B: via Conda / Mamba (Recommended)
Perfect for an isolated scientific environment:
# Create environment from the provided file
mamba env create -f environment.yml
mamba activate humanin_env
# Finalize setup
humanin-finder setup
2. Run Discovery Pipeline
humanin-finder predict -i genome.fasta -o results --hmm --rescue
3. Biological Interpretation
# Get a summary of your findings from the AI Specialist
humanin-finder agent --results results_results.csv
📖 Documentation
Detailed information is available in the paper/ directory:
- 🛠️ Software Paper: Architecture and methodology for JOSS.
- 🏗️ Scientific Report: Evolutionary dynamics of Humanin in Primates.
🤝 Contributing
Contributions are welcome! Please see our CONTRIBUTING.md for details.
📄 License
This project is licensed under the MIT License - see the LICENSE file for details.
Developed by LaBiOmicS - Laboratory of Bioinformatics and Omics Sciences. Institution: Universidade de Mogi das Cruzes (UMC)
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file humaninfinder-1.0.9.tar.gz.
File metadata
- Download URL: humaninfinder-1.0.9.tar.gz
- Upload date:
- Size: 175.4 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7c199f5fc302e7d4940666c8d42d35449b432eac8121324466f21b574cbe12c7
|
|
| MD5 |
03a381360abbac011768bb6f7c38fc53
|
|
| BLAKE2b-256 |
e432178ea59aa888ad5992959f2730dc82dc6833305fb1447f004736756c0a96
|
File details
Details for the file humaninfinder-1.0.9-py3-none-any.whl.
File metadata
- Download URL: humaninfinder-1.0.9-py3-none-any.whl
- Upload date:
- Size: 171.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
05a2cb71fa24053380917f7133c2fa273a4c1a1881df3352fafeafb91fbd63e8
|
|
| MD5 |
9749ec267a1bed1ef743edc6cd3b65ee
|
|
| BLAKE2b-256 |
1c7118ce9eb152410413ce608d247e25fe983e544771b2bb957453fa94f298a0
|
























