XGBoost for label-imbalanced data: XGBoost with weighted and focal loss functions
Project description
Imbalance-Xgboost
This software includes the codes of Weighted Loss and Focal Loss [1] implementations for Xgboost [2](<\url> https://github.com/dmlc/xgboost) in binary classification problems. The principal reason for us to use Weighted and Focal Loss functions is to address the problem of label-imbalanced data. The original Xgboost program provides a convinient way to customize the loss function, but one will be needing to compute the first and second order derivatives to implement them. The major contribution of the software is the drivation of the gradients and the implementations of them.
Software Update
Version 0.8.1: The package now supports early stopping, you can specify this by early_stopping_rounds when initializing the object.
Version Notification
From version 0.7.0 on Imbalance-XGBoost starts to support higher versions of XGBoost and removes supports of versions earlier than 0.4a30(XGBoost>=0.4a30). This contradicts with the previous requirement of XGBoost<=0.4a30. Please choose the version that fits your system accordingly.
Starting from version 0.8.1, the package now requires xgboost to have a newer version of >=1.1.1. This is due to some changes on deprecated arguments of the XGBoost.
Installation
Installing with Pypi is the easiest way, you can run:
pip install imbalance-xgboost
If you have multiple versions of Python, make sure you're using Python 3 (run with pip3 install imbalance-xgboost). The program are designated for Python 3.5 and 3.6. That being said, an (incomplete) test does not find any compatible issue on Python 3.7 and 3.8.
The package has hard depedency on numpy, sklearn and xgboost.
Usage
To use the wrapper, one needs to import imbalance_xgboost from module imxgboost.imbalance_xgb. An example is given as bellow:
from imxgboost.imbalance_xgb import imbalance_xgboost as imb_xgb
The specific loss function could be set through special_objective parameter. Specificly, one could construct a booster with:
xgboster = imb_xgb(special_objective='focal')
for focal loss and
xgboster = imb_xgb(special_objective='weighted')
for weighted loss. The prarameters $\alpha$ and $\gamma$ can be specified by giving a value when constructing the object. In addition, the class is designed to be compatible with scikit-learn package, and you can treat it as a sk-learn classifier object. Thus, it will be easy to use methods in Sklearn such as GridsearchCV to perform grid search for the parameters of focal and weighted loss functions.
from sklearn.model_selection import GridSearchCV
xgboster_focal = imb_xgb(special_objective='focal')
xgboster_weight = imb_xgb(special_objective='weighted')
CV_focal_booster = GridSearchCV(xgboster_focal, {"focal_gamma":[1.0,1.5,2.0,2.5,3.0]})
CV_weight_booster = GridSearchCV(xgboster_weight, {"imbalance_alpha":[1.5,2.0,2.5,3.0,4.0]})
The data fed to the booster should be of numpy type and following the convention of:
x: [nData, nDim]
y: [nData,]
In other words, the x_input should be row-major and labels should be flat.
And finally, one could fit the data with Cross-validation and retreive the optimal model:
CV_focal_booster.fit(records, labels)
CV_weight_booster.fit(records, labels)
opt_focal_booster = CV_focal_booster.best_estimator_
opt_weight_booster = CV_weight_booster.best_estimator_
After getting the optimal booster, one will be able to make predictions. There are following methods to make predictions with imabalnce-xgboost:
Method predict
raw_output = opt_focal_booster.predict(data_x, y=None)
This will return the value of 'zi' before applying sigmoid.
Method predict_sigmoid
sigmoid_output = opt_focal_booster.predict_sigmoid(data_x, y=None)
This will return the \hat{y} value, which is p(y=1|x) for 2-lcass classification.
Method predict_determine
class_output = opt_focal_booster.predict_determine(data_x, y=None)
This will return the predicted logit, which 0 or 1 in the 2-class scenario.
Method predict_two_class
prob_output = opt_focal_booster.predict_two_class(data_x, y=None)
This will return the predicted probability of 2 classes, in the form of [nData * 2]. The first column is the probability of classifying the datapoint to 0 and the second column is the prob of classifying as 1.
To assistant the evluation of classification results, the package provides a score function score_eval_func() with multiple metrics. One can use make_scorer() method in sk-learn and functools to specify the evaluation score. The method will be compatible with sk-learn cross validation and model selection processes.
import functools
from sklearn.metrics import make_scorer
from sklearn.model_selection import LeaveOneOut, cross_validate
# retrieve the best parameters
xgboost_opt_param = CV_focal_booster.best_params_
# instantialize an imbalance-xgboost instance
xgboost_opt = imb_xgb(special_objective='focal', **xgboost_opt_param)
# cross-validation
# initialize the splitter
loo_splitter = LeaveOneOut()
# initialize the score evalutation function by feeding the 'mode' argument
# 'mode' can be [\'accuracy\', \'precision\',\'recall\',\'f1\',\'MCC\']
score_eval_func = functools.partial(xgboost_opt.score_eval_func, mode='accuracy')
# Leave-One cross validation
loo_info_dict = cross_validate(xgboost_opt, X=x, y=y, cv=loo_splitter, scoring=make_scorer(score_eval_func))
In the new version, we can also collect the information of the confusion matrix through the correct_eval_func provided. This enables the users to evluate the metrics like accuracy, precision, and recall for the average/overall test sets in the cross-validation process.
# initialize the correctness evalutation function by feeding the 'mode' argument
# 'mode' can be ['TP', 'TN', 'FP', 'FN']
TP_eval_func = functools.partial(xgboost_opt.score_eval_func, mode='TP')
TN_eval_func = functools.partial(xgboost_opt.score_eval_func, mode='FP')
FP_eval_func = functools.partial(xgboost_opt.score_eval_func, mode='TN')
FN_eval_func = functools.partial(xgboost_opt.score_eval_func, mode='FN')
# define the score function dictionary
score_dict = {'TP': make_scorer(TP_eval_func),
'FP': make_scorer(TN_eval_func),
'TN': make_scorer(FP_eval_func),
'FN': make_scorer(FN_eval_func)}
# Leave-One cross validation
loo_info_dict = cross_validate(xgboost_opt, X=x, y=y, cv=loo_splitter, scoring=score_dict)
overall_tp = np.sum(loo_info_dict['test_TP']).astype('float')
More soring function may be added in later versions.
Theories and derivatives
You don't have to understand the equations if you find they are hard to grasp, you can simply use it with the API. However, for the purpose of understanding, the derivatives of the two loss functions are listed.
For both of the loss functions, since the task is 2-class classification, the activation would be sigmoid:
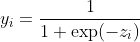
And bellow the two types of loss will be discussed respectively.
1. Weighted Imbalance (Cross-entropoy) Loss
And combining with $\hat{y}$, which are the true labels, the weighted imbalance loss for 2-class data could be denoted as:
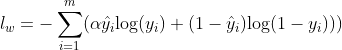
Where $\alpha$ is the 'imbalance factor'. And $\alpha$ value greater than 1 means to put extra loss on 'classifying 1 as 0'.
The gradient would be:
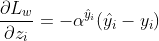
And the second order gradient would be:
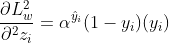
2. Focal Loss
The focal loss is proposed in [1] and the expression of it would be:
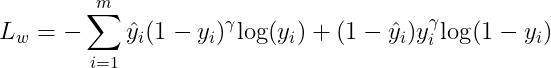
The first order gradient would be:
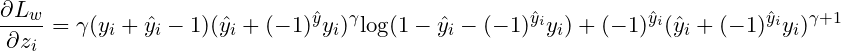
And the second order gradient would be a little bit complex. To simplify the expression, we firstly denotes the terms in the 1-st order gradient as the following notations:
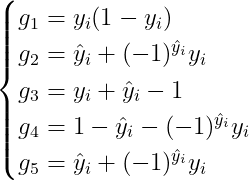
Using the above notations, the 1-st order drivative will be:
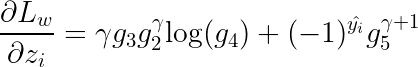
Then the 2-nd order derivative will be:
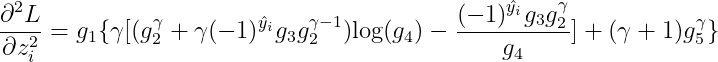
Paper Citation
If you use this package in your research please cite our paper:
@misc{wang2019imbalancexgboost,
title={Imbalance-XGBoost: Leveraging Weighted and Focal Losses for Binary Label-Imbalanced Classification with XGBoost},
author={Chen Wang and Chengyuan Deng and Suzhen Wang},
year={2019},
eprint={1908.01672},
archivePrefix={arXiv},
primaryClass={cs.LG}
}
More information on the software:
@author: Chen Wang, Dept. of Computer Science, School of Art and Science, Rutgers University (previously affiliated with University College London, Sichuan University and Northwestern Polytechnical University)
@version: 0.7.4
References
[1] Lin, Tsung-Yi, Priyal Goyal, Ross Girshick, Kaiming He, and Piotr Dollár. "Focal loss for dense object detection." IEEE transactions on pattern analysis and machine intelligence (2018).
[2] Chen, Tianqi, and Carlos Guestrin. "Xgboost: A scalable tree boosting system." In Proceedings of the 22nd acm sigkdd international conference on knowledge discovery and data mining, pp. 785-794. ACM, 2016.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file imbalance-xgboost-0.8.1.tar.gz.
File metadata
- Download URL: imbalance-xgboost-0.8.1.tar.gz
- Upload date:
- Size: 12.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.3.0 pkginfo/1.7.0 requests/2.25.1 setuptools/41.0.1 requests-toolbelt/0.9.1 tqdm/4.56.0 CPython/3.7.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
fa4fd1f8f9689ba0dd3e3032d24f8e7dba82af3b70e3ede04f9b47a4a06315e6
|
|
| MD5 |
2b21b31dc2f9a88c02e87960bd0cfa23
|
|
| BLAKE2b-256 |
35767772df6ab39e66d66bcf908ea2b2623961e0c8c6f4a45c94d6e6728ab786
|
File details
Details for the file imbalance_xgboost-0.8.1-py3-none-any.whl.
File metadata
- Download URL: imbalance_xgboost-0.8.1-py3-none-any.whl
- Upload date:
- Size: 15.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.3.0 pkginfo/1.7.0 requests/2.25.1 setuptools/41.0.1 requests-toolbelt/0.9.1 tqdm/4.56.0 CPython/3.7.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8a728a81f5cd7b851f686d8e605289ce2f19d1846ebc8c57d14b390ccdd389ce
|
|
| MD5 |
0ad250bf856ff2e515112b3d899febc3
|
|
| BLAKE2b-256 |
cb7e1ce879a4d3c1f8a19cc2e98f907ebefaf17808b5dbd106155ea52e7c3ffd
|











