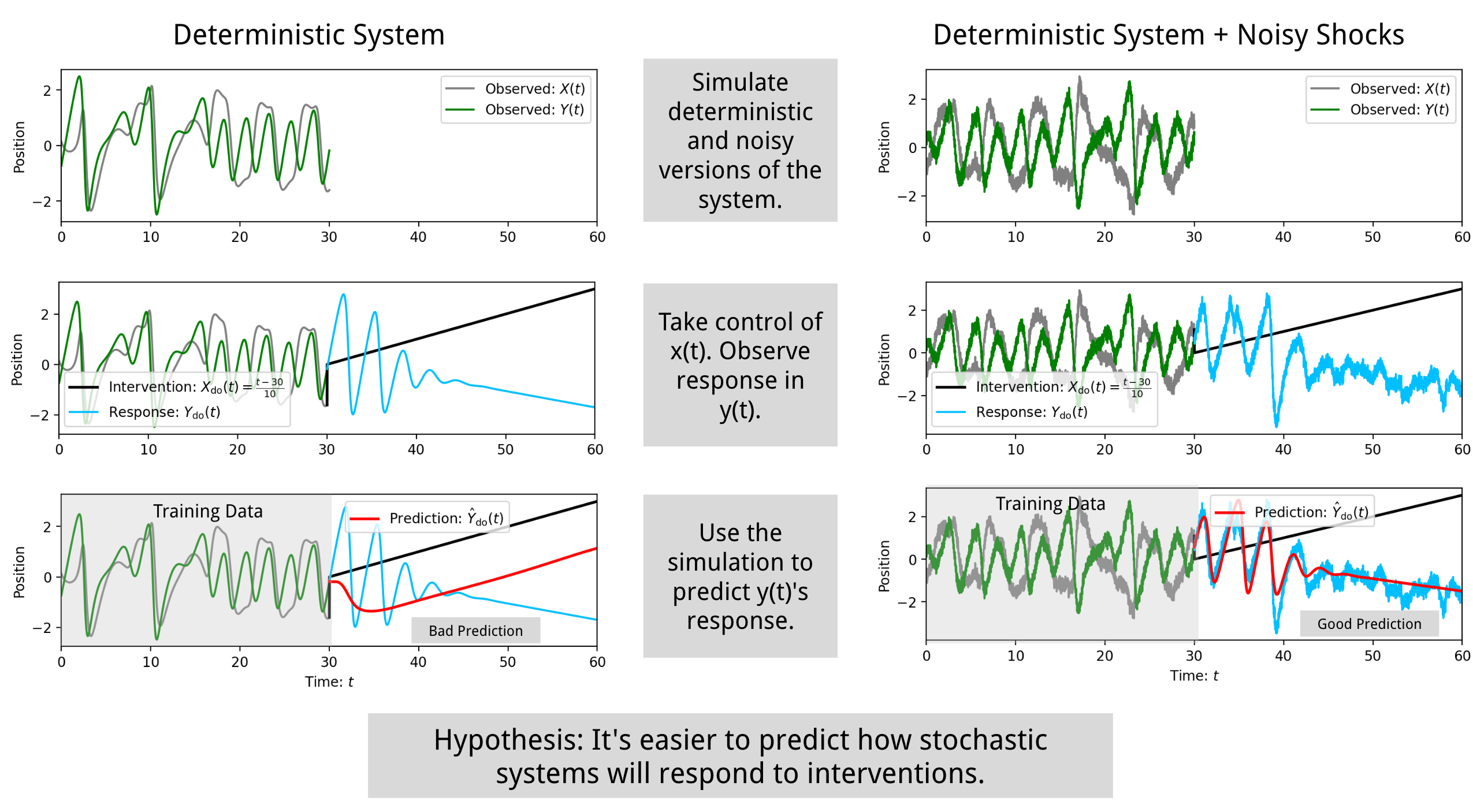
Predicting how complex systems will respond to interventions.
Project description
🌀 Interfere
A Python package for modeling and predicting the response of complex dynamic systems to interventions.
Overview
Interfere is a research-oriented Python package that addresses a fundamental question in complex systems: When can we predict how complex systems will respond to interventions? This package provides tools for:
- Modeling dynamic nonlinear multivariate stochastic systems.
- Simulating and analyzing how such systems respond to interventions.
- Generating complex dynamic counterfactuals.
- Studying causal relationships in complex systems.
Interfere Benchmark Dataset (Download)
The image above depicts the uninterrupted trajectories of sixty dynamic models in blue and their response to a particular intervention in red. This data is available for download as the Interfere Benchmark 1.1.1. It can be used to benchmark a forecasting method's ability to predict the response of a dynamic system to interventions.
Documentation
Full documentation is built with MkDocs and published at https://djpasseyjr.github.io/interfere/.
Installation
From pip
pip install interfere
From Local Clone
git clone https://github.com/djpasseyjr/interfere.git
cd interfere
pip install .
Quick Start
The Interfere package is designed around three main tasks: counterfactual simulation, predictive method optimization, and prediction. Here's a complete example using the SINDy (Sparse Identification of Nonlinear Dynamics) method:
1. Counterfactual Simulation
First, let's create and simulate a dynamic model:
import numpy as np
import interfere
import optuna
# Set up simulation parameters
initial_cond = np.random.rand(3)
t_train = np.arange(0, 10, 0.05)
dynamics = interfere.dynamics.Belozyorov3DQuad(sigma=0.5)
# Generate trajectory
sim_states = dynamics.simulate(t_train, initial_cond)
2. Applying an Intervention
Next, we'll apply an intervention to one component of the system:
# Time points for the intervention simulation
test_t = np.arange(t_train[-1], 15, 0.05)
# Intervention initialization
intervention = interfere.SignalIntervention(iv_idxs=1, signals=np.sin)
# Simulate intervention
interv_states = dynamics.simulate(
test_t,
prior_states=sim_states,
intervention=intervention,
)
3. Model Optimization and Prediction
Using the generated data, we can run hyperparameter optimization with a forecasting method. All forecasting methods come with reasonable hyperparameter ranges built in.
# Select the SINDy method for hyperparameter optimization.
method_type = interfere.SINDy
# Create an objective function that aims to minimize cross validation error
# over different hyper parameter configurations for SINDy
cv_obj = interfere.CrossValObjective(
method_type=method_type,
data=sim_states,
times=t_train,
train_window_percent=0.3,
num_folds=5,
exog_idxs=intervention.iv_idxs,
)
# Run the study using optuna.
study = optuna.create_study()
study.optimize(cv_obj, n_trials=25)
# Collect the best hyperparameters into a dictionary.
best_param_dict = study.best_params
4. Intervention Response Prediction
Using the best parameters found, we can fit the forecasting method to pre-intervention data and then make a prediction about how the system will respond to the intervention.
# Initialize SINDy with the best perfoming parameters.
method = interfere.SINDy(**study.best_params)
# Use an intervention helper function to split the pre-intervention data
# into endogenous and exogenous columns.
Y_endog, Y_exog = intervention.split_exog(sim_states)
# Fit SINDy to the pre-intervention data.
method.fit(t_train, Y_endog, Y_exog)
# Use the inherited interfere.ForecastMethod.simulate() method
# To simulate intervention response using SINDy
pred_traj = method.simulate(
test_t, prior_states=sim_states, intervention=intervention
)
The SINDy method identifies the underlying dynamics of the system using sparse regression techniques, making it particularly effective for discovering interpretable mathematical models of complex systems.
Dependencies
- Basic:
pip install interfere - Full forecasting methods:
pip install "interfere[methods]" - Developer / experimental features:
pip install "interfere[dev]"
Example Use
The package can be used to simulate and analyze how systems respond to interventions. For example, it can model the effect of stochasticity on intervention response forecasting:
Documentation
For a more detailed explanation of the purpose of the package refer to paper.pdf.
Contributing
Contributions are welcome! To contribute code, make your own local fork of the repository.
Then install the full developer deps using pip install ".[dev]". (The full dependencies are pretty big. Use a
virtual environment, so you can delete it when you are done.)
After you write code, auto-format it with black at the top level of the repo:
$ black interfere
Then you can run the linter and fix any linter errors (also at the top level):
$ flake8 interfere
Build docs
Docs use MkDocs (mkdocs.yml). From the repo root:
$ mkdocs serve # preview at http://127.0.0.1:8000
$ mkdocs build # output in site/
Add Tests
If you are adding a dynamic model or forecasting method, the test suite has a series of prebuilt tests.
Dynamic Model Tests
Add a factory function in tests/sample_models.py, then import it and append your model instance to the MODELS list in tests/test_dynamics.py:
# In sample_models.py
def my_model() -> interfere.dynamics.MyDynamics:
return interfere.dynamics.MyDynamics(...)
# In test_dynamics.py: add to imports, then
MODELS = [
...
my_model()
]
Run tests for a specific model by index:
pytest tests -k "model7"
(The above will only run tests for the 8th model in the MODELS list.)
Forecasting Method Tests
Add your method class to the METHODS list in tests/test_methods.py:
# In test_methods.py
METHODS = [
...
interfere.methods.YourMethod,
]
Run tests for a specific method by name:
pytest tests -k "YourMethod"
Optional dependencies (GitHub forks)
Some features need maintainer forks that are not on PyPI (they cannot be declared in PyPI package metadata). Install them for local development and before running the full test suite.
SURD (surd) — SURD_SINDy
- Module:
interfere/_methods/restricted_sindy.py(exported viainterfere.methods) - Uses
surd.surd_parallelto restrict SINDy terms by information decomposition - Also needs Tigramite:
pip install interfere[methods]
pip install git+https://github.com/djpasseyjr/surd.git
pyclustering fork — oscillatory neural dynamics
- Module:
interfere/dynamics/pyclustering_models.py(not imported by default) - Models:
HodgkinHuxleyPyclustering,LEGIONPyclustering,StuartLandauKuramoto
pip install pyclustering@git+https://github.com/djpasseyjr/pyclustering
Running Full Tests
The full test suite takes over an hour. When contributing, just make sure you add tests for new code. The full suite will run as part of the automated checks with your pull request.
If you like, you can install the full set of dependencies and run tests locally like this:
git clone https://github.com/djpasseyjr/interfere.git
cd interfere
pip install pyclustering@git+https://github.com/djpasseyjr/pyclustering
pip install git+https://github.com/djpasseyjr/surd.git
pip install ".[dev]"
python -m pytest -v tests
(The full dependencies are pretty big. Use a virtual environment.)
Building the docs
Install dev dependencies (which include mkdocs), then build and serve the site locally:
pip install ".[dev]"
mkdocs serve
Open http://127.0.0.1:8000 to view the docs. To build static files only: mkdocs build.
License
This project is licensed under the MIT License - see the LICENSE file for details.
Contact
- Author: DJ Passey (djpassey@unc.edu)
- Institution: University of North Carolina at Chapel Hill
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file interfere-1.0.2.tar.gz.
File metadata
- Download URL: interfere-1.0.2.tar.gz
- Upload date:
- Size: 7.7 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.9.6
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1aeac3612accab8ce355559fc8a1a39cdedf56a4f7979384843dab8620f87b2d
|
|
| MD5 |
2ce4e581d99cd39ba58d558efcada25c
|
|
| BLAKE2b-256 |
fb53ce47f6b0cba155588d104f526616509b7c8360babeaefac6ea3efc1957c2
|
File details
Details for the file interfere-1.0.2-py3-none-any.whl.
File metadata
- Download URL: interfere-1.0.2-py3-none-any.whl
- Upload date:
- Size: 7.7 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.9.6
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
6e4082005f70fc421bdc10f44256b7ac86a9153a3dacfbb0335a1c2ca8aebb63
|
|
| MD5 |
e43b7a7d6bbb0afd66dc3ec62a997c39
|
|
| BLAKE2b-256 |
b30003d06aae450ba00f7cb942c76bf951bde6546733288eb7197b1792c2f2d3
|