No project description provided
Project description


A toolbox to analyze and interact with Ca imaging data, developed within the Cell and Tissue Networks research group led by prof. Marjan Slak-Rupnik at the Medical University of Vienna.
https://user-images.githubusercontent.com/2512087/162633046-b26d7c49-3501-4e78-9a33-433119157537.mp4
A typical experiment involving imaging of pancreatic slices in our lab concerns a single field of view showing up to hundreds of cells, in a recording of at least several, often dozens, gigabytes. Current tools (i.e. ImageJ) rely on loading the recording, or its part, into memory, for viewing, analysis, and processing. It also requires laborious and long human engagement. We have developed a set of interdependent tools to automatize as much as possible the analysis pipeline.
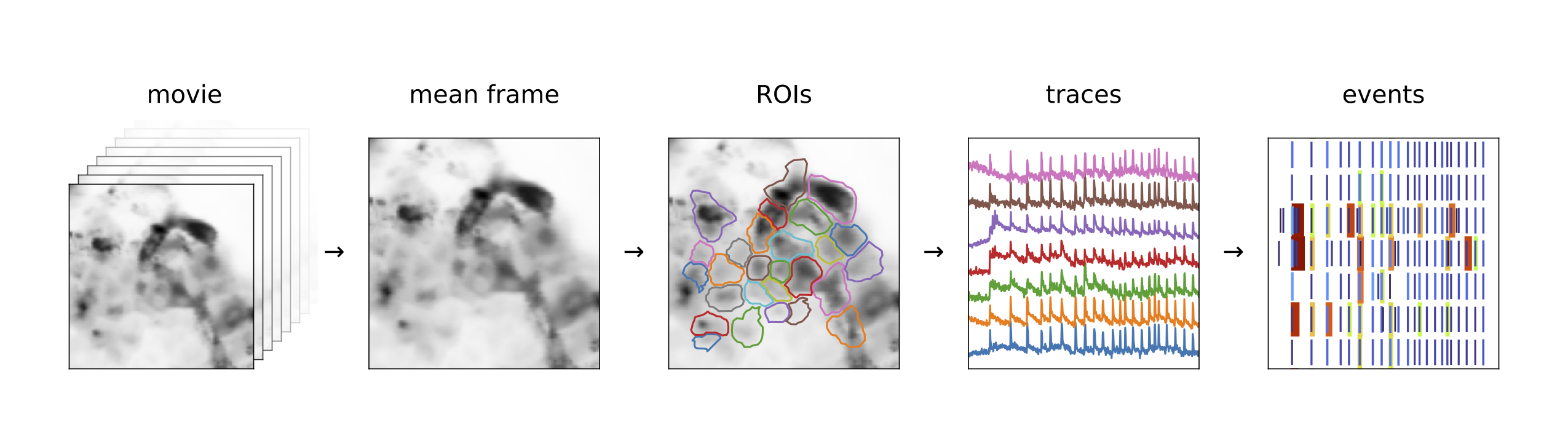
- The main elements of our pipeline are the following:
Automatic detection of regions of interest (ROIs);
Transformation of ROI time traces into standard score (“z-score”) and correction for filtering distortion;
Representation of the phenotype for each trace (height, auc, or halfwidth statistics, event rate…).
Quantification of the effect magnitude for acute pharmacological manipulations, and/or different experimental branches (different mice, or diet, of genetic manipulations).
Documentation
The usage of the framework in practical terms is documented in the original repository at https://github.com/szarma/Physio_Ca/
Features
One of our most used tools is the _’’roi examiner’’_ for live interaction with the ROIs and their traces within a jupyter notebook.
We have a similar app to examine line scans.
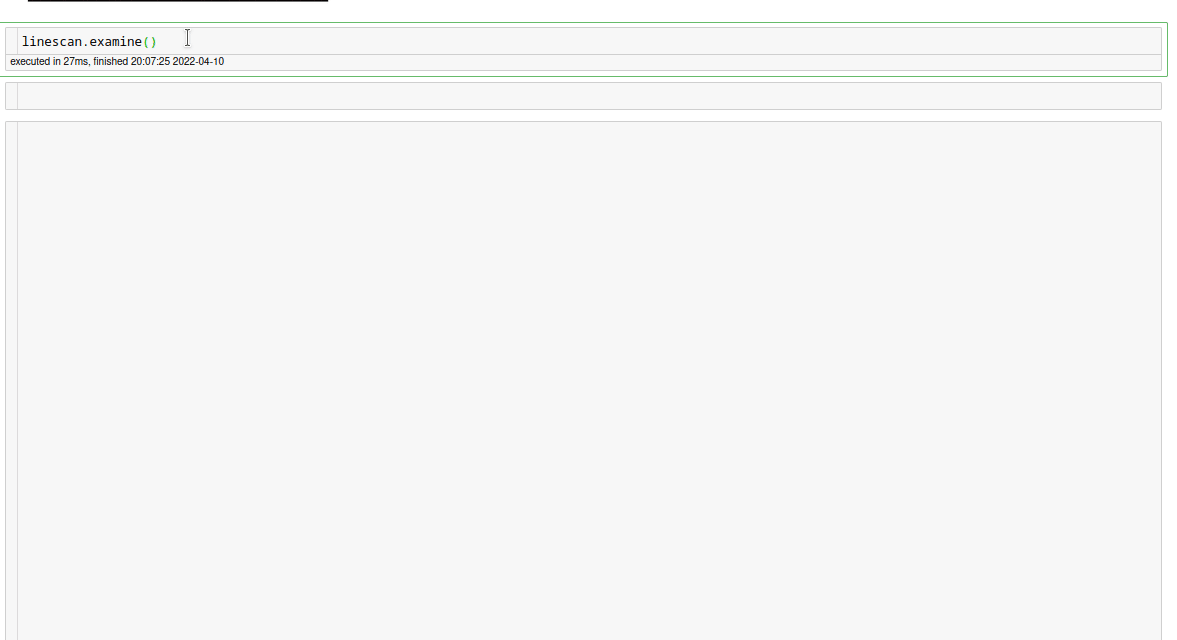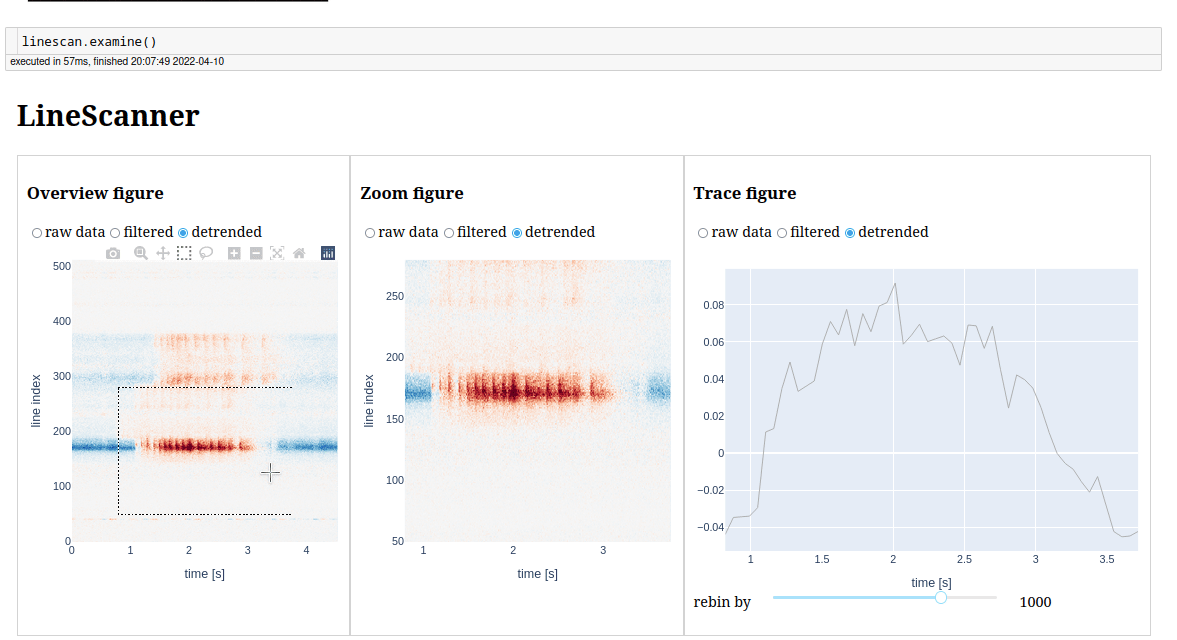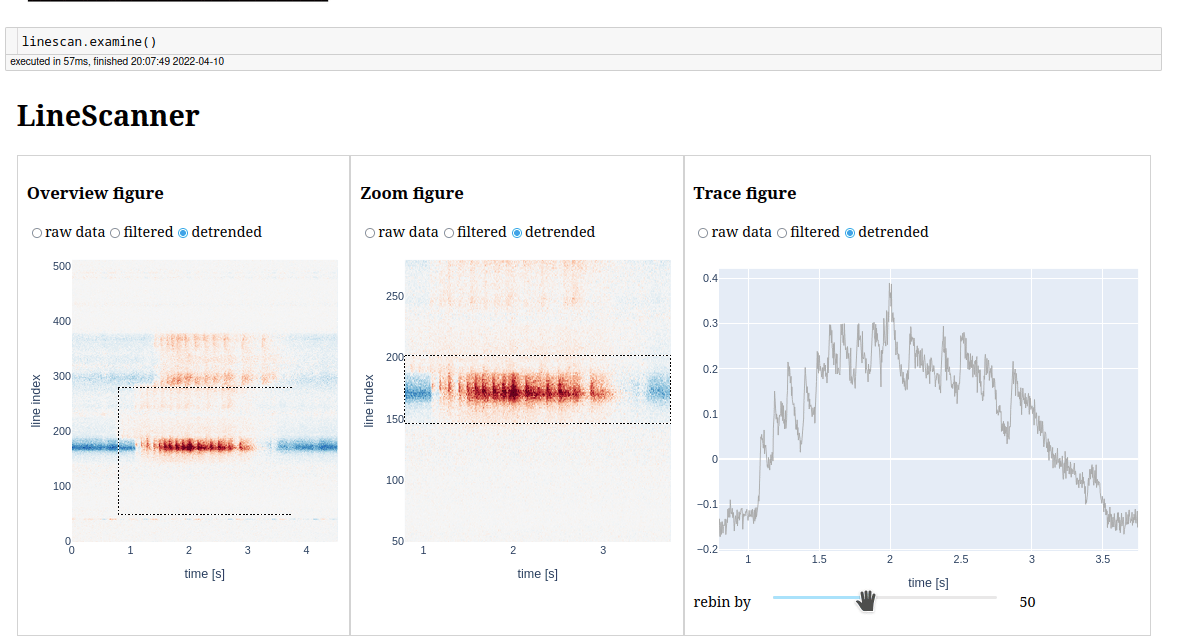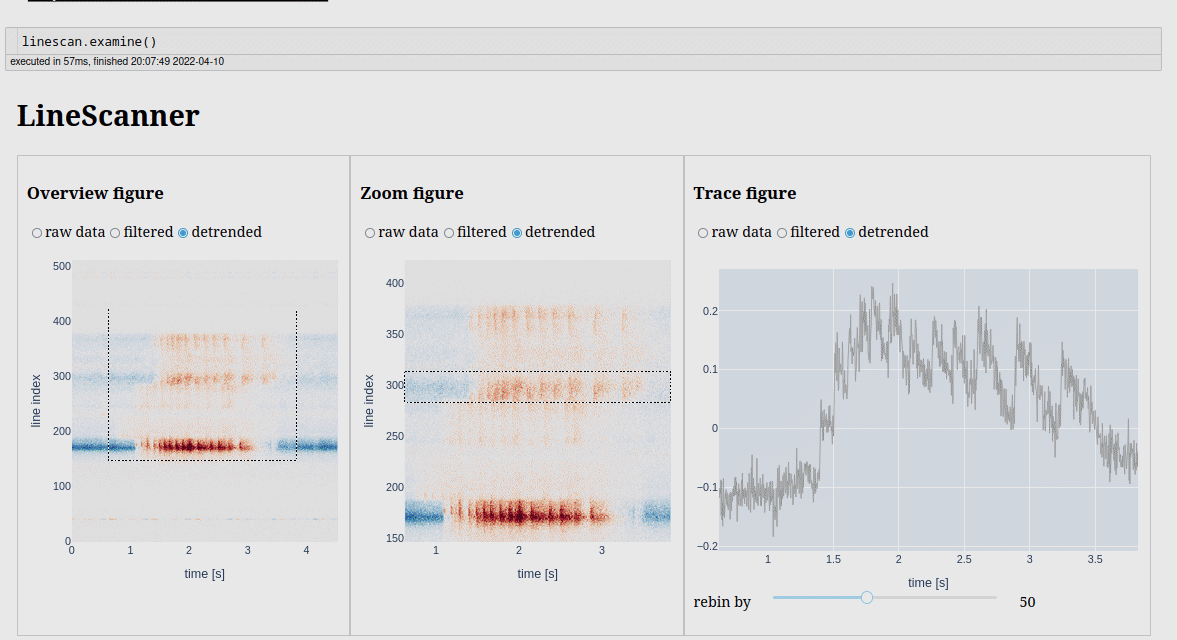
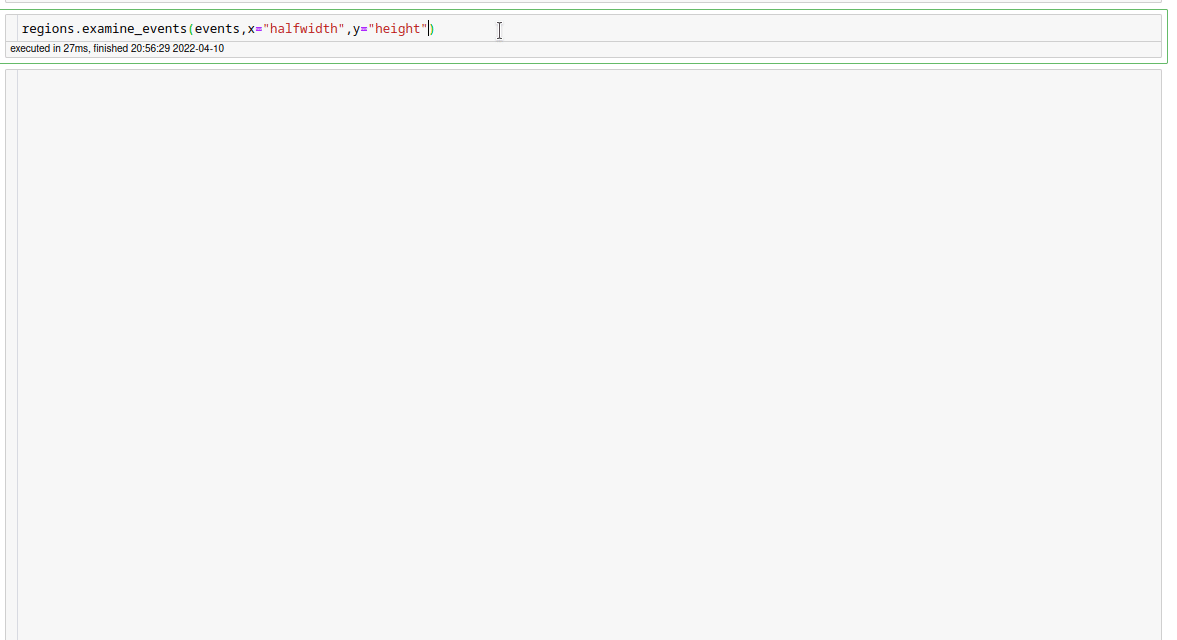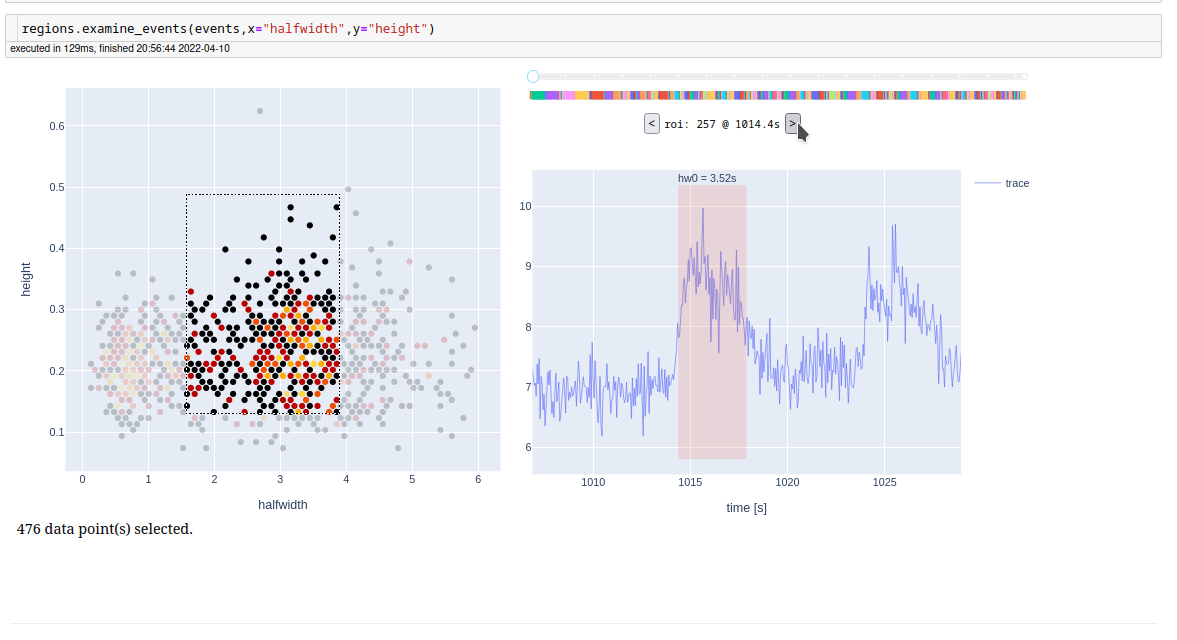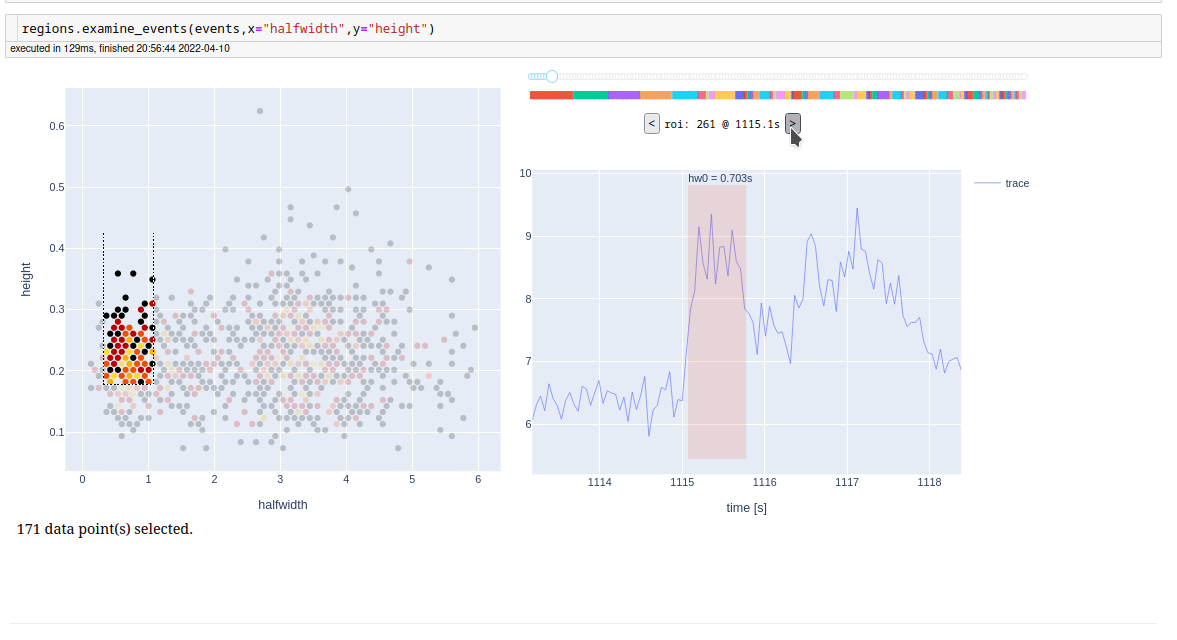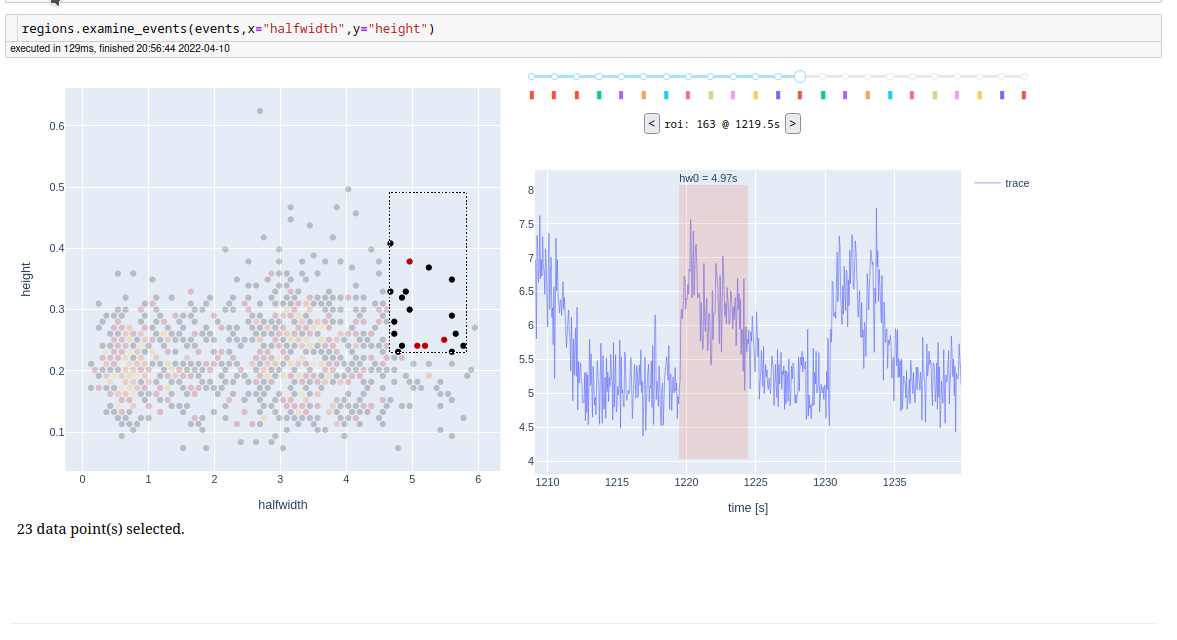
For examine detected events, one at a time, we also have a app.
We have also built a dashboard for fast intreaction with our storage filesystem. Given a folder, it finds all processed recordings in it and its subfolders, collects metadata and presents it in a table form. It further enables entering of the experimental protocol, and additional data, which are then also searchable. It also provides a link to an automaticaly generated notebook for a brief glimpse into actual results of an experiment. See demo on youtube (https://youtu.be/tj4TjL_PJ1Q).
Installation
We are working on an easy way of installation. For this purpose we use poetry as install tool. However, there will be ready-to-go packages available on PyPI via pip. For easy deployment, there will be a docker image as well. Documentation will be updated as soon as it is ready.
Docker
A dockerfile is included in the root of the framework. It contains everything to run python code in the base environment. It can be built with the following command:
docker build -t ctn_server .If you do not want to build it yourself there is a prebuilt version on docker-hub. It can be pulled simly by:
docker pull hannsen/cell-tissue-networks_server:latestAs an example, to run the server with custom data and access it in a shell you can use it like this:
docker run -it -v /path/to/real/data:/data:rw hannsen/cell-tissue-networks_server:latest /bin/bashProject details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file islets-0.7.5.tar.gz.
File metadata
- Download URL: islets-0.7.5.tar.gz
- Upload date:
- Size: 140.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.1.14 CPython/3.10.5 Linux/5.19.1-3-MANJARO
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e138cea5ae23a6db7f4f99dc455b6e41748966d16fc38ba97d3bfe99e46758f1
|
|
| MD5 |
e40f5628aa9695ae84ffe6aa8a5cf73a
|
|
| BLAKE2b-256 |
9e6b6b81bd4a0eb80f9e9d614fe5113fd356af8b47041af79d52a8ef41b66d8a
|
File details
Details for the file islets-0.7.5-py3-none-any.whl.
File metadata
- Download URL: islets-0.7.5-py3-none-any.whl
- Upload date:
- Size: 150.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.1.14 CPython/3.10.5 Linux/5.19.1-3-MANJARO
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cf4bfd715d79b1890ecb48bc9378471785c268a7f8fa4573f01b7692ab9adc9c
|
|
| MD5 |
f3ef5baa635b38bf6729404cd751624c
|
|
| BLAKE2b-256 |
615319aa60c6088b300861ebe41de487341873b6f632fe7e7d4e56cd6bd0579c
|










