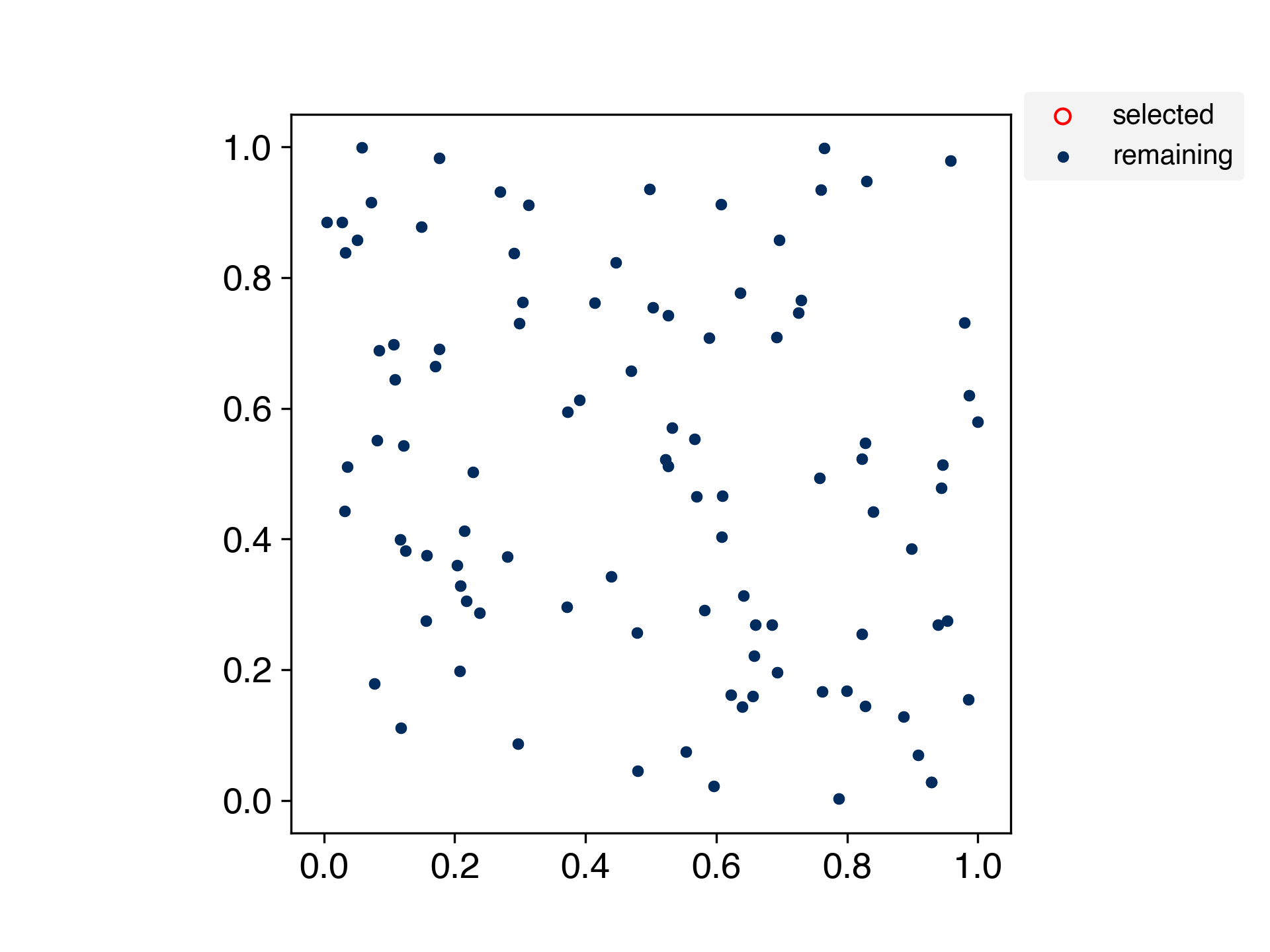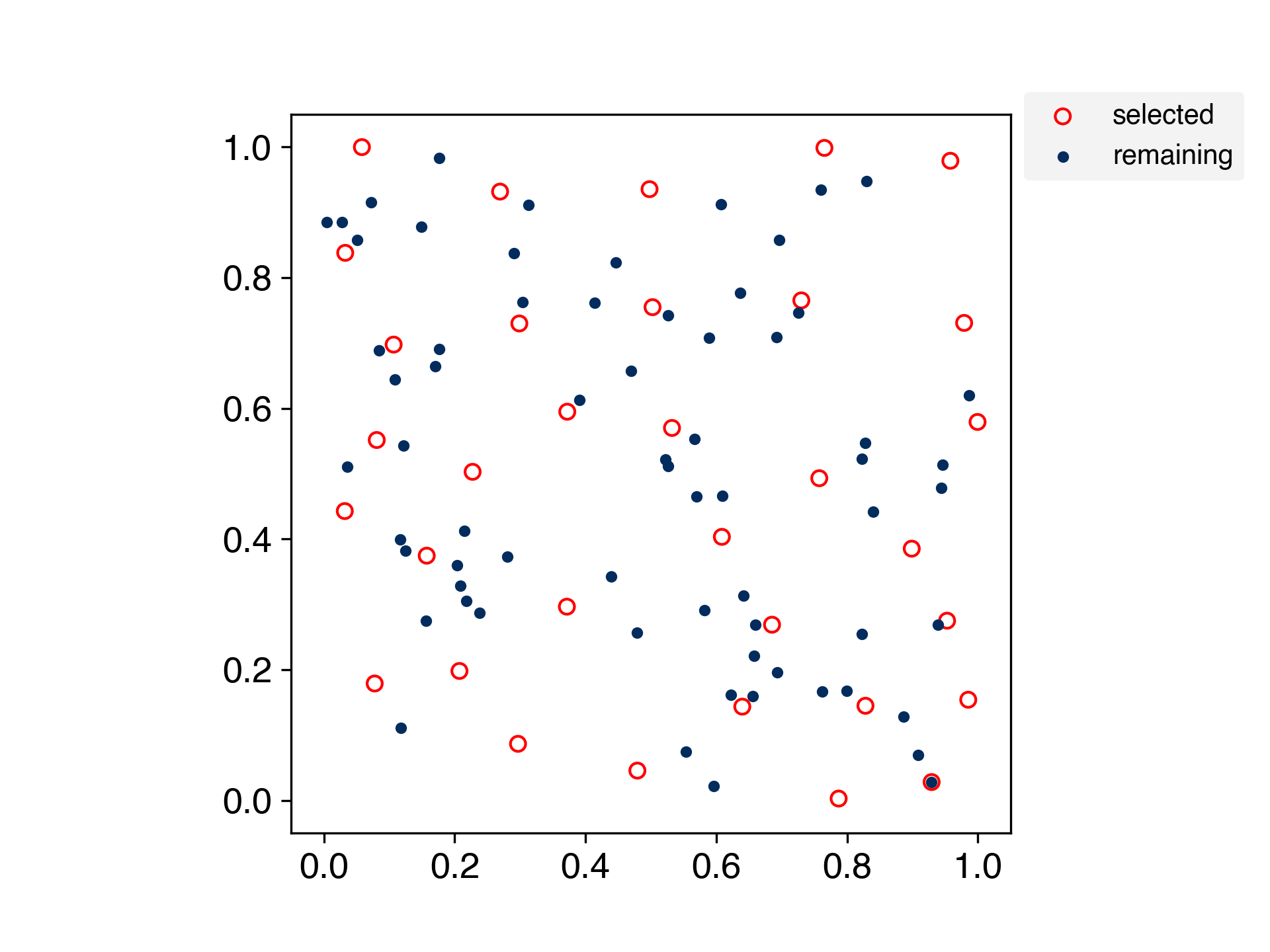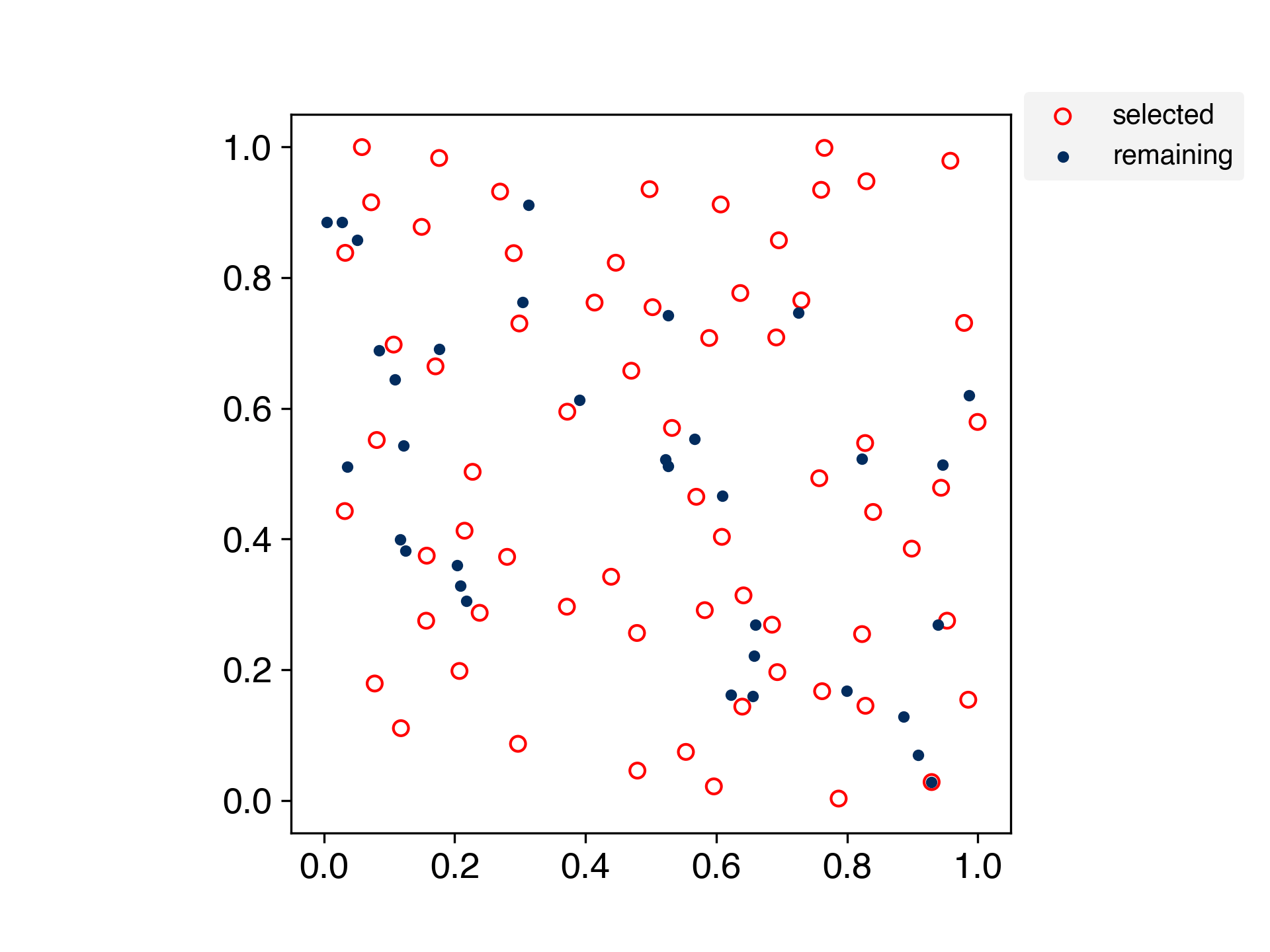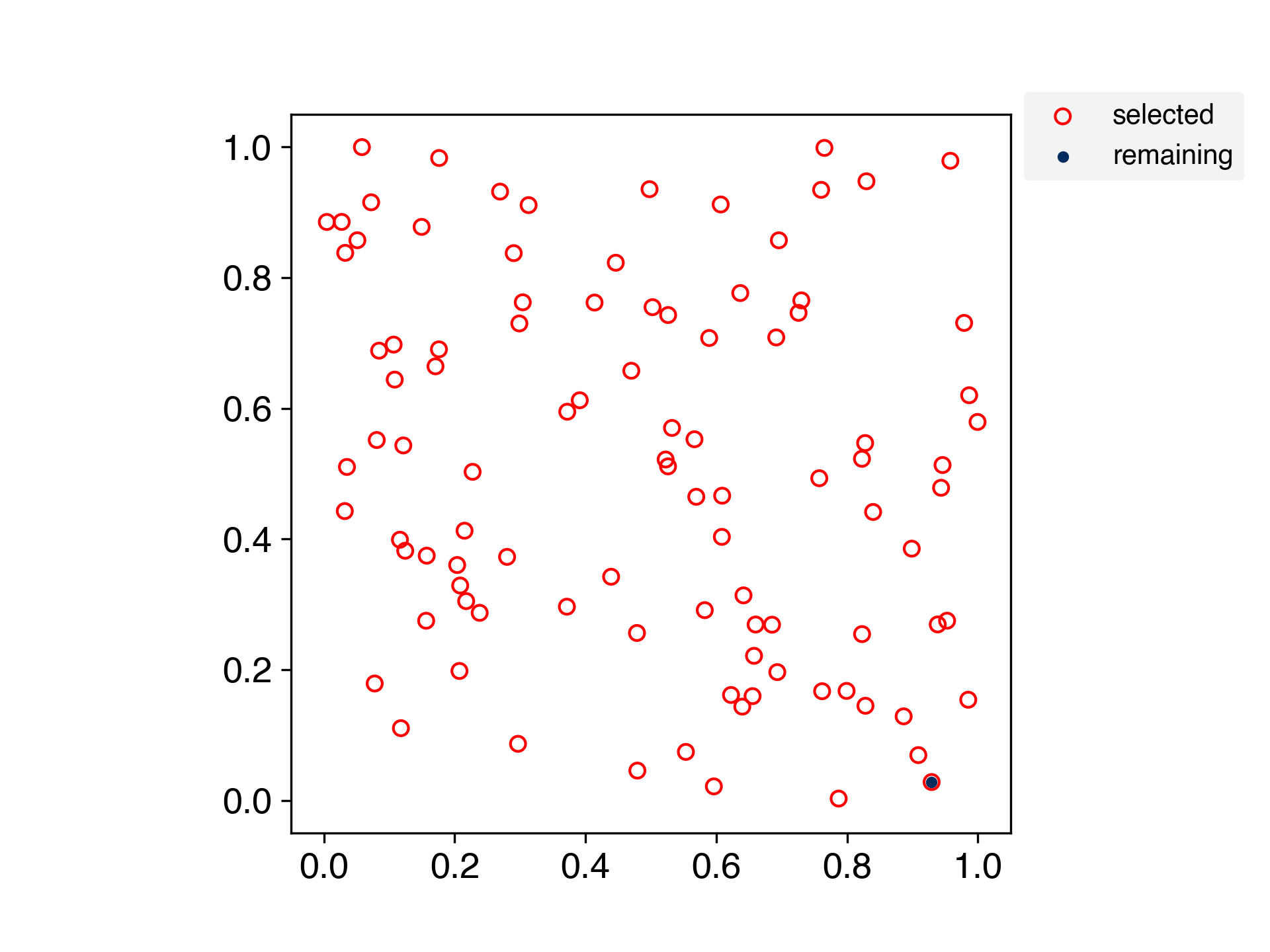
A method for selecting samples by spreading the training data evenly.
Project description
Kennard Stone
What is this?
This is an algorithm for evenly partitioning data in a scikit-learn-like interface.
(See References for details of the algorithm.)
How to install
PyPI
pip install kennard-stone
The project site is here.
Anaconda
conda install -c conda-forge kennard-stone
The project site is here.
You need numpy>=1.20 and scikit-learn to run.
How to use
You can use them like scikit-learn.
See examples for details.
In the following, X denotes an arbitrary explanatory variable and y an arbitrary objective variable.
And, estimator indicates an arbitrary prediction model that conforms to scikit-learn.
train_test_split
kennard_stone
from kennard_stone import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
scikit-learn
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, random_state=334
)
KFold
kennard_stone
from kennard_stone import KFold
# Always shuffled and uniquely determined for a data set.
kf = KFold(n_splits=5)
for i_train, i_test in kf.split(X, y):
X_train = X[i_train]
y_train = y[i_train]
X_test = X[i_test]
y_test = y[i_test]
scikit-learn
from sklearn.model_selection import KFold
kf = KFold(n_splits=5, shuffle=True, random_state=334)
for i_train, i_test in kf.split(X, y):
X_train = X[i_train]
y_train = y[i_train]
X_test = X[i_test]
y_test = y[i_test]
Other usages
If you ever specify cv in scikit-learn, you can assign KFold objects to it and apply it to various functions.
An example is cross_validate.
kennard_stone
from kennard_stone import KFold
from sklearn.model_selection import cross_validate
kf = KFold(n_splits=5)
print(cross_validate(estimator, X, y, cv=kf))
scikit-learn
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_validate
kf = KFold(n_splits=5, shuffle=True, random_state=334)
print(cross_validate(estimator, X, y, cv=kf))
OR
from sklearn.model_selection import cross_validate
print(cross_validate(estimator, X, y, cv=5))
Notes
There is no notion of random_state or shuffle because the partitioning is determined uniquely for the dataset.
If these arguments are included, they do not cause an error. They simply have no effect on the result. Please be careful.
If you want to run the notebook in examples directory,
you will need to additionally install pandas, matplotlib, seaborn, tqdm, and jupyter other than the packages in requirements.txt.
Distance metrics
See the documentation of
scipy.spatial.distance.pdisthttps://docs.scipy.org/doc/scipy/reference/generated/scipy.spatial.distance.pdist.htmlsklearn.metrics.pairwise_distanceshttps://scikit-learn.org/stable/modules/generated/sklearn.metrics.pairwise_distances.html
Valid values for metric are:
- From scikit-learn: ['cityblock', 'cosine', 'euclidean', 'l1', 'l2', 'manhattan']. These metrics support sparse matrix inputs. ['nan_euclidean'] but it does not yet support sparse matrices.
- From scipy.spatial.distance: ['braycurtis', 'canberra', 'chebyshev', 'correlation', 'dice', 'hamming', 'jaccard', 'mahalanobis', 'minkowski', 'rogerstanimoto', 'russellrao', 'seuclidean', 'sokalmichener', 'sokalsneath', 'sqeuclidean', 'yule'] See the documentation for scipy.spatial.distance for details on these metrics. These metrics do not support sparse matrix inputs.
, by default "euclidean"
Parallelization (since v2.1.0)
This algorithm is very computationally intensive and takes a lot of time.
To solve this problem, I have implemented parallelization and optimized the algorithm since v2.1.0.
n_jobs can be specified for parallelization as in the scikit-learn-like api.
# parallelization KFold
kf = KFold(n_splits=5, n_jobs=-1)
# parallelization train_test_split
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, n_jobs=-1
)
The parallelization is used when calculating the distance matrix,
so it doesn't conflict with something like cross_validate in parallel when using KFold.
# OK: does not conflict each other
cross_validate(estimator, X, y, cv=KFold(5, n_jobs=-1), n_jobs=-1)
Using GPU
If you have a GPU and have installed pytorch, you can use it to calculate Minkowski distances (Manhattan, Euclidean, and Chebyshev distances).
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, device="cuda"
)
LICENSE
MIT Licence
Copyright (c) 2021 yu9824
References
Papers
- R. W. Kennard & L. A. Stone (1969) Computer Aided Design of Experiments, Technometrics, 11:1, 137-148, DOI: 10.1080/00401706.1969.10490666
Sites
- https://datachemeng.com/trainingtestdivision/ (Japanese site)
Histories
v2.0.0 (deprecated)
- Define Extended Kennard-Stone algorithm (multi-class) i.e. Improve KFold algorithm.
- Delete
alternateargument inKFold. - Delete requirements of
pandas.
v2.0.1
- Fix bug with Python3.7.
v2.1.0 (deprecated)
- Optimize algorithm
- Deal with Large number of data.
- parallel calculation when calculating distance (Add
n_jobsargument) - replacing recursive functions with for-loops
- parallel calculation when calculating distance (Add
- Add other than "euclidean" calculation methods (Add
metricargument)
v2.1.1 (deprecated)
- Fix bug when
metric="nan_euclidean".
v2.1.2 (deprecated)
- Fix details.
- Update docstrings and typings.
v2.1.3 (deprecated)
- Fix details.
- Update some typings. (You have access to a list of strings that can be used in the metric.)
v2.1.4
- Fix bug when metric=="seuclidean" and "mahalanobis"
- Add some tests to check all metrics.
- Add requirements numpy>=1.20
v2.1.5
- Delete "klusinski" metric to support scipy>=1.11
v2.1.6
- Improve typing in
kennard_stone.train_test_split - Add some docstrings.
v2.2.0
- Supports GPU calculations. (when metric is 'euclidean', 'manhattan', 'chebyshev' and 'minkowski')
- Supports Python 3.12
v2.2.1
- Fix setup.cfg
- Update 'typing'
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file kennard_stone-3.0.1.tar.gz.
File metadata
- Download URL: kennard_stone-3.0.1.tar.gz
- Upload date:
- Size: 19.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9050695546ec0b04dcd667fa2ce9db6f08cabd9124e6b89719e05f211ab30666
|
|
| MD5 |
d42864f4ced6fecebe2a5fb280f045dd
|
|
| BLAKE2b-256 |
20dc6ec59ff435fc09babd75f43ae12974ba8edbaae125e9c35289b5c0fbe2ec
|
Provenance
The following attestation bundles were made for kennard_stone-3.0.1.tar.gz:
Publisher:
release-pypi.yml on yu9824/kennard_stone
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
kennard_stone-3.0.1.tar.gz -
Subject digest:
9050695546ec0b04dcd667fa2ce9db6f08cabd9124e6b89719e05f211ab30666 - Sigstore transparency entry: 444287052
- Sigstore integration time:
-
Permalink:
yu9824/kennard_stone@63ca4397830bebc6594abd5e95fae39cf022ab9b -
Branch / Tag:
refs/tags/v3.0.1 - Owner: https://github.com/yu9824
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release-pypi.yml@63ca4397830bebc6594abd5e95fae39cf022ab9b -
Trigger Event:
push
-
Statement type:
File details
Details for the file kennard_stone-3.0.1-py3-none-any.whl.
File metadata
- Download URL: kennard_stone-3.0.1-py3-none-any.whl
- Upload date:
- Size: 18.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
0823ea78cda671d94679320eb4654a904786c79bf9586cbb76dac9d12d4a3fdb
|
|
| MD5 |
81e01f15eeaea302d32d40ee094dbc43
|
|
| BLAKE2b-256 |
577adc3576523a11ac4c691f530f7676765e9ff9dfb29fcc7a7d014ebdee4c23
|
Provenance
The following attestation bundles were made for kennard_stone-3.0.1-py3-none-any.whl:
Publisher:
release-pypi.yml on yu9824/kennard_stone
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
kennard_stone-3.0.1-py3-none-any.whl -
Subject digest:
0823ea78cda671d94679320eb4654a904786c79bf9586cbb76dac9d12d4a3fdb - Sigstore transparency entry: 444287064
- Sigstore integration time:
-
Permalink:
yu9824/kennard_stone@63ca4397830bebc6594abd5e95fae39cf022ab9b -
Branch / Tag:
refs/tags/v3.0.1 - Owner: https://github.com/yu9824
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release-pypi.yml@63ca4397830bebc6594abd5e95fae39cf022ab9b -
Trigger Event:
push
-
Statement type: