Feature extractor from noisy time series
Project description
light-curve processing toolbox for Python
A Python wrapper for the light-curve-feature and
light-curve-dmdt Rust crates, providing high-performance
time-series feature extraction for astrophysics.
Quick start
pip install 'light-curve[full]'
import light_curve as lc
import numpy as np
rng = np.random.default_rng(0)
n = 100 # observations per light curve
# Observation times in days (unevenly sampled, must be sorted)
t = np.sort(rng.uniform(0, 100, n))
# Magnitudes with a slight linear fade and measurement noise
m = 15.0 + 0.01 * t + rng.normal(0, 0.1, n)
err = np.full(n, 0.1)
# Combine features into a single extractor evaluated in one pass
extractor = lc.Extractor(lc.Amplitude(), lc.BeyondNStd(nstd=1), lc.LinearFit())
result = extractor(t, m, err)
print('\n'.join(f"{name} = {value:.4f}" for name, value in zip(extractor.names, result)))
# Extract a feature from 1000 light curves in parallel
light_curves = [
(np.sort(rng.uniform(0, 100, n)), 15.0 + rng.normal(0, 0.2, n), np.full(n, 0.1))
for _ in range(1000)
]
amplitudes = lc.Amplitude().many(light_curves)
print(f"Amplitude: mean = {np.mean(amplitudes):.3f} mag, std = {np.std(amplitudes):.3f} mag")
For conda, alternative package names, and platform-specific installation notes, see the Installation section.
Feature extractors
Most classes implement feature extractors useful for astrophysical source classification and characterization based on light curves.
To list all available feature extractors:
import light_curve as lc
print([x for x in dir(lc) if hasattr(getattr(lc, x), "names")])
To read the documentation for a specific extractor:
import light_curve as lc
help(lc.BazinFit)
Available features
See the complete list of available feature extractors and their documentation in the
light-curve-feature Rust crate docs.
Italic names are experimental features.
While we usually say "magnitude" and use "m" as the time-series value, some features are designed for
flux light curves.
The Flux/magnitude column indicates whether a feature should be used with flux light curves only, magnitude light curves only, or either.
The transform=True column shows the default transformation applied to the output when transform=True
is passed to the extractor constructor (default is transform=None, which does not apply any
transformations).
Transformations are monotonic functions intended to reduce dynamic range and improve the suitability of
raw feature values for machine learning.
You can also select a transformation by name string (e.g. transform="lg") for supported features;
available names are "identity" (no-op), "lg" (base-10 logarithm), "arcsinh" (inverse hyperbolic sine),
"clipped_lg" (base-10 logarithm with values below the minimum positive float clipped to a floor),
"ln1p" (natural logarithm of 1 + x), and "sqrt".
For parametric fit features (BazinFit, LinexpFit, VillarFit), transform=True applies a
feature-specific transformation described under Parametric fit features.
Features marked "N/A" do not implement transformation.
| Feature name | Description | Min data points | Features number | Flux/magnitude | `transform=True` |
|---|---|---|---|---|---|
| Amplitude | Half amplitude of magnitude: $\displaystyle \frac{\max (m)-\min (m)}{2}$ |
1 | 1 | Flux or magn | identity |
| AndersonDarlingNormal | Unbiased Anderson–Darling normality test statistic:
$\displaystyle \left( 1+\frac{4}{N} -\frac{25}{N^{2}}\right) \times$
$\times \left( -N-\frac{1}{N}\sum\limits_{i=0}^{N-1} (2i+1)\ln \Phi _{i} +(2(N-i)-1)\ln (1-\Phi _{i} )\right) ,$ where $\Phi _{i\ } \equiv \Phi (( m_{i} \ -\ \langle m\rangle ) /\sigma _{m})$ is the commutative distribution function of the standard normal distribution, $N-$ the number of observations, $\langle m\rangle -$ mean magnitude and $\sigma _{m} =\sqrt{\sum\limits_{i=0}^{N-1}( m_{i} -\langle m\rangle )^{2} /( N-1) \ }$ is the magnitude standard deviation |
4 | 1 | Flux or magn | lg |
| BazinFit | Five fit parameters and goodness of fit (reduced $\chi ^{2}$) of the Bazin function developed for core-collapsed supernovae:
$\displaystyle f(t)=A\frac{\mathrm{e}^{-(t-t_{0} )/\tau _{fall}}}{1+\mathrm{e}^{-(t-t_{0} )/\tau _{rise}}} +B,$ where $f(t)-$ flux observation |
6 | 1 | Flux only | $A$ → mag, $B/A$, $t_0$ dropped, $\chi^2$ → $\ln(1+\chi^2)$ |
| BeyondNStd | Fraction of observations beyond $n\sigma _{m}$ from the mean magnitude $\langle m\rangle $:
$\displaystyle \frac{\sum _{i} I_{|m-\langle m\rangle | >n\sigma _{m}} (m_{i} )}{N},$ where $I-$ an indicator function |
2 | 1 | Flux or magn | identity |
| ColorOfMedian (experimental) |
Magnitude difference between medians of two bands | 2 | 1 | Magn only | identity |
| Cusum | A range of cumulative sums:
$\displaystyle \max(S) - \min(S),$ where$\displaystyle S_{j} \equiv \frac{1}{N\sigma_{m}} \sum_{i=0}^{j} (m_{i} - \langle m\rangle), \quad j \in \lbrace 1..N-1 \rbrace\;$ |
2 | 1 | Flux or magn | identity |
| Eta | Von Neummann $\eta $:
$\displaystyle \eta \equiv \frac{1}{(N-1)\sigma _{m}^{2}}\sum\limits _{i=0}^{N-2} (m_{i+1} -m_{i} )^{2}$ |
2 | 1 | Flux or magn | identity |
| EtaE | Modernisation of Eta for unevenly spaced time series:
$\displaystyle \eta ^{e} \equiv \frac{(t_{N-1} -t_{0} )^{2}}{(N-1)^{3}}\frac{\sum\limits_{i=0}^{N-2}\left(\frac{m_{i+1} -m_{i}}{t_{i+1} -t_{i}}\right)^{2}}{\sigma _{m}^{2}}$ |
2 | 1 | Flux or magn | lg |
| ExcessVariance | Measure of the variability amplitude:
$\displaystyle \frac{\sigma _{m}^{2} -\langle \delta ^{2} \rangle }{\langle m\rangle ^{2}},$ where $\langle \delta ^{2} \rangle -$ mean squared error |
2 | 1 | Flux only | identity |
| FluxNNotDetBeforeFd (experimental) |
Number of non-detections before the first detection | 2 | 1 | Flux only | identity |
| InterPercentileRange | $\displaystyle Q(1-p)-Q(p),$ where $Q(n)$ and $Q(d)-$ $n$-th and $d$-th quantile of magnitude sample |
1 | 1 | Flux or magn | identity |
| Kurtosis | Excess kurtosis of magnitude:
$\displaystyle \frac{N(N+1)}{(N-1)(N-2)(N-3)}\frac{\sum _{i} (m_{i} -\langle m\rangle )^{4}}{\sigma _{m}^{2}} -3\frac{(N+1)^{2}}{(N-2)(N-3)}$ |
4 | 1 | Flux or magn | arcsinh |
| LinearFit | The slope, its error and reduced $\chi ^{2}$ of the light curve in the linear fit of a magnitude light curve with respect to the observation error $\{\delta _{i}\}$:
$\displaystyle m_{i} \ =\ c\ +\ \text{slope} \ t_{i} \ +\ \delta _{i} \varepsilon _{i} ,$ where $c$ is a constant, $\{\varepsilon _{i}\}$ are standard distributed random variables |
3 | 3 | Magn only | identity |
| LinearTrend | The slope and its error of the light curve in the linear fit of a magnitude light curve without respect to the observation error $\{\delta _{i}\}$:
$\displaystyle m_{i} \ =\ c\ +\ \text{slope} \ t_{i} \ +\ \Sigma \varepsilon _{i} ,$ where $c$ and $\Sigma$ are constants, $\{\varepsilon _{i}\}$ are standard distributed random variables. |
2 | 2 | Magn only | identity |
| MagnitudeNNotDetBeforeFd (experimental) |
Number of non-detections before the first detection | 2 | 1 | Magn only | identity |
| MagnitudePercentageRatio | Magnitude percentage ratio:
$\displaystyle \frac{Q(1-n)-Q(n)}{Q(1-d)-Q(d)}$ |
1 | 1 | Flux or magn | identity |
| MaximumSlope | Maximum slope between two consecutive observations:
$\displaystyle \max_{i=0\dotsc N-2}\left| \frac{m_{i+1} -m_{i}}{t_{i+1} -t_{i}}\right|$ |
2 | 1 | Flux or magn | clipped_lg |
| Mean | Mean magnitude:
$\displaystyle \langle m\rangle =\frac{1}{N}\sum\limits _{i} m_{i}$ |
1 | 1 | Flux or magn | identity |
| MeanVariance | Standard deviation to mean ratio:
$\displaystyle \frac{\sigma _{m}}{\langle m\rangle }$ |
2 | 1 | Flux only | identity |
| Median | Median magnitude | 1 | 1 | Flux or magn | identity |
| MedianAbsoluteDeviation | Median of the absolute value of the difference between magnitude and its median:
$\displaystyle \mathrm{Median} (|m_{i} -\mathrm{Median} (m)|)$ |
1 | 1 | Flux or magn | identity |
| MedianBufferRangePercentage | Fraction of points within $\displaystyle \mathrm{Median} (m)\pm q\times (\max (m)-\min (m))/2$ |
1 | 1 | Flux or magn | identity |
| OtsuSplit | Difference of subset means, standard deviation of the lower subset, standard deviation of the upper
subset and lower-to-all observation count ratio for two subsets of magnitudes obtained by Otsu's method split.
Otsu's method is used to perform automatic thresholding. The algorithm returns a single threshold that separates values into two classes. This threshold is determined by minimizing intra-class intensity variance $\sigma^2_{W}=w_0\sigma^2_0+w_1\sigma^2_1$, or equivalently, by maximizing inter-class variance $\sigma^2_{B}=w_0 w_1 (\mu_1-\mu_0)^2$. When multiple extrema exist, the algorithm returns the minimum threshold. |
2 | 4 | Flux or magn | N/A |
| PercentAmplitude | Maximum deviation of magnitude from its median:
$\displaystyle \max_{i} |m_{i} \ -\ \text{Median}( m) |$ |
1 | 1 | Flux or magn | identity |
| PercentDifferenceMagnitudePercentile | Ratio of $p$-th inter-percentile range to the median:
$\displaystyle \frac{Q( 1-p) -Q( p)}{\text{Median}( m)}$ |
1 | 1 | Flux only | clipped_lg |
| LinexpFit | Four fit parameters and goodness of fit (reduced $\chi^{2}$) of the Linexp function developed for core-collapsed supernovae:
$\displaystyle f(t)=A\frac{t-t_{0}}{\tau}\exp\!\left(\frac{t-t_{0}}{\tau}\right)+B$ |
5 | 5 | Flux only | $A$ → mag, $B/A$, $t_0$ dropped, $\chi^2$ → $\ln(1+\chi^2)$ |
| RainbowFit (experimental) |
Seven fit parameters and goodness of fit (reduced $\chi ^{2}$). The Rainbow method is developed and detailed in Russeil+23. This implementation is suited for transient objects. Bolometric flux and temperature functions are customizable; by default, Bazin and logistic functions are used:
$\displaystyle F_{\nu}(t, \nu) = \frac{\pi\,B\left(T(t),\nu\right)}{\sigma_\mathrm{SB}\,T(t)^{4}} \times F_\mathrm{bol}(t),$ where $F_{\nu}(t, \nu)-$ flux observation at a given wavelength |
6 | 1 | Flux only | identity |
| ReducedChi2 | Reduced $\chi ^{2}$ of magnitude measurements:
$\displaystyle \frac{1}{N-1}\sum _{i}\left(\frac{m_{i} -\overline{m}}{\delta _{i}}\right)^{2} ,$ where $\overline{m} -$ weighted mean magnitude |
2 | 1 | Flux or magn | ln1x |
| Roms (experimental) |
Robust median statistic: $\displaystyle \frac1{N-1} \sum_{i=0}^{N-1} \frac{|m_i - \mathrm{median}(m_i)|}{\sigma_i}$ |
2 | 1 | Flux or magn | identity |
| Skew | Skewness of magnitude:
$\displaystyle \frac{N}{(N-1)(N-2)}\frac{\sum _{i} (m_{i} -\langle m\rangle )^{3}}{\sigma _{m}^{3}}$ |
3 | 1 | Flux or magn | arcsinh |
| StandardDeviation | Standard deviation of magnitude:
$\displaystyle \sigma _{m} \equiv \sqrt{\sum _{i} (m_{i} -\langle m\rangle )^{2} /(N-1)}$ |
2 | 1 | Flux or magn | identity |
| StetsonK | Stetson K coefficient describing light curve shape:
$\displaystyle \frac{\sum _{i}\left| \frac{m_{i} -\overline{m} }{\delta _{i}}\right| }{\sqrt{N\ \chi ^{2}}}$ |
2 | 1 | Flux or magn | identity |
| VillarFit | Seven fit parameters and goodness of fit (reduced $\chi ^{2}$) of the Villar function developed for supernovae classification:
$f(t)=c+\frac{A}{1+\exp\frac{-(t-t_{0} )}{\tau _{rise}}} \times f_{fall}(t),$ $f_{fall}(t) = 1-\frac{\nu (t-t_{0} )}{\gamma }, ~~~ t< t_{0} +\gamma,$ $f_{fall}(t) = (1-\nu )\exp\frac{-(t-t_{0} -\gamma )}{\tau _{fall}}, ~~~ t \geq t_{0} + \gamma.$ where $f(t) -$ flux observation, $A, \gamma , \tau _{rise} , \tau _{fall} >0$, $\nu \in [0;1)$Here we introduce a new dimensionless parameter $\nu$ instead of the plateau slope $\beta$ from the original paper: $\nu \equiv -\beta \gamma /A$ |
8 | 8 | Flux only | $A$ → mag, $c/A$, $t_0$ dropped, $\chi^2$ → $\ln(1+\chi^2)$ |
| WeightedMean | Weighted mean magnitude:
$\displaystyle \overline{m} \equiv \frac{\sum _{i} m_{i} /\delta _{i}^{2}}{\sum _{i} 1/\delta _{i}^{2}}$ |
1 | 1 | Flux or magn | identity |
Meta-features
Meta-features accept other feature extractors and apply them to pre-processed data.
Periodogram
This feature transforms time-series data into the Lomb-Scargle periodogram, providing an estimate of the power
spectrum. The peaks argument specifies how many of the most significant spectral density peaks to return.
For
each peak, its period and signal-to-noise ratio are returned:
$$ \text{signal to noise of peak} \equiv \frac{P(\omega_\mathrm{peak}) - \langle P(\omega) \rangle}{\sigma_{P(\omega)}} $$
The optional features argument accepts a list of additional feature extractors, which are applied to the
power
spectrum: frequency is passed as "time," power spectrum is passed as "magnitude," and no uncertainties are
set.
Bins
Bins the time series into windows of width $\mathrm{window}$ with respect to some $\mathrm{offset}$. The $j$-th bin spans $[j \cdot \mathrm{window} + \mathrm{offset},\ (j + 1) \cdot \mathrm{window} + \mathrm{offset}]$.
The binned time series is defined by $$t_j^* = (j + \frac12) \cdot \mathrm{window} + \mathrm{offset},$$ $$m_j^* = \frac{\sum{m_i / \delta_i^2}}{\sum{\delta_i^{-2}}},$$ $$\delta_j^* = \frac{N_j}{\sum{\delta_i^{-2}}},$$ where $N_j$ is the number of observations in the bin and all sums are over observations within that bin.
Parametric fit features
BazinFit, LinexpFit, and VillarFit fit parametric functions to transient flux light curves (see
Available features for the function definitions). Unlike other extractors,
they require an explicit algorithm argument selecting the optimization method.
The available algorithms depend on the compile-time Cargo features (see Build from source):
| Algorithm | Requires | Description |
|---|---|---|
"mcmc" |
always available | MCMC ensemble sampler; robust but slow |
"ceres" |
ceres-source or ceres-system |
Ceres trust-region solver; fast, gradient-based |
"mcmc-ceres" |
ceres-source or ceres-system |
MCMC exploration followed by Ceres refinement |
"lmsder" |
gsl |
Levenberg-Marquardt via GSL |
"mcmc-lmsder" |
gsl |
MCMC exploration followed by LMSDER refinement |
The hybrid "mcmc-ceres" and "mcmc-lmsder" algorithms run MCMC first to broadly explore the
parameter space, then hand off to the gradient-based solver for precise convergence. This typically
gives the best balance of robustness and final accuracy.
By default, initial parameter values and bounds are estimated from the data. Override them with the
init and bounds arguments (supported by MCMC-based algorithms only), each a list of values or
Nones to keep data-derived defaults for individual parameters. BazinFit has 5 parameters, LinexpFit has
4,
VillarFit has 7; run help() on the respective class for the parameter order.
The ln_prior argument sets the MCMC prior. It accepts None / "no" (flat prior, the default),
a named string literal, or a list of light_curve.ln_prior.LnPrior1D objects for per-parameter
distributions (see help(light_curve.ln_prior) for the available distribution types).
VillarFit additionally supports "hosseinzadeh2020", a prior from Hosseinzadeh et al. 2020 that
encodes supernova physics constraints; it assumes time values are in days.
Pass transform=True to convert the raw fit parameters to a magnitude-like representation: the
amplitude becomes a magnitude (assuming input flux in janskys), the baseline is normalized by the amplitude,
the
reference time is
dropped, and the reduced chi^2 is log-scaled. Run help(lc.BazinFit) for the exact definition.
For the experimental multi-band analogue, see Rainbow Fit below.
Multi-band features
As of v0.8, experimental extractors support multi-band light-curve inputs.
import numpy as np
from light_curve.light_curve_py import LinearFit
t = np.arange(20, dtype=float)
m = np.arange(20, dtype=float)
sigma = np.full_like(t, 0.1)
bands = np.array(["g"] * 10 + ["r"] * 10)
feature = LinearFit(bands=["g", "r"])
values = feature(t, m, sigma, bands)
print(values)
Rainbow Fit
Rainbow (Russeil+23) is a black-body parametric model for transient light
curves. By default, it uses the Bazin function as a model for bolometric flux evolution and a logistic
function
for temperature evolution. The user may customize the model by providing their own functions for bolometric
flux
and temperature evolution. This example demonstrates the reconstruction of a synthetic light curve with this
model.
RainbowFit requires the iminuit package.
import numpy as np
from light_curve.light_curve_py import RainbowFit
def bb_nu(wave_aa, T):
"""Black-body spectral model"""
nu = 3e10 / (wave_aa * 1e-8)
return 2 * 6.626e-27 * nu ** 3 / 3e10 ** 2 / np.expm1(6.626e-27 * nu / (1.38e-16 * T))
# Effective wavelengths in Angstrom
band_wave_aa = {"g": 4770.0, "r": 6231.0, "i": 7625.0, "z": 9134.0}
# Parameter values
reference_time = 60000.0 # time close to the peak time
# Bolometric flux model parameters
amplitude = 1.0 # bolometric flux semiamplitude, arbitrary (non-spectral) flux/luminosity units
rise_time = 5.0 # exponential growth timescale, days
fall_time = 30.0 # exponential decay timescale, days
# Temperature model parameters
Tmin = 5e3 # temperature on +infinite time, kelvins
delta_T = 10e3 # (Tmin + delta_T) is temperature on -infinite time, kelvins
k_sig = 4.0 # temperature evolution timescale, days
rng = np.random.default_rng(0)
t = np.sort(rng.uniform(reference_time - 3 * rise_time, reference_time + 3 * fall_time, 1000))
band = rng.choice(list(band_wave_aa), size=len(t))
waves = np.array([band_wave_aa[b] for b in band])
# Temperature evolution is a sigmoid function
temp = Tmin + delta_T / (1.0 + np.exp((t - reference_time) / k_sig))
# Bolometric flux evolution is the Bazin function
lum = amplitude * np.exp(-(t - reference_time) / fall_time) / (
1.0 + np.exp(-(t - reference_time) / rise_time))
# Spectral flux density for each given pair of time and passband
flux = np.pi * bb_nu(waves, temp) / (5.67e-5 * temp ** 4) * lum
# S/N = 5 for minimum flux, scale for Poisson noise
flux_err = np.sqrt(flux * np.min(flux) / 5.0)
flux += rng.normal(0.0, flux_err)
feature = RainbowFit.from_angstrom(band_wave_aa, with_baseline=False)
values = feature(t, flux, sigma=flux_err, band=band)
print(dict(zip(feature.names, values)))
print(f"Goodness of fit: {values[-1]}")
Note that while the data generation above uses approximate physical constant values, RainbowFit uses CODATA
2018
values internally.
Experimental extractors
The package consists of two parts: a wrapper for the
light-curve-feature Rust crate (light_curve_ext
sub-package)
and a pure-Python sub-package light_curve_py.
We use the Python implementation to test the Rust implementation and to develop new experimental extractors.
The Python implementation is significantly slower for most extractors and does not provide the same full
feature set as the Rust implementation, but it does include some extractors you may find useful.
You can use extractors from either implementation directly:
import numpy as np
from numpy.testing import assert_allclose
from light_curve.light_curve_ext import LinearTrend as RustLinearTrend
from light_curve.light_curve_py import LinearTrend as PythonLinearTrend
rust_fe = RustLinearTrend()
py_fe = PythonLinearTrend()
n = 100
t = np.sort(np.random.normal(size=n))
m = 3.14 * t - 2.16 + np.random.normal(size=n)
assert_allclose(rust_fe(t, m), py_fe(t, m),
err_msg="Python and Rust implementations must provide the same result")
This should print a warning about the experimental status of the Python class.
Performance
The package is designed for high throughput. The following techniques help extract features with minimal overhead.
sorted and check parameters
The sorted=True argument tells the extractor that t is already sorted in ascending order, and
check=False
disables validation of NaN/inf values. The defaults are sorted=None (the array will be validated and an
error raised
if unsorted) and
check=True. Passing invalid inputs without validation can cause incorrect results or crashes.
Batch processing with .many()
Each extractor has a .many() method that processes a collection of light curves in parallel across all
available CPU cores by default. It accepts either a list of (t, m[, sigma]) tuples or an Arrow array
(see below).
import light_curve as lc
import numpy as np
rng = np.random.default_rng(0)
# A list of (t, m, sigma) tuples, one per light curve
light_curves = [
(np.sort(rng.random(50)), rng.random(50), rng.random(50) * 0.1)
for _ in range(1000)
]
results = lc.Amplitude().many(light_curves)
Arrow input
The .many() method also accepts Arrow arrays instead of a list of tuples, enabling zero-copy data access
from Arrow-compatible libraries. The input must be a List<Struct<t, m[, sigma]>> array where all fields
share the same float dtype. Any library implementing the
Arrow C Data Interface is supported.
All light curves are processed in parallel using Rust threads, bypassing the GIL entirely.
nested-pandas
nested-pandas extends pandas with nested column support,
useful for working with catalog data like ZTF or Rubin LSST.
Install with pip install nested-pandas s3fs universal-pathlib.
import light_curve as lc
import nested_pandas as npd
import numpy as np
import pyarrow as pa
from upath import UPath
# Read a ZTF DR23 HATS partition with nested light curves
s3_path = UPath(
"s3://ipac-irsa-ztf/contributed/dr23/lc/hats/ztf_dr23_lc-hats/dataset/Norder=6/Dir=30000/Npix=34623/part0.snappy.parquet",
anon=True,
)
ndf = npd.read_parquet(
s3_path,
columns=["objectid", "lightcurve.hmjd", "lightcurve.mag", "lightcurve.magerr"],
)
# Select rows with more than 10 observations
ndf = ndf.loc[ndf["lightcurve"].list_lengths > 10]
# We must have subcolumns to be the same dtype, so we convert time to float32.
ndf["lightcurve.t"] = np.asarray(ndf["lightcurve.hmjd"] - 58000, dtype=np.float32)
# Extract features directly from the Arrow-backed nested column
feature = lc.Extractor(lc.Amplitude(), lc.EtaE(), lc.InterPercentileRange(quantile=0.25))
result = feature.many(pa.array(ndf["lightcurve"]), n_jobs=-1, arrow_fields=["t", "mag", "magerr"])
# Assign features back to the dataframe
ndf = ndf.assign(**dict(zip(feature.names, result.T)))
print(ndf.head())
PyArrow
PyArrow is the reference Python implementation of Apache Arrow.
Install with pip install pyarrow.
import light_curve as lc
import numpy as np
import pyarrow as pa
rng = np.random.default_rng(42)
n_lc = 10
# Build a PyArrow List<Struct<t, m, sigma>> array
struct_type = pa.struct([("t", pa.float64()), ("m", pa.float64()), ("sigma", pa.float64())])
lcs_arrow = pa.array(
[
[{"t": t, "m": m, "sigma": s} for t, m, s in zip(*[np.sort(rng.random(50)) for _ in range(3)])]
for _ in range(n_lc)
],
type=pa.list_(struct_type),
)
feature = lc.Extractor(lc.Kurtosis(), lc.Skew(), lc.ReducedChi2())
result = feature.many(lcs_arrow, sorted=True, check=False, n_jobs=4, arrow_fields=["t", "m", "sigma"])
print(f"Features: {feature.names}")
print(f"Results shape: {result.shape}")
Polars
Polars is a fast DataFrame library built on Arrow.
Install with pip install polars.
import light_curve as lc
import numpy as np
import polars as pl
rng = np.random.default_rng(42)
# Start with a flat DataFrame, as you might get from a database or CSV
object_id = np.repeat(np.arange(10), 50)
t = np.sort(rng.random(500))
m = rng.random(500)
sigma = rng.random(500)
df = pl.DataFrame({"object_id": object_id, "t": t, "m": m, "sigma": sigma})
# Group by object_id into List<Struct<t, m, sigma>>
nested = df.group_by("object_id").agg(pl.struct("t", "m", "sigma").alias("lc"))
feature = lc.Extractor(lc.Amplitude(), lc.BeyondNStd(nstd=2), lc.LinearFit(), lc.StetsonK())
result = feature.many(nested["lc"], sorted=True, check=False, n_jobs=1, arrow_fields=["t", "m", "sigma"])
# Join feature columns back to the nested DataFrame
nested = nested.with_columns(
[pl.Series(name, result[:, i]) for i, name in enumerate(feature.names)]
)
print(nested)
Benchmarks
Run all benchmarks from the Python project folder with
python3 -mpytest --benchmark-enable tests/test_w_bench.py, or with slow benchmarks disabled:
python3 -mpytest -m "not (nobs or multi)" --benchmark-enable tests/test_w_bench.py.
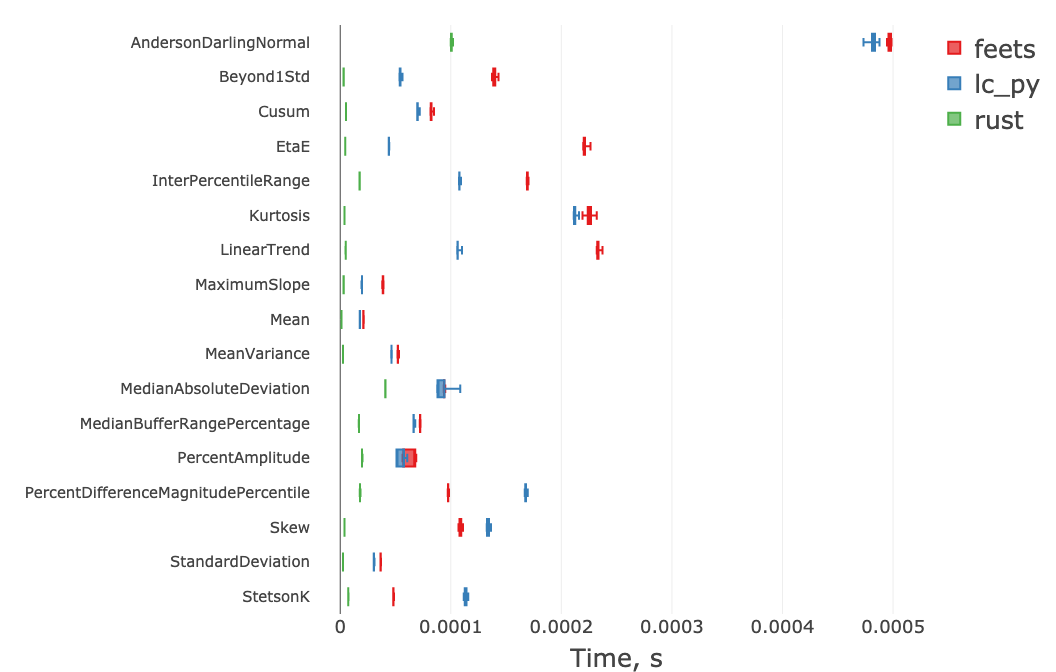
Below we benchmark the Rust implementation (rust) against the
feets
package and our own Python implementation (lc_py) for a light curve with n=1000 observations.
The Rust implementation outperforms the alternatives by a factor of 1.5–50.
This allows extracting a large set of "cheap" features in well under one millisecond for n=1000.
The performance of parametric fits (BazinFit and VillarFit) and Periodogram depends on their parameters,
but the typical extraction time including these features is 20–50 ms for a few hundred observations.
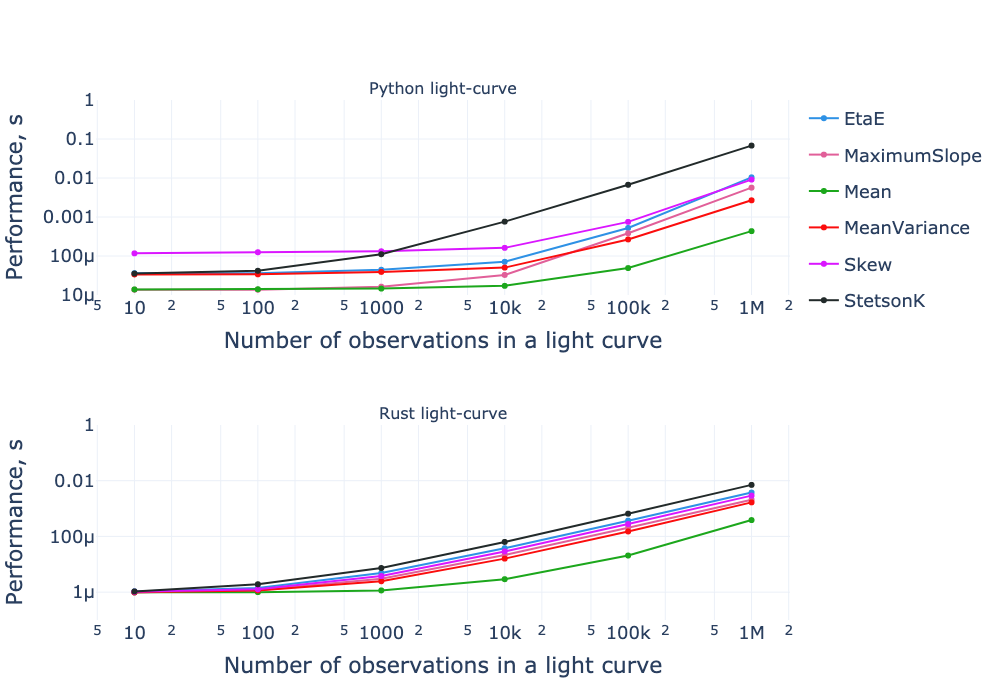
Benchmark results for both the pure-Python and Rust implementations as a function of the number of observations. Both axes are on a logarithmic scale.
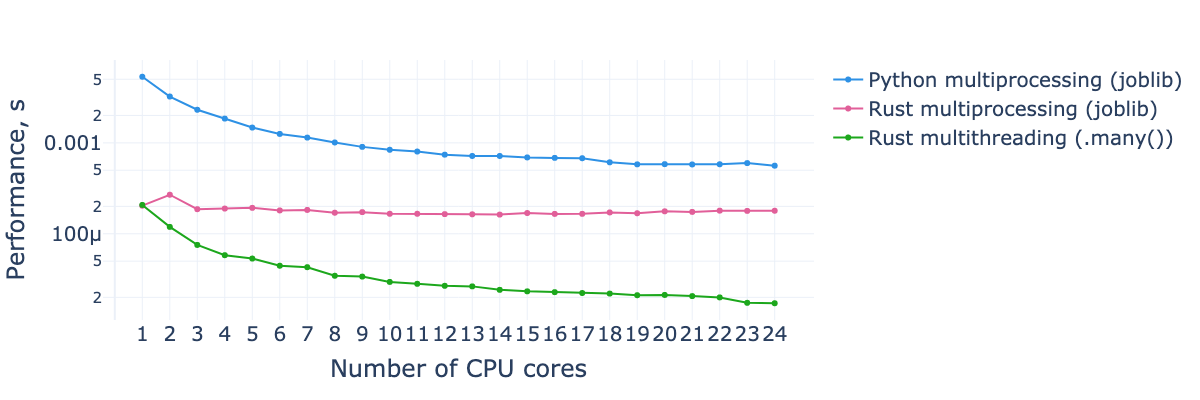
Processing time per light curve for the feature subset from the first benchmark, as a function of the number of CPU cores used. The dataset consists of 10,000 light curves with 1,000 observations each.
See the benchmarks described in more detail in "Performant feature extraction for photometric time series".
Light curve embeddings (light_curve.embed)
The light_curve.embed submodule provides pretrained neural network models that map raw photometric
time series to dense vector embeddings suitable for downstream machine learning tasks such as
classification, anomaly detection, and similarity search.
The examples below use from_hf() to download model weights from HuggingFace Hub
(cached locally after the first download), which requires onnxruntime and huggingface-hub:
pip install light-curve onnxruntime huggingface_hub
Models can also be loaded directly from a local ONNX file via from_onnx_file(),
without huggingface-hub.
See the onnxruntime install guide for GPU and platform-specific
packages (onnxruntime-gpu, onnxruntime-directml, etc.).
Hardware-specific options such as execution providers can be passed via ort_session_kwargs
(see onnxruntime Python API).
Single-band models: Astromer
Astromer1 (Donoso-Oliva et al. 2023)
and Astromer2 (Donoso-Oliva et al. 2026)
are models pretrained on MACHO light curves.
They accept irregularly-sampled (time, mag) pairs and return 256-dimensional embeddings.
import numpy as np
from light_curve.embed import Astromer2
model = Astromer2.from_hf(output="mean")
# The default reduction splits long light curves into non-overlapping windows;
# pass e.g. reduction="beginning" to always take the first 200 observations.
rng = np.random.default_rng(0)
time = np.sort(rng.uniform(0, 500, 120)).astype(np.float64)
mag = rng.normal(15, 0.5, 120).astype(np.float64)
# Returns shape (n_bands, n_subsamples, seq_size, embed_dim)
embedding = model(time, mag)
print(embedding.shape) # (1, 1, 1, 256)
Multi-band light curves can be embedded per-band by passing a list of band labels to the bands
constructor argument:
import numpy as np
from light_curve.embed import Astromer2
model = Astromer2.from_hf(output="mean", bands=["g", "r"])
rng = np.random.default_rng(1)
n = 80
time = np.sort(rng.uniform(0, 300, n)).astype(np.float64)
mag = rng.normal(15, 0.5, n).astype(np.float64)
band = np.array(["g", "r"] * (n // 2))
# n_bands=2: one embedding per band
embedding = model(time, mag, band=band)
print(embedding.shape) # (2, 1, 1, 256)
Multi-band model: ATCAT
ATCAT (Tung 2025) is a model
trained on ELAsTiCC light curves. It processes all six LSST bands jointly and returns
384-dimensional embeddings. Inputs are flux, flux-error, time, and integer band indices (ugrizY → 012345):
import numpy as np
from light_curve.embed import ATCAT
model = ATCAT.from_hf(output="last")
rng = np.random.default_rng(2)
n = 120
time = np.sort(rng.uniform(0, 500, n)).astype(np.float32)
flux = rng.normal(100, 10, n).astype(np.float32)
flux_err = np.full(n, 5.0, dtype=np.float32)
band = np.array([i % 6 for i in range(n)]) # ugrizY → 012345
embedding = model(time, flux, flux_err, band)
print(embedding.shape) # (1, 1, 1, 384)
Input fluxes should be in AB units. The default zero-point is 31.4 (LSST nJy); set mag_zp=27.5
for ELAsTiCC / SNANA FITS data or mag_zp=8.9 for Jy.
Example: nearest-neighbour search on ZTF DR23
The following example reads a ZTF DR23 HATS pixel directly from the public S3
bucket using
nested-pandas and s3fs
(install all dependencies for this example with
pip install huggingface_hub light-curve nested-pandas onnxruntime s3fs scipy), embeds all well-observed
light curves with Astromer2, and finds the closest neighbour to a given object by cosine distance.
This particular pixel contains ~50k objects passing the quality cuts; embedding takes roughly
12 minutes on M2 Pro (~15 ms per object).
import numpy as np
import nested_pandas as npd
from scipy.spatial.distance import cdist
from upath import UPath
from light_curve.embed import Astromer2
TARGET_OID = 680213300009232 # ZTF r-band light curve
MIN_OBS = 1000
nf = npd.read_parquet(UPath(
"s3://ipac-irsa-ztf/contributed/dr23/lc/hats/ztf_dr23_lc-hats/dataset/Norder=5/Dir=0/Npix=2378/",
anon=True,
))
nf = nf.query("lightcurve.catflags == 0").query(f"lightcurve.list_lengths >= {MIN_OBS}")
model = Astromer2.from_hf(reduction="beginning")
def embed_row(row):
return {"embedding.values": model(row["lightcurve.hmjd"], row["lightcurve.mag"]).squeeze()}
nf = nf.map_rows(embed_row, columns=["lightcurve.hmjd", "lightcurve.mag"], append_columns=True)
oids = nf["objectid"].to_numpy()
matrix = nf["embedding.values"].to_numpy().reshape(len(nf), -1)
query_idx = np.where(oids == TARGET_OID)[0][0]
distances = cdist(matrix[query_idx: query_idx + 1], matrix, metric="cosine")[0]
distances[query_idx] = np.inf
best_idx = np.argmin(distances)
print(f"Nearest neighbour: OID {oids[best_idx]}, cosine distance {distances[best_idx]:.6f}")
# Nearest neighbour: OID 680113300005170, cosine distance 0.000046
The nearest neighbour is OID 680113300005170 — the same physical
object (HZ Her / Her X-1, an X-ray binary) observed in the g
-band,
recovered automatically from an r-band query through embedding similarity.
See the light curve on SNAD Viewer.
dm-dt map
In addition to the feature extractors above, the package provides a separate dm–dt mapping tool. The dm–dt map is a 2-D histogram of magnitude differences (dm) versus time differences (dt) for all pairs of observations in a light curve. It is commonly used as an input representation for machine learning classifiers rather than as a scalar feature.
The DmDt class (a Python wrapper for the
light-curve-dmdt Rust crate) implements this mapper,
following Mahabal et al. 2011 and
Soraisam et al. 2020.
import numpy as np
from light_curve import DmDt
from numpy.testing import assert_array_equal
dmdt = DmDt.from_borders(min_lgdt=0, max_lgdt=np.log10(3), max_abs_dm=3, lgdt_size=2, dm_size=4, norm=[])
t = np.array([0, 1, 2], dtype=np.float32)
m = np.array([0, 1, 2], dtype=np.float32)
desired = np.array(
[
[0, 0, 2, 0],
[0, 0, 0, 1],
]
)
actual = dmdt.points(t, m)
assert_array_equal(actual, desired)
Installation
pip install 'light-curve[full]'
# or
conda install -c conda-forge light-curve-python
The full extra installs all optional Python dependencies required by experimental features.
We also provide a light-curve-python package as an alias for light-curve[full].
The minimum supported Python version is 3.10. Binary CPython distributions are available via PyPI and Anaconda for various platforms and architectures. On PyPI, we publish wheels for the stable CPython ABI, ensuring compatibility with future CPython 3 versions.
Support matrix
| Arch \ OS | Linux glibc 2.17+ | Linux musl 1.2+ | macOS | Windows |
|---|---|---|---|---|
| x86-64 | PyPI (MKL), conda | PyPI (MKL) | PyPI macOS 15+, conda | PyPI, conda (conda: no GSL) |
| i686 | src | src | — | not tested |
| aarch64 | PyPI | PyPI | PyPI macOS 14+, conda | not tested |
| ppc64le | src | not tested (no Rust toolchain) | — | — |
- PyPI / conda: A binary wheel or package is available on pypi.org or anaconda.org.
Local building is not required; the only prerequisite is a recent version of
piporconda. For Linux x86-64, PyPI wheels are built with Intel MKL for improved periodogram performance, which is not the default build option. For Windows x86-64, the conda package excludes GSL support (LMSDER fitting unavailable); PyPI wheels include both Ceres and GSL. - src: The package has been confirmed to build and pass unit tests locally, but CI does not test or publish packages for this platform. See the Build from source section for details. Please open an issue or pull request if you encounter build problems or would like us to distribute packages for these platforms.
- not tested: Building from source has not been tested. Please report build status via issue, PR, or email.
macOS wheels require relatively recent OS versions; please open an issue if you need support for older macOS versions (see https://github.com/light-curve/light-curve-python/issues/376 for details).
We no longer publish PyPy wheels (#345) or the PPC64le CPython glibc wheel (#479); please open an issue if you need either.
Free-threaded Python is supported when built from source for Python 3.14+
(officially supported in Python 3.14+).
No pre-built distributions are currently provided; please comment on the relevant issues if you need them:
PyPI binary wheel issue,
conda-forge package issue.
Note that for expensive features, the GIL-enabled interpreter with .many() achieves performance on par with
the free-threaded interpreter using Python threads. For inexpensive extractors, .many() still reduces
Rust–Python interaction overhead significantly.
Developer guide
Prepare environment
Install a recent Rust toolchain and Python 3.10 or higher.
It is recommended to use rustup to install the Rust toolchain and keep it updated
with rustup update.
Clone the repository, then create and activate a virtual environment:
git clone https://github.com/light-curve/light-curve-python.git
cd light-curve-python/light-curve
python3 -m venv venv
source venv/bin/activate
Install the package in editable mode (see Build from source for more details):
python -mpip install maturin
# --release takes longer to build but produces a faster package
# Add other Cargo flags if needed, e.g. --no-default-features --features=ceres-source
maturin develop --group dev
Run this command during initial setup. On subsequent runs, activate the environment with
source venv/bin/activate
and rebuild Rust code with maturin develop. Python-only changes require no rebuild. The --group dev flag
(run from the light-curve/ directory) is only needed once to install development dependencies.
It is also recommended to install pre-commit hooks to keep the codebase clean.
Install via pip (see the documentation for other options), then run:
pre-commit install
Run tests and benchmarks
All test dependencies are installed with maturin develop --group dev. Run the tests with:
python -mpytest
Benchmarks are disabled by default; enable them with --benchmark-enable:
python -mpytest --benchmark-enable
See the Benchmarks section in Performance for more details.
Build from source
Dependencies and Cargo features
The package has a number of compile-time Cargo features, mostly controlling which C/C++ dependencies are used.
Pass the desired features to maturin with --features; it is also recommended to use
--no-default-features
to avoid building unnecessary dependencies.
Available features:
abi3(default) — enables CPython ABI3 compatibility. Disable for other interpreters or if you have a specific reason (our benchmarks show no performance difference). ABI3 is not supported by free-threaded CPython or PyPy.ceres-source(default) — builds Ceres solver from source. Requires a C++ compiler and CMake. Used as an optional optimization backend forBazinFitandVillarFit.ceres-system— links against a system-installed Ceres dynamic library instead of building from source.mkl— enables the FFTW interface with the Intel MKL backend for the fast periodogram. Intel MKL is downloaded automatically during the build. Highly recommended for Intel CPUs. Without this feature, the pure-Rust RustFFT backend is used.gsl(default) — enables GNU Scientific Library support. Requires a compatible GSL version installed on your system. Used as an optional optimization backend forBazinFitandVillarFit.mimalloc(default) — enables the mimalloc memory allocator. Benchmarks show up to a 2× speedup for some cheap features, though it may increase memory usage.
Build with maturin
maturin is the recommended tool for building Rust-backed Python packages. This example builds with minimal system dependencies:
python -mpip install maturin
maturin build --release --locked --no-default-features --features=abi3,mimalloc
The --release flag enables release mode (slower build, faster execution), --locked ensures reproducible
builds, and --no-default-features --features=abi3,mimalloc selects only ABI3 compatibility and the mimalloc
allocator.
Build with build
You can also build the package with build:
python -mpip install build
MATURIN_PEP517_ARGS="--locked --no-default-features --features=abi3,mimalloc" python -m build
Build with cibuildwheel
cibuildwheel builds wheels on CI servers; we use it with GitHub Actions. You can also use it locally to build wheels for your platform (replace the identifier with one from the list of supported platforms):
python -mpip install cibuildwheel
python -m cibuildwheel --only=cp310-manylinux_x86_64
Note that different Cargo feature sets are used for different platforms, as defined in pyproject.toml.
Windows wheels must be built on Windows; Linux wheels can be built on any platform with Docker (Qemu may be
needed for cross-architecture builds); macOS wheels must be built on macOS.
macOS builds also require MACOSX_DEPLOYMENT_TARGET to match the current macOS version, since Homebrew
dependencies are built against it:
export MACOSX_DEPLOYMENT_TARGET=$(sw_vers -productVersion | awk -F '.' '{print $1"."0}')
python -m cibuildwheel --only=cp310-macosx_arm64
unset MACOSX_DEPLOYMENT_TARGET
Since we use ABI3 compatibility, a wheel built for CPython 3.10 works with any later CPython 3 version.
Citation
If you found this project useful for your research, please cite Malanchev et al., 2021:
@ARTICLE{2021MNRAS.502.5147M,
author = {{Malanchev}, K.~L. and {Pruzhinskaya}, M.~V. and {Korolev}, V.~S. and {Aleo}, P.~D. and {Kornilov}, M.~V. and {Ishida}, E.~E.~O. and {Krushinsky}, V.~V. and {Mondon}, F. and {Sreejith}, S. and {Volnova}, A.~A. and {Belinski}, A.~A. and {Dodin}, A.~V. and {Tatarnikov}, A.~M. and {Zheltoukhov}, S.~G. and {(The SNAD Team)}},
title = "{Anomaly detection in the Zwicky Transient Facility DR3}",
journal = {\mnras},
keywords = {methods: data analysis, astronomical data bases: miscellaneous, stars: variables: general, Astrophysics - Instrumentation and Methods for Astrophysics, Astrophysics - Solar and Stellar Astrophysics},
year = 2021,
month = apr,
volume = {502},
number = {4},
pages = {5147-5175},
doi = {10.1093/mnras/stab316},
archivePrefix = {arXiv},
eprint = {2012.01419},
primaryClass = {astro-ph.IM},
adsurl = {https://ui.adsabs.harvard.edu/abs/2021MNRAS.502.5147M},
adsnote = {Provided by the SAO/NASA Astrophysics Data System}
}
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distributions
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file light_curve-0.12.4.tar.gz.
File metadata
- Download URL: light_curve-0.12.4.tar.gz
- Upload date:
- Size: 468.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
33d0ea79cc054d8d33550cba9c2f23ddb89dc14324b8280314f4861b5e9a0a91
|
|
| MD5 |
ff2cebc15658b00a1d3a43a30f2f8ee5
|
|
| BLAKE2b-256 |
a2655b5c1182426c4267605ac131a0926328c7a7e4c80db3c7b9781d034dedaa
|
File details
Details for the file light_curve-0.12.4-cp310-abi3-win_amd64.whl.
File metadata
- Download URL: light_curve-0.12.4-cp310-abi3-win_amd64.whl
- Upload date:
- Size: 6.3 MB
- Tags: CPython 3.10+, Windows x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
dd2ef22155291620362bc64944fb59ab5851ae0b267e6e35c3a9cdc79479dc03
|
|
| MD5 |
765cb0a12cad6e9f61f1a03c018d23a9
|
|
| BLAKE2b-256 |
2dfa3848ae6e62299f152f2bb0b3073a9ddd549652bb3a7d698afaa02592c8c7
|
File details
Details for the file light_curve-0.12.4-cp310-abi3-musllinux_1_2_x86_64.whl.
File metadata
- Download URL: light_curve-0.12.4-cp310-abi3-musllinux_1_2_x86_64.whl
- Upload date:
- Size: 31.8 MB
- Tags: CPython 3.10+, musllinux: musl 1.2+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c33b3511f2010884a86d559408a0e4667be2eeb9c6a0acf3fdca452a4fa39fee
|
|
| MD5 |
6ac8da3988c8abd9f750c8faf2bfb833
|
|
| BLAKE2b-256 |
4750bac6ea0e19827d3a892f63a98c37f40585305f3fc402c6caf499092a6536
|
File details
Details for the file light_curve-0.12.4-cp310-abi3-musllinux_1_2_aarch64.whl.
File metadata
- Download URL: light_curve-0.12.4-cp310-abi3-musllinux_1_2_aarch64.whl
- Upload date:
- Size: 8.0 MB
- Tags: CPython 3.10+, musllinux: musl 1.2+ ARM64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
457cf055b7f98eedadc713275990089129f1282852abc79dd697b7ab84a24a7b
|
|
| MD5 |
362cc57116655e8b93d5ccd393a74c50
|
|
| BLAKE2b-256 |
684b5c600104c0e31ad267eddab502631cf87a1ff7d09e263de212c85ba310ac
|
File details
Details for the file light_curve-0.12.4-cp310-abi3-manylinux2014_x86_64.manylinux_2_17_x86_64.whl.
File metadata
- Download URL: light_curve-0.12.4-cp310-abi3-manylinux2014_x86_64.manylinux_2_17_x86_64.whl
- Upload date:
- Size: 45.8 MB
- Tags: CPython 3.10+, manylinux: glibc 2.17+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8322ad19e517219a592d0e90cf23e7789a6adfc4f3f77c4ed0af45f68c604e17
|
|
| MD5 |
06cae8c37be2bd8308b664888cf328fa
|
|
| BLAKE2b-256 |
3b1140c945c6128f63746c12a9c05d7404cc34db4a6913f64bd3df079469bc89
|
File details
Details for the file light_curve-0.12.4-cp310-abi3-manylinux2014_aarch64.manylinux_2_17_aarch64.whl.
File metadata
- Download URL: light_curve-0.12.4-cp310-abi3-manylinux2014_aarch64.manylinux_2_17_aarch64.whl
- Upload date:
- Size: 6.1 MB
- Tags: CPython 3.10+, manylinux: glibc 2.17+ ARM64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9c63dcb9ba7184beba22c3d352d7abd181f6eb0eb2bae90479f9610ed986b0f0
|
|
| MD5 |
083d67399d7d654ec8d0bdda0de440eb
|
|
| BLAKE2b-256 |
083ca0b76de303e2168619f52c6b5c80bac17fe178fec88e3f9ad47a28a7265f
|
File details
Details for the file light_curve-0.12.4-cp310-abi3-macosx_15_0_x86_64.whl.
File metadata
- Download URL: light_curve-0.12.4-cp310-abi3-macosx_15_0_x86_64.whl
- Upload date:
- Size: 7.6 MB
- Tags: CPython 3.10+, macOS 15.0+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5ebcd1210a02011350bc8fd68fc1d532c0c7062ce585c1fc0fd32bd0da324ee7
|
|
| MD5 |
e202f9dfbe61c68ca6a8f99a2242f9f7
|
|
| BLAKE2b-256 |
dee9d402a09c6223ecc0f7b86c1a0ea70371dd2e09edac4c4a4916cd70eadf5e
|
File details
Details for the file light_curve-0.12.4-cp310-abi3-macosx_14_0_arm64.whl.
File metadata
- Download URL: light_curve-0.12.4-cp310-abi3-macosx_14_0_arm64.whl
- Upload date:
- Size: 6.4 MB
- Tags: CPython 3.10+, macOS 14.0+ ARM64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
0ab51c9b5abeb73d5c2c0ddb6090f01c8f5a320ee6ece18f041858719378b31d
|
|
| MD5 |
5e36370895b28771374b459556d38732
|
|
| BLAKE2b-256 |
38958cf16ca5e4ac130df428eaaea44de04f31684722d06a4465e54a3b853847
|