Dynamic feature distillation framework for robust zero-shot LLM annotation.
Project description
LLM-scCurator

Cite: bioRxiv DOI: 10.64898/2025.12.28.696778 • Zenodo (v0.1.0) DOI: 10.5281/zenodo.17970494
🚀 Overview
LLM-scCurator is a Large Language Model–based curator for single-cell and spatial transcriptomics. It performs noise-aware marker distillation—suppressing technical programs (e.g., ribosomal/mitochondrial), clonotype signals (TCR/Ig), and stress signatures while rescuing lineage markers—and applies leakage-safe lineage filters before prompting an LLM. It supports hierarchical (coarse-to-fine) annotation for single-cell and spatial transcriptomics data. See the documentation for full tutorials and API reference: https://llm-sccurator.readthedocs.io/
Key Features
- 🛡️ Noise-aware filtering: Automatically removes lineage-specific noise (TCR/Ig) and state-dependent noise (ribosomal/mitochondrial).
- 🧠 Context-aware inference: Automatically infers lineage context (e.g., "T cell") to guide LLM reasoning.
- 🔬 Hierarchical discovery: One-line function to dissect complex tissues into major lineages and fine-grained subtypes.
- 🌍 Spatial ready: Validated on scRNA-seq (10x) and spatial transcriptomics (Xenium, Visium).
📦 Installation
-
Option A (recommended): Install from PyPI
pip install llm-sc-curator
(See PyPI project page: https://pypi.org/project/llm-sc-curator/)
-
Option B: Install from GitHub (development)
# 1. Clone the repository git clone https://github.com/kenflab/LLM-scCurator.git # 2. Navigate to the directory cd LLM-scCurator # 3. Install the package (and dependencies) pip install .
Notes:
If you already have a Scanpy/Seurat pipeline environment, you can install it into that environment.
🐳 Docker (official environment)
We provide an official Docker environment (Python + R + Jupyter), sufficient to run LLM-scCurator and most paper figure generation.
-
Option A: Prebuilt image (recommended)
Use the published image from GitHub Container Registry (GHCR).
# from the repo root (optional, for notebooks / file access) docker pull ghcr.io/kenflab/llm-sc-curator:official
Run Jupyter:
docker run --rm -it \ -p 8888:8888 \ -v "$PWD":/work \ -e GEMINI_API_KEY \ -e OPENAI_API_KEY \ ghcr.io/kenflab/llm-sc-curator:officialOpen Jupyter: http://localhost:8888
(Use the token printed in the container logs.)
Notes:For manuscript reproducibility, we also provide versioned tags (e.g., :0.1.0). Prefer a version tag when matching a paper release.
-
Option B: Build locally (development)
# Option B1: Build locally with Compose # from the repo root docker compose -f docker/docker-compose.yml build docker compose -f docker/docker-compose.yml up
Open Jupyter: http://localhost:8888
Workspace mount: /work# Option B2: Build locally without Compose (alternative) # from the repo root docker build -f docker/Dockerfile -t llm-sc-curator:official .
Run Jupyter:
docker run --rm -it \ -p 8888:8888 \ -v "$PWD":/work \ -e GEMINI_API_KEY \ -e OPENAI_API_KEY \ llm-sc-curator:officialOpen Jupyter: http://localhost:8888
🖥️ Apptainer / Singularity (HPC)
-
Option A: Prebuilt image (recommended)
Use the published image from GitHub Container Registry (GHCR).
apptainer build llm-sc-curator.sif docker://ghcr.io/kenflab/llm-sc-curator:official
-
Option B: a .sif from the Docker image (development)
docker compose -f docker/docker-compose.yml build apptainer build llm-sc-curator.sif docker-daemon://llm-sc-curator:official
Run Jupyter (either image):
apptainer exec --cleanenv \
--bind "$PWD":/work \
llm-sc-curator.sif \
bash -lc 'jupyter lab --ip=0.0.0.0 --port=8888 --no-browser
🔒 Privacy
We respect the sensitivity of clinical and biological data. LLM-scCurator is architected to ensure that raw expression matrices and cell-level metadata never leave your local environment.
- Local execution: All heavy lifting—preprocessing, confounding gene removal, and feature ranking—occurs locally on your machine.
- Minimal transmission: When interfacing with external LLM APIs, the system transmits only anonymized, cluster-level marker lists (e.g., top 50 ranked gene symbols) and basic tissue context.
- User control: You retain control over any additional background information (e.g., disease state, treatment conditions, and platform) provided via custom prompts. Please review your institution’s data policy and the LLM provider’s terms before sending any information to external LLM APIs.
⚡ Quick Start
🐍 For Python / Scanpy Users
- Set your API key (simplest: paste in the notebook)
import scanpy as sc
from llm_sc_curator import LLMscCurator
GEMINI_API_KEY = "PASTE_YOUR_KEY_HERE"
# OPENAI_API_KEY = "PASTE_YOUR_KEY_HERE" # optional
# Load your data
adata = sc.read_h5ad("my_data.h5ad")
# Initialize with your API Key (Google AI Studio)
curator = LLMscCurator(api_key=GEMINI_API_KEY)
curator.set_global_context(adata)
- Run LLM-scCurator
-
Option A: hierarchical discovery mode(iterative coarse-to-fine clustering and labeling)
# Fully automated hierarchical annotation (includes clustering) adata = curator.run_hierarchical_discovery(adata) # Visualize sc.pl.umap(adata, color=['major_type', 'fine_type'])
-
Option B: Annotate your existing clusters (cluster → table/CSV → per-cell labels)
Use this when you already have clusters (e.g., Seuratseurat_clusters,Leiden, etc.) and want to annotate each cluster once, then propagate labels to cells.# v0.1.1+ from llm_sc_curator import ( export_cluster_annotation_table, apply_cluster_map_to_cells, ) cluster_col = "seurat_clusters" # change if needed # 1) Annotate each cluster (once) clusters = sorted(adata.obs[cluster_col].astype(str).unique()) cluster_results = {} genes_by_cluster = {} for cl in clusters: genes = curator.curate_features( adata, group_col=cluster_col, target_group=str(cl), use_statistics=True, ) genes_by_cluster[str(cl)] = genes or [] if genes: cluster_results[str(cl)] = curator.annotate(genes, use_auto_context=True) else: cluster_results[str(cl)] = { "cell_type": "NoGenes", "confidence": "Low", "reasoning": "Curated gene list empty", } # 2) Export a shareable cluster table (CSV/DataFrame) df_cluster = export_cluster_annotation_table( adata, cluster_col=cluster_col, cluster_results=cluster_results, genes_by_cluster=genes_by_cluster, prefix="Curated", ) df_cluster.to_csv("cluster_curated_map.csv", index=False) # 3) Propagate cluster labels to per-cell labels apply_cluster_map_to_cells( adata, cluster_col=cluster_col, df_cluster=df_cluster, label_col="Curated_CellType", new_col="Curated_CellType", )
Notes: > Manuscript results correspond to v0.1.0; later minor releases add user-facing utilities without changing core behavior.
📊 For R / Seurat Users
You can use LLM-scCurator in two ways:
-
Option A (recommended): Export → run in Python We provide a helper script
examples/R/export_to_curator.Rto export your Seurat object seamlessly for processing in Python.source("examples/R/export_to_curator.R") Rscript examples/R/export_to_curator.R \ --in_rds path/to/seurat_object.rds \ --outdir out_seurat \ --cluster_col seurat_clusters
Output:
counts.mtx(raw counts; recommended)features.tsv(gene list)obs.csv(cell metadata; includes seurat_clusters)umap.csv(optional, if available)
Notes:
- The folder will contain: counts.mtx, features.tsv, obs.csv (and umap.csv if available).
- Then continue in the Python/Colab tutorial to run LLM-scCurator and write cluster_curated_map.csv,
- which can be re-imported into Seurat for plotting.
-
Option B: Run from R via reticulate (advanced)
If you prefer to stay in R, you can invoke the Python package via reticulate (Python-in-R). This is more sensitive to Python environment configuration, so we recommend Option A for most users.
- Use the official Docker (Python + R + Jupyter) and follow the step-by-step tutorial notebook: 📓
examples/R/run_llm_sccurator_R_reticulate.ipynb
The notebook includes:- Use LLM-scCurator for robust marker distillation (no API key required)
- 🔑 Optional: Requires an API key for LLM-scCurator annotation .
- Use the official Docker (Python + R + Jupyter) and follow the step-by-step tutorial notebook: 📓
📄 Manuscript reproduction
For manuscript-facing verification (benchmarks, figures, and Source Data), use the versioned assets under paper/. See paper/README.md for the primary instructions.
Notes:
- Figures are supported by exported Source Data in
paper/source_data/(seepaper/FIGURE_MAP.csvfor panel → file mapping).- Re-running LLM/API calls or external reference annotators is optional; LLM API outputs may vary across runs even with temperature=0.
- For transparency, we include read-only provenance notebooks with example run logs in
paper/notebooks/
📓 Colab notebooks
-
Python / Scanpy quickstart (recommended: colab_quickstart.ipynb)
-
☝️ Runs end-to-end on a public Scanpy dataset (no API key required by default).- 🔑 Optional: If an API key is provided (replace
GEMINI_API_KEY = "YOUR_KEY_HERE"), the notebook can also run LLM-scCurator automatic hierarchical cell annotation.
- 🔑 Optional: If an API key is provided (replace
-
OpenAI quickstart (OpenAI backend: colab_quickstart_openai.ipynb)
-
☝️ Same workflow as the Python / Scanpy quickstart, but configured for the OpenAI backend.- 🔑 Optional: If an API key is provided (replace
OPENAI_API_KEY= "YOUR_KEY_HERE"), the notebook can also run LLM-scCurator automatic hierarchical cell annotation.OPENAI_API_KEYrequires OpenAI API billing (paid API credits).
- 🔑 Optional: If an API key is provided (replace
-
-
R / Seurat quickstart (export → Python LLM-scCurator → back to Seurat: colab_quickstart_R.ipynb)
☝️ Runs a minimal Seurat workflow in R, exports a Seurat object to an AnnData-ready folder, runs LLM-scCurator in Python, then re-imports labels into Seurat for visualization and marker sanity checks.- 🔑 Optional: Requires an API key for LLM-scCurator annotation (same setup as above).
- Recommended for Seurat users who want to keep Seurat clustering/UMAP but use LLM-scCurator for robust marker distillation and annotation.
🔑 Backends (LLM API keys) Setup
Set your provider API key as an environment variable:
GEMINI_API_KEYfor Google GeminiOPENAI_API_KEYfor OpenAI API
See each provider’s documentation for how to obtain an API key and for current usage policies.
-
Option A (Gemini steps):
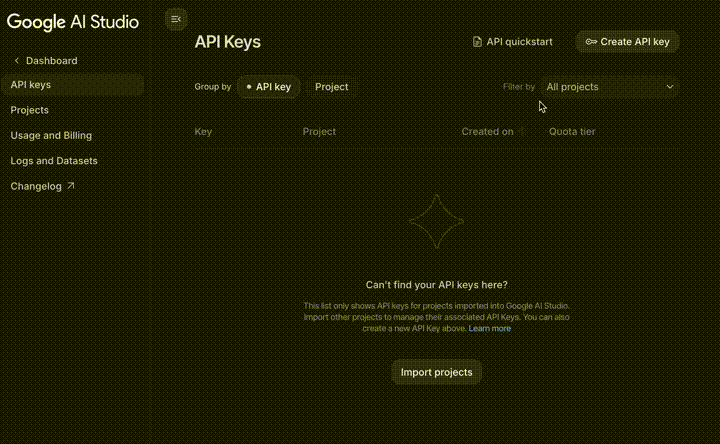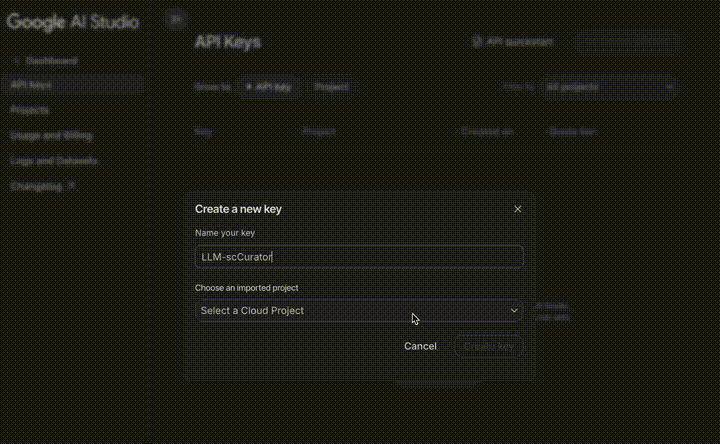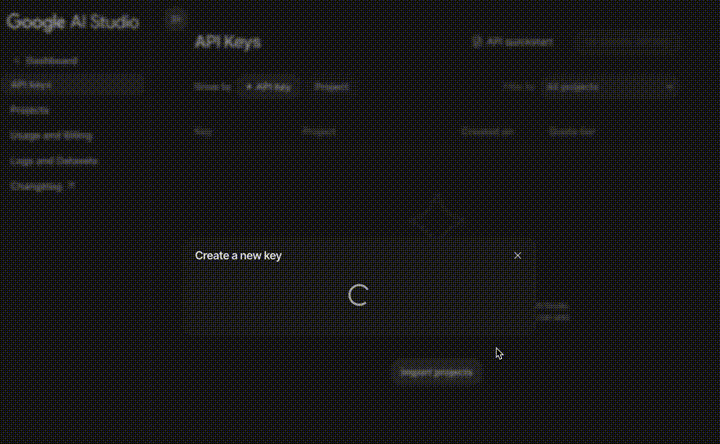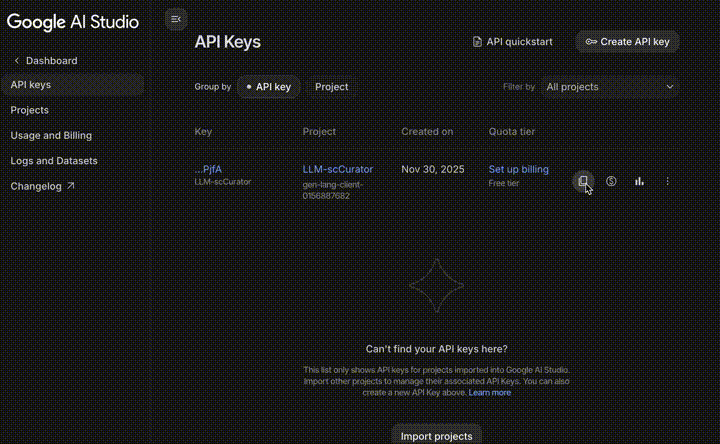
- Go to Google AI Studio.
- Log in with your Google Account.
- Click Get API key (top-left) $\rightarrow$ Create API key.
- Copy the key and use it in your code.
-
Option B (OpenAI steps):
- Go to OpenAI Platform.
- Log in with your OpenAI Account.
- Click Create new secret key $\rightarrow$ Create secret key.
- Copy the key and use it in your code.
Notes:
Google Gemini can be used within its free-tier limits.
OpenAI API usage requires enabling billing (paid API credits); ChatGPT subscriptions (e.g. Plus) do NOT include API usage.
Citation
- bioRxiv preprint: 10.64898/2025.12.28.696778
- Zenodo archive (v0.1.0): 10.5281/zenodo.17970494
- GitHub release tag: v0.1.0
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file llm_sc_curator-0.1.1.tar.gz.
File metadata
- Download URL: llm_sc_curator-0.1.1.tar.gz
- Upload date:
- Size: 55.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
47b9c18a43c4c81a3f72729af43cc853238fe9c23377ccec0376cde72506f028
|
|
| MD5 |
90b2a8958a720586077eb8077b2d2c34
|
|
| BLAKE2b-256 |
d5852366502c546fd3ead9292bbff9d936fdcebe05460ea019e60039897b80be
|
Provenance
The following attestation bundles were made for llm_sc_curator-0.1.1.tar.gz:
Publisher:
pypi-release.yml on kenflab/LLM-scCurator
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
llm_sc_curator-0.1.1.tar.gz -
Subject digest:
47b9c18a43c4c81a3f72729af43cc853238fe9c23377ccec0376cde72506f028 - Sigstore transparency entry: 786318201
- Sigstore integration time:
-
Permalink:
kenflab/LLM-scCurator@7ba06643c4a4a13ab4825e46dc2448076efe50b3 -
Branch / Tag:
refs/tags/v.0.1.1 - Owner: https://github.com/kenflab
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
pypi-release.yml@7ba06643c4a4a13ab4825e46dc2448076efe50b3 -
Trigger Event:
push
-
Statement type:
File details
Details for the file llm_sc_curator-0.1.1-py3-none-any.whl.
File metadata
- Download URL: llm_sc_curator-0.1.1-py3-none-any.whl
- Upload date:
- Size: 57.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8cc934579b7c3c5e1748ae8b30a7ed9be353ec350d569322f4c01bdb38989e31
|
|
| MD5 |
c48ea541ac411c893edf50b62cdf5c12
|
|
| BLAKE2b-256 |
3831b9fe428ea0720df3a467e60811d8dfcd544d514736c6db1b67638525b9f5
|
Provenance
The following attestation bundles were made for llm_sc_curator-0.1.1-py3-none-any.whl:
Publisher:
pypi-release.yml on kenflab/LLM-scCurator
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
llm_sc_curator-0.1.1-py3-none-any.whl -
Subject digest:
8cc934579b7c3c5e1748ae8b30a7ed9be353ec350d569322f4c01bdb38989e31 - Sigstore transparency entry: 786318211
- Sigstore integration time:
-
Permalink:
kenflab/LLM-scCurator@7ba06643c4a4a13ab4825e46dc2448076efe50b3 -
Branch / Tag:
refs/tags/v.0.1.1 - Owner: https://github.com/kenflab
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
pypi-release.yml@7ba06643c4a4a13ab4825e46dc2448076efe50b3 -
Trigger Event:
push
-
Statement type: