Performs large-scale CMF on flexible data layouts.
Project description
Large-scale Collective Matrix Factorization (lsCMF)
This is a package implementing the data integration methodology described in "Large-scale Data Integration using Matrix Denoising and Geometric Factor Matching" (Held, 2024, arXiv:2405.10036 [stat.ME]).
Install development version
To install the development version of the package run
pip install git+https://github.com/cyianor/lscmf.git
Install stable version
To install the stable version of the package run
pip install lscmf
Usage example
A simple usage example is shown below:
import lscmf
from numpy.random import default_rng
# Control randomness
rng = default_rng(42)
# Simulate some data
# - `viewdims`: Dimensions of each view
# - `factor_scales`: The strength/singular value of each factor.
# The diagonal of the D matrices in the paper.
# - `snr`: Signal-to-noise ratio of the noise added to each true signal
#
# The function below generates orthogonal matrices V_i and uses the
# supplied D_ij to form signal matrices V_i D_ij V_j^T. Noise with
# residual variance controlled by the signal-to-noise ratio is added.
xs_sim = lscmf.simulate(
viewdims={0: 500, 1: 250, 2: 250},
factor_scales={
(0, 1): [3.0, 2.5, 2.0, 0.0, 0.0],
(0, 2): [2.8, 0.0, 0.0, 2.0, 0.0],
(1, 2): [1.2, 0.0, 5.0, 0.0, 1.1],
},
snr=1.0,
rng=rng,
)
# `xs_sim` is a dictionary containing
# - "xs_truth", the true signal matrices
# - "xs", the noisy data
# - "vs", the simulated orthogonal factors
# Create the lscmf object and fit the model to data
est = lscmf.LargeScaleCMF().fit(xs_sim["xs"])
Estimates of model parameters are then contained in the LargeScaleCMF object.
The estimated singular values can be accessed as shown below.
est.ds_
{(0,
1): array([-2.98687823, 0. , 1.96015864, 0. , 2.47498787]),
(0,
2): array([-2.78861131, 1.96604697, 0. , 0. , 0. ]),
(1, 2): array([1.13272996, 0. , 4.98917765, 1.00988163, 0. ])}
The estimated factors can be accessed as follows, e.g., for view 0 the first factor is
import matplotlib.pyplot as plt
cos_angle = (est.vs_[1][:, 2] * xs_sim["vs"][1][:, 4]).sum()
print(f"Scalar product between estimated and true factor: {cos_angle:.2f}")
fig = plt.figure(figsize=(8, 3), dpi=100)
ax = fig.add_subplot(111)
# Negate integrated factor since it was estimated as -v instead of v
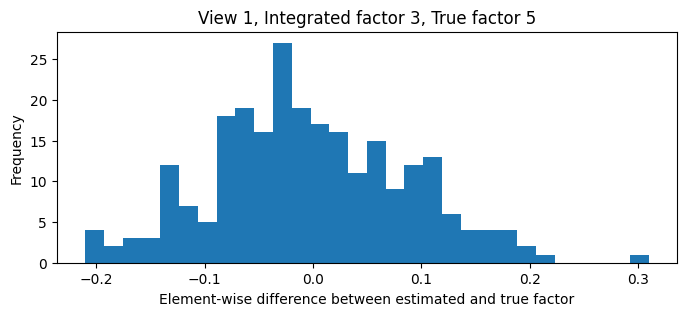
ax.hist((-est.vs_[1][:, 2]) - xs_sim["vs"][1][:, 4], bins=30)
ax.set_xlabel("Element-wise difference between estimated and true factor")
ax.set_ylabel("Frequency");
ax.set_title("View 1, Integrated factor 3, True factor 5");
Scalar product between estimated and true factor: 0.00
A raw graph-based interface exists as well.
# Create a view graph to hold the data layout
G = lscmf.ViewGraph()
# Add data
# - `names` need to be provided as an iterable.
# These are in general arbitrary, however, in case of
# repeated layers, each layer requires a unique name.
# - `xs` is an iterable to the input data in the same order as
# `names`
# - `viewrels` is an iterable containing tuples describing the
# relationships between views contained in data matrices.
G.add_data_from(["x01", "x02", "x12"], xs_sim["xs"].values(), [(0, 1), (0, 2), (1, 2)])
# Once data is added to the view graph, joint matrices for each
# view need to be formed and denoising needs to be performed.
# Different types of shrinkers can be used for denoising and they
# depend on the type of loss assumed for reconstruction of the
# signal. See Gavish and Donoho (2017) for details.
# In the paper, Frobenius loss is assumed, and therefore the resulting
# `FrobeniusShrinker` is used here.
lscmf.precompute(G, lscmf.FrobeniusShrinker)
# Finally, matching of factors for each view and merging of the
# factor match graphs is performed. This function returns
# the final merged factor match graph.
H = lscmf.match_factors(G)
Matches in the factor match graph can be investigated. MatchNodes contain a data_edge, which is a MultiEdge(name, viewrel) corresponding to an input matrix during view graph construction, and a factor which is the factor in the data_edge. Keys of the dictionary are integrated factors.
In the example below, MatchNode(data_edge=MultiEdge(x01, (0, 1)), factor=0) is the first factor in data matrix $X_{01}$ which is being associated with integrated factor 0.
The numbering of integrated factors is arbitrary and may be non-consecutive.
H.graph
defaultdict(set,
{0: {MatchNode(data_edge=MultiEdge(x01, (0, 1)), factor=0),
MatchNode(data_edge=MultiEdge(x02, (0, 2)), factor=0),
MatchNode(data_edge=MultiEdge(x12, (1, 2)), factor=1)},
2: {MatchNode(data_edge=MultiEdge(x02, (0, 2)), factor=1)},
1: {MatchNode(data_edge=MultiEdge(x01, (0, 1)), factor=2),
MatchNode(data_edge=MultiEdge(x12, (1, 2)), factor=0)},
3: {MatchNode(data_edge=MultiEdge(x12, (1, 2)), factor=2)},
4: {MatchNode(data_edge=MultiEdge(x01, (0, 1)), factor=1)}})
Reconstruction of $D_{ij}$ and $V_i$ is performed in LargeScaleCMF.fit() and it is recommended to use that interface unless the graph interface is required for a specific reason.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file lscmf-0.2.1.post1.tar.gz.
File metadata
- Download URL: lscmf-0.2.1.post1.tar.gz
- Upload date:
- Size: 58.8 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: python-httpx/0.28.1
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5ee2b3415548ccf4790d3948da26084e76945d4753ddb4271ab3488b67df3442
|
|
| MD5 |
74d5b755af86153db1af7f383056075e
|
|
| BLAKE2b-256 |
fe99fa81a7ed2e7d9817129aadb6e306ad7a39c20a2a8e83e8962a3f7c5633e1
|
File details
Details for the file lscmf-0.2.1.post1-py3-none-any.whl.
File metadata
- Download URL: lscmf-0.2.1.post1-py3-none-any.whl
- Upload date:
- Size: 25.9 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: python-httpx/0.28.1
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
df32a966db05b5722a1f6708b4f745b536d94db9840d4cc84041765e4825c3e6
|
|
| MD5 |
b21c015b2a4c3998c71021666a8e8ac3
|
|
| BLAKE2b-256 |
7b773670e220f1d6a3debee9b344f74c848c66fa7131a2173219edc76348b8ed
|