3D Neural Network for Lung Cancer Risk Prediction on CT Volumes
Project description
# 3D Neural Network for Lung Cancer Risk Prediction on CT Volumes
## Overview
This repository contains my implementation of the “full-volume” model from the paper:
[End-to-end lung cancer screening with three-dimensional deep learning on low-dose chest computed tomography.](https://doi.org/10.1038/s41591-019-0447-x)<br/> Ardila, D., Kiraly, A.P., Bharadwaj, S. et al. Nat Med 25, 954–961 (2019).
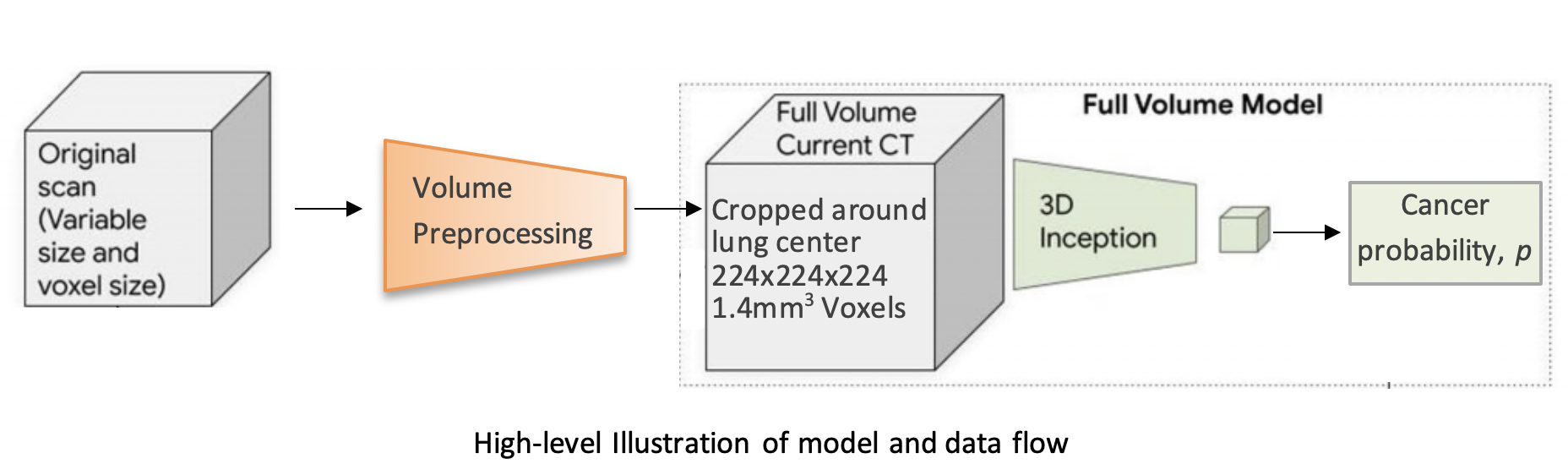
The model uses a three-dimensional (3D) CNN to perform end-to-end analysis of whole-CT volumes, using LDCT volumes with pathology-confirmed cancer as training data. The CNN architecture is an Inflated 3D ConvNet (I3D) ([Carreira and Zisserman](http://openaccess.thecvf.com/content_cvpr_2017/html/Carreira_Quo_Vadis_Action_CVPR_2017_paper.html)).
### Data
We use the NLST dataset which contains chest LDCT volumes with pathology-confirmed cancer evaluations. For description and access to the dataset refer to the [NCI website](https://biometry.nci.nih.gov/cdas/learn/nlst/images/).
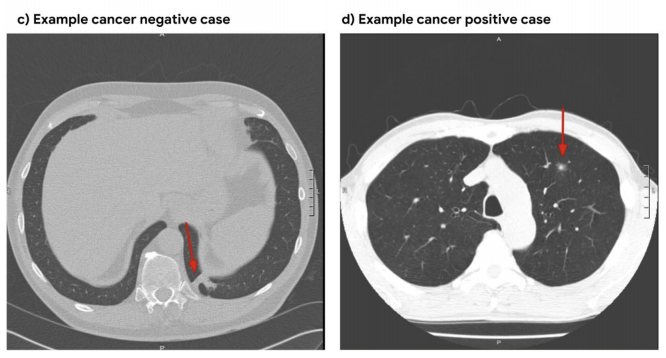
Sample data comes from the [Lung Image Database Consortium image collection (LIDC-IDRI)<sup>1</sup>](https://wiki.cancerimagingarchive.net/display/Public/LIDC-IDRI).
## Running the code
### Setup
`bash git clone https://github.com/danielkorat/Lung-Cancer-Risk-Prediction cd Lung-Cancer-Risk-Prediction pip install -U pip pip install -r requirements.txt `
The main.py module contains training (fine-tuning) and inference procedures. The inputs are preprocessed CT volumes, as produced by preprocess.py. For usage example, refer to the arguments’ description and default values in the bottom of main.py.
For an example of running out-of-the box inference: [[Notebook](https://github.com/danielkorat/Lung-Cancer-Risk-Prediction/blob/master/notebooks/inference.ipynb)] [[Colab](https://colab.research.google.com/drive/1nWFFiFI43W7aClax0fjR3OEepTAW5Opw?usp=sharing)]
### Data Preprocessing
The main.py module operates only on preprocessed volumes, produced by preprocess.py. Each CT volume in NLST is a folder of DICOM files (one file per slice). The preprocess.py module accepts a directory path/to/data containing multiple CT volumes, performs several preprocessing steps, and writes each volume as an .npz file in path/to/data_preprocssed. The preprocessing steps include methods from [this](https://www.kaggle.com/gzuidhof/full-preprocessing-tutorial/notebook) tutorial and include:
Resampling to a 1.5mm voxel size (slow)
Coarse lung segmentation – used to compute lung center for alignment and reduction of problem space
To save storage space, the following preprocessing steps are performed online (during training/inference):
Windowing – clip pixel values to focus on lung volume
RGB normalization
### Provided checkpoint
By default, our fine-tuned model checkpoint is downloaded in main.py and the model is then initialized with its weights. Due to limited storage and compute time, we trained on a small subset of NLST containing 1,045 volumes (34% positive). Nevertheless, we still achieved a very high AUC score of 0.892 on a validation set of 448 volumes. This is comparable to the original paper’s AUC for the full-volume model (see the paper’s supplemtary material), trained on 47,974 volumes (1.34% positive).
To train this model we first initialized by bootstrapping the filters from the [ImageNet pre-trained 2D Inception-v1 model]((http://download.tensorflow.org/models/inception_v1_2016_08_28.tar.gz)) into 3D, as described in the I3D paper. It was then fine-tuned on the preprocessed CT volumes to predict cancer within 1 year (binary classification). Each of these volumes was a large region cropped around the center of the bounding box, as determined by lung segmentation in the preprocessing step.
For the training setup, we set the dropout keep_prob to 0.7, and trained in mini-batches of size of 2 (due to limited GPU memory). We used tf.train.AdamOptimizer with a small learning rate of 5e-5, (due to the small batch size) and stopped the training before overfitting started around epoch 37. The focal loss function from the paper is provided in the code, but we did not experience improved results using it, compared to cross-entropy loss which was used instead. The likely reason is that our dataset was more balanced than the original paper’s.
The follwoing plots show loss, AUC, and accuracy progression during training, along with ROC curves for selected epochs:
<img src=”https://raw.githubusercontent.com/danielkorat/Lung-Cancer-Risk-Prediction/master/figures/loss.png” width=”786” height=”420”> <img src=”https://raw.githubusercontent.com/danielkorat/Lung-Cancer-Risk-Prediction/master/figures/auc_and_accuracy.png” width=”786” height=”420”>
<img src=”https://raw.githubusercontent.com/danielkorat/Lung-Cancer-Risk-Prediction/master/figures/epoch_10.png” width=”270” height=”270”><img src=”https://raw.githubusercontent.com/danielkorat/Lung-Cancer-Risk-Prediction/master/figures/epoch_20.png” width=”270” height=”270”><img src=”https://raw.githubusercontent.com/danielkorat/Lung-Cancer-Risk-Prediction/master/figures/epoch_32.png” width=”270” height=”270”>
### Acknowledgments
The author thanks the National Cancer Institute for access to NCI’s data collected by the National Screening Trial (NLST). The statements contained herein are solely those of the author and do not represent or imply concurrence or endorsement by NCI.
<sup>1</sup> Armato III, SG; McLennan, G; Bidaut, L; McNitt-Gray, MF; Meyer, CR; Reeves, AP; Zhao, B; Aberle, DR; Henschke, CI; Hoffman, Eric A; Kazerooni, EA; MacMahon, H; van Beek, EJR; Yankelevitz, D; Biancardi, AM; Bland, PH; Brown, MS; Engelmann, RM; Laderach, GE; Max, D; Pais, RC; Qing, DPY; Roberts, RY; Smith, AR; Starkey, A; Batra, P; Caligiuri, P; Farooqi, Ali; Gladish, GW; Jude, CM; Munden, RF; Petkovska, I; Quint, LE; Schwartz, LH; Sundaram, B; Dodd, LE; Fenimore, C; Gur, D; Petrick, N; Freymann, J; Kirby, J; Hughes, B; Casteele, AV; Gupte, S; Sallam, M; Heath, MD; Kuhn, MH; Dharaiya, E; Burns, R; Fryd, DS; Salganicoff, M; Anand, V; Shreter, U; Vastagh, S; Croft, BY; Clarke, LP. (2015). Data From LIDC-IDRI. The Cancer Imaging Archive. http://doi.org/10.7937/K9/TCIA.2015.LO9QL9SX
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file lungs-0.1.0.tar.gz.
File metadata
- Download URL: lungs-0.1.0.tar.gz
- Upload date:
- Size: 21.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.2.0 pkginfo/1.5.0.1 requests/2.23.0 setuptools/47.3.1 requests-toolbelt/0.9.1 tqdm/4.43.0 CPython/3.6.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
61c123c11cdd87f4be314ae203fd077e317016797463cf6b4e2d83236b1197f7
|
|
| MD5 |
f85a5c8edb119e629f30cc5e7df2aa0f
|
|
| BLAKE2b-256 |
ad478a25f9337ff868de95234d63be4b0bcf8b485d5e44ce03028afd07a7b0b6
|
File details
Details for the file lungs-0.1.0-py3-none-any.whl.
File metadata
- Download URL: lungs-0.1.0-py3-none-any.whl
- Upload date:
- Size: 33.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.2.0 pkginfo/1.5.0.1 requests/2.23.0 setuptools/47.3.1 requests-toolbelt/0.9.1 tqdm/4.43.0 CPython/3.6.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d77f3256e2a588e4530ace5b7888ee5fc9bd6ae71e1aaccaf9bd705fb1963722
|
|
| MD5 |
bacc742940af7e3a91b657a5aea5d9e0
|
|
| BLAKE2b-256 |
1ceccec96e277ce3705a3a133428b76e87d188c8819ee34dd65f83c81048f42e
|











