Containment analysis for (periodic) point clouds.
Project description

MDVContainment v2.0.0
Robust and fast containment charaterization for (periodic) point clouds and voxel masks for binary density labeling.
Molecular Dynamics Voxel Containment yields a robust characterization of the inside and outside hierarchy for point clouds in periodic spaces of dimensionality three or lower (e.g. R^3/Z^3).
Using an MDAnalysis atomgroup and resolution, a density grid is created by pbc aware binning of the particle positions. This density grid is segmented using connected components, and graph logic is utilized to solve the topological identification of containment (insides and outsides). The final output is a set of Directed Acyclic Graphs (DAGs) running from the largest container to the smallest (from outside inwards in graph space). This containment logic can then be used to analyse or manipulate the systems.
Any complex configuration of (non)periodic segments is supported by this algorithm in a fast, robust, unambiguous, deterministic and rot+trans invarient (up to voxel discretization) manner.
[!NOTE] MDVContainment is undergoing a functional overhaul, supporting integer labeling (e.g. MDVLeafletSegmentation labeling)! When the code is ready to be tested a branch will be created called
integer_containmentwhich will probably become the new main branch over time.
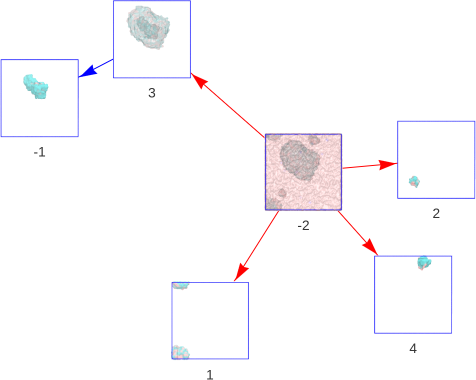
Figure 1 | Containment hierarchy in self-assembled acyl chain bicelles. The main solvent (segment -2) is the most outer segment in this system. It containes three non-periodic segments (1, 2, 3), where segment 1 is split over the periodic boundary. Segment 3 contains a bubble of inner solvent (segment -1).
$~$
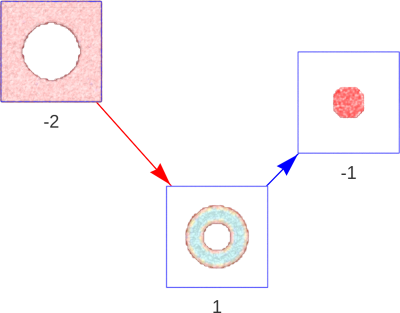
Figure 2 | A periodic hollow cylinder in solution. The cylinder (segment 1) splits the solution into two segments (segment -2, -1), the solid cylinder inside the hollow cylinder (segment -1), and all of the space outside of the cylinder (segment -2). Both cylindrical segments (1 and -1) are said to be contained by the solvent segment (-2), although only the hollow cylinder (1) is a child of the solvent segment (-2).
$~$
Citation and License
@article {Bruininks2025.08.06.668936,
author = {Bruininks, Bart M. H. and Vattulainen, Ilpo},
title = {Classification of containment hierarchy for point clouds in periodic space},
elocation-id = {2025.08.06.668936},
year = {2025},
doi = {10.1101/2025.08.06.668936},
publisher = {Cold Spring Harbor Laboratory},
URL = {https://www.biorxiv.org/content/early/2025/08/09/2025.08.06.668936},
eprint = {https://www.biorxiv.org/content/early/2025/08/09/2025.08.06.668936.full.pdf},
journal = {bioRxiv}
}
Please cite this work if you use it for scientific publications. It helps me to continue to work on this kind of software, thanks! On that note, if you would offer me a job to work on this, I would take your offer very seriously.
MDVContainment is available under the Apache-2.0 license.
Requirements
MDVContainment has been tested to work with python >= 3.12.
Installation
Using pypi
Install v2.0.0 in the current python environment:
pip install mdvcontainment==v2.0.0
Using git
Direct install from the github main branch into the current python environment:
pip install git+https://github.com/BartBruininks/mdvcontainment
Create a folder in a custom location using git clone:
git clone git@github.com:BartBruininks/mdvcontainment.git
cd mdvcontainment
pip install .
[!IMPORTANT] If you need any help with MDVContainment or have ideas for future functionalities, please raise an issue!
Minimal example CG Martini
Calculate the containment
Input
# Import the required libraries
import MDAnalysis as mda
from mdvcontainment import Containment
# Import the structure file
path = 'your_structure.pdb' # Or any MDA supported structures file
universe = mda.Universe(path)
selection_string = 'not resname W WF ION' # Useful for CG Martini
selection = universe.select_atoms(selection_string)
# Run the containment analysis
containment = Containment(selection, resolution=0.5, closing=True)
# Show the containment graph with voxel counts
print(containment)
[!NOTE] For atomistic structures use
closing=Falseat 0.5 nm resolution. Take a look at closing (link to wikipedia) to learn more about what it does.
Output
Containment Graph with 3 components (component: nm^3: rank):
└── [-2: 7350: 3]
└── [1: 477: 0]
└── [-1: 65: 0]
Plot the compositions
Input
# Plot the compositions
composition, fig, axs = cl.analyze_composition(containment, mode='names') # or 'resnames' / 'molar'
Output
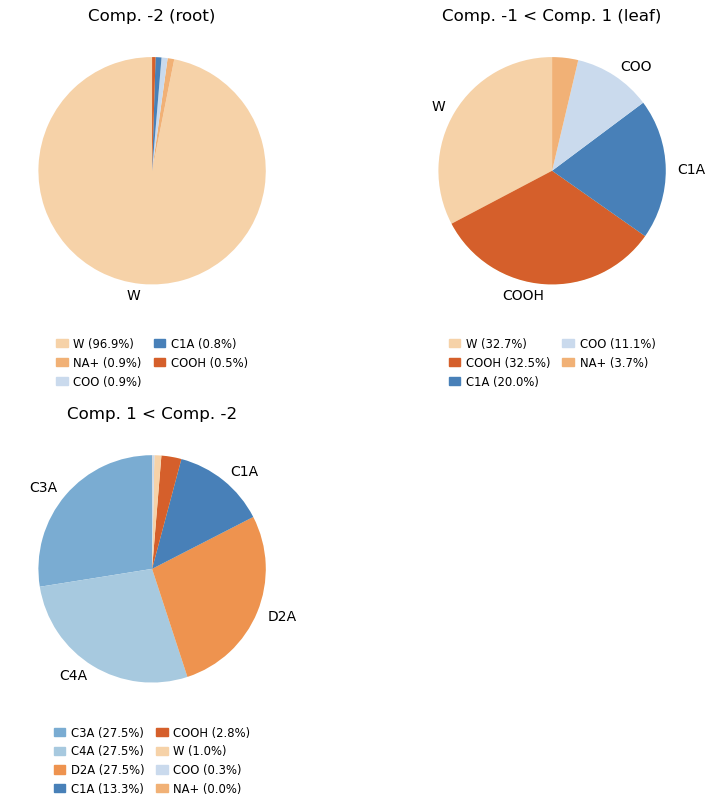
Extensive examples
For worked examples in jupyter notebooks, take a look at the examples/notebooks folder. Some example structure files are added under examples/structures.
I still need to add this tutorial to this repo, but for now a very detailed and up to date tutorial can be found on the cgmartini website here.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file mdvcontainment-2.0.0.post1.tar.gz.
File metadata
- Download URL: mdvcontainment-2.0.0.post1.tar.gz
- Upload date:
- Size: 371.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8f5223dc5c6f38ab373ca0b1c36445596823a224cbba0a966253127bf0e970bc
|
|
| MD5 |
4f59ab7385ee3bc9de509179be4575e5
|
|
| BLAKE2b-256 |
b85e89146e0485ebecb2a4e5ce37cdfc9da343288e7c226bca07e9f8c530a032
|
File details
Details for the file mdvcontainment-2.0.0.post1-cp314-cp314-macosx_11_0_arm64.whl.
File metadata
- Download URL: mdvcontainment-2.0.0.post1-cp314-cp314-macosx_11_0_arm64.whl
- Upload date:
- Size: 154.1 kB
- Tags: CPython 3.14, macOS 11.0+ ARM64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1556b65b1f1af49543ee00a86195deb661c505393cfb73d9ed6a69251ec5e7c6
|
|
| MD5 |
9a97cbc028edc2adcbf2851043ec02b2
|
|
| BLAKE2b-256 |
92fffddadeb39683317e4851f244a0f3e7689383c778e4524420f8e0f5a64caf
|











