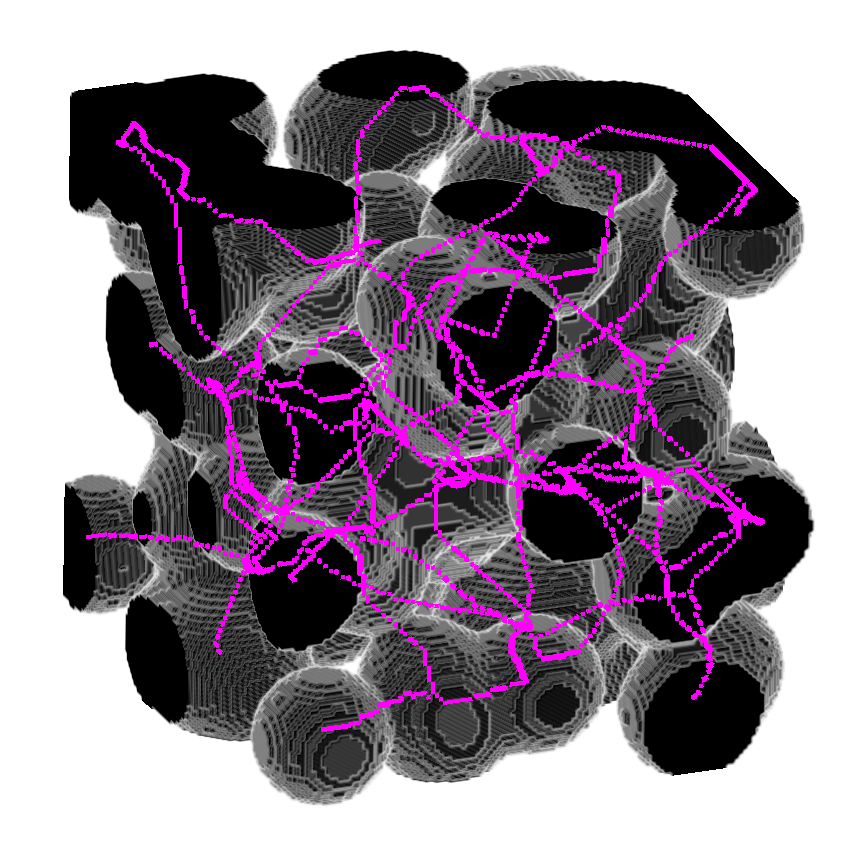
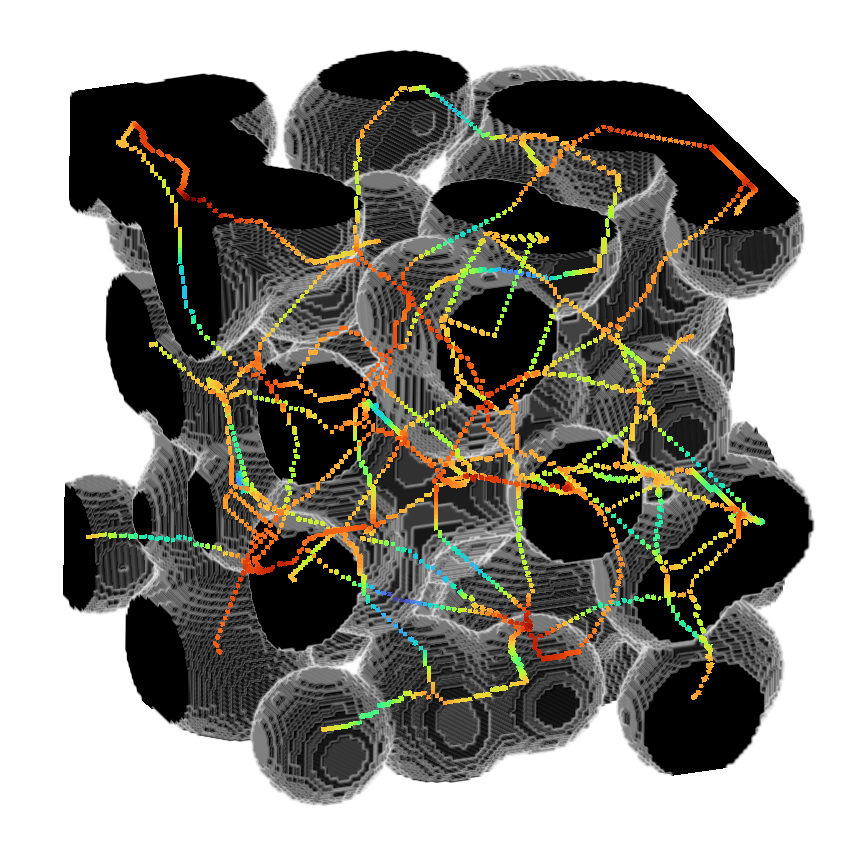
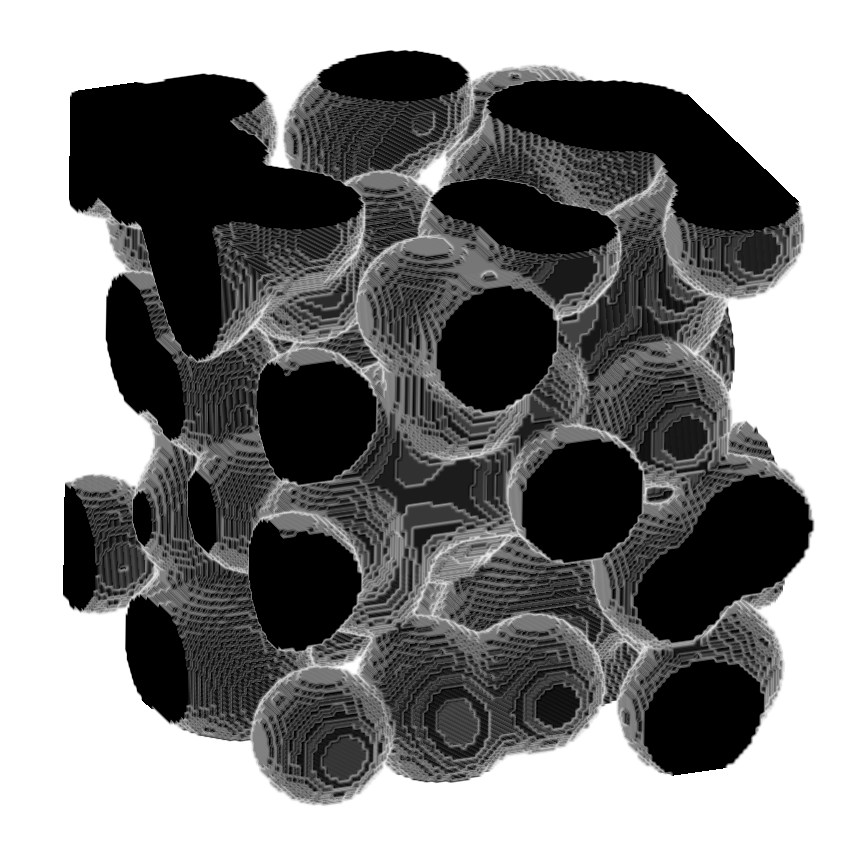
Module to perform 3D skeletonization using Medial Axis Transform
Project description
medialaxis3d
This package extends the scikit-image function medial_axis to the 3D case.
Install
pip install medialaxis3d
Dependencies
Automatically installed with pip:
numpyscipycython
Optional only for visualization
napari
Documentation
WIP
Quickstart
Use it without returning the medial distance.
>>> import numpy as np
>>> import skimage as ski
>>> import medialaxis3d
>>> import napari
>>> rng = np.random.default_rng(1278)
>>> image = ski.data.binary_blobs(length = 128,
>>> blob_size_fraction = 0.2,
>>> n_dim = 3,
>>> volume_fraction = 0.6,
>>> rng = rng)
>>> skeleton = medialaxis3d.medial_axis_3d(image,
>>> return_distance = False,
>>> size = 8,
>>> rng = rng)
>>> viewer = napari.Viewer()
>>> viewer.add_image(image,
>>> rendering = "attenuated_mip",
>>> attenuation = 0.5,
>>> scale = [1, 1, 1])
>>> viewer.add_image(skeleton,
>>> interpolation3d = "nearest",
>>> colormap = "magenta",
>>> scale = [1, 1, 1])
>>> napari.run()

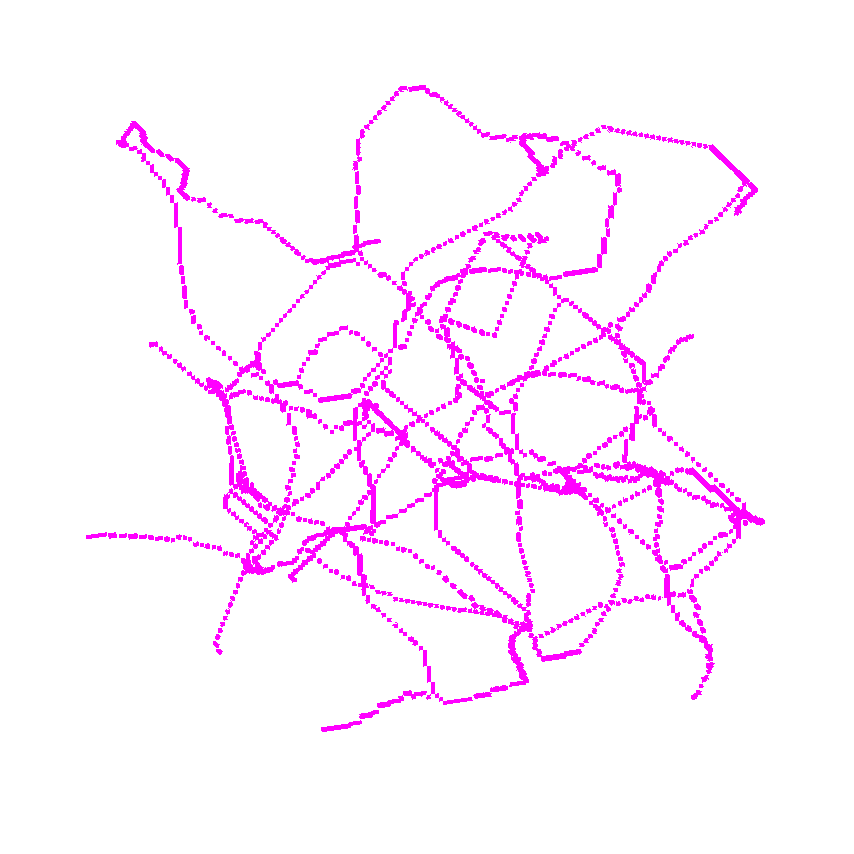
or use it to return the distance as well.
>>> import numpy as np
>>> import skimage as ski
>>> import medialaxis3d
>>> import napari
>>> rng = np.random.default_rng(1278)
>>> image = ski.data.binary_blobs(length = 128,
blob_size_fraction = 0.2,
n_dim = 3,
volume_fraction = 0.6,
rng = rng)
>>> skeleton, distance = medialaxis3d.medial_axis_3d(image,
>>> return_distance = True,
>>> size = 8,
>>> rng = rng)
>>> viewer = napari.Viewer()
>>> viewer.add_image(image,
>>> rendering = "attenuated_mip",
>>> attenuation = 0.5,
>>> scale = [1, 1, 1])
>>> viewer.add_image(skeleton*distance,
>>> interpolation3d = "nearest",
>>> colormap = "turbo",
>>> scale = [1, 1, 1])
>>> napari.run()

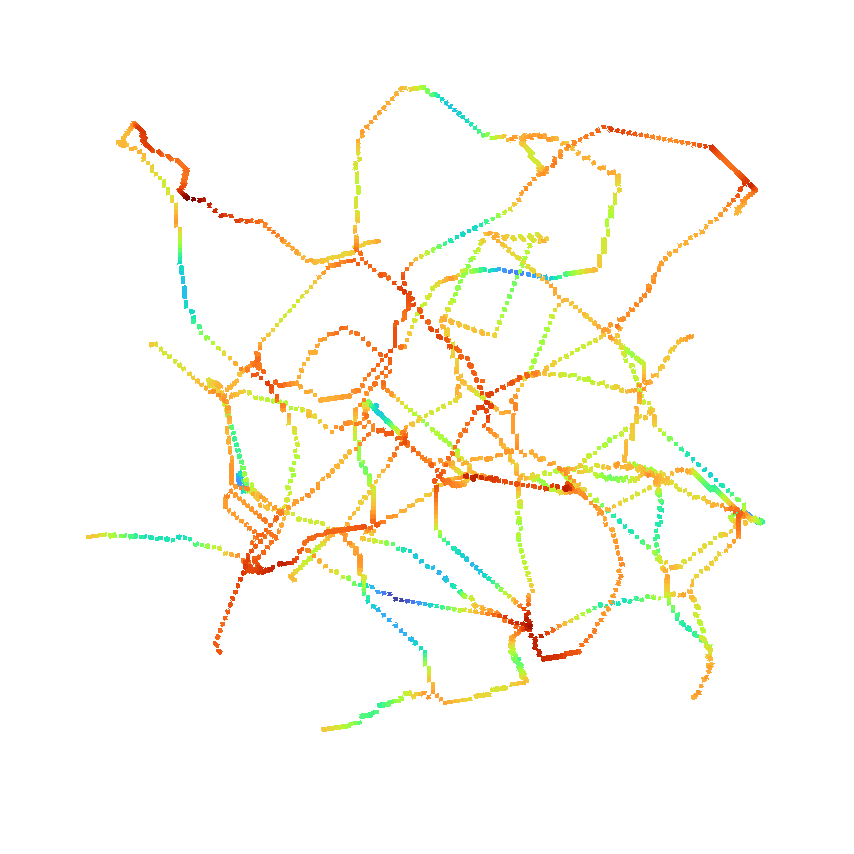
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file medialaxis3d-1.0.3.tar.gz.
File metadata
- Download URL: medialaxis3d-1.0.3.tar.gz
- Upload date:
- Size: 3.0 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1ee35c6711706663c9091bbdd6ab8f7924fe734525e4c036305d70a4b7cf5ec3
|
|
| MD5 |
4ced815ae5c34b01b5f65c278867ccf2
|
|
| BLAKE2b-256 |
306a92d5df10256266ce6c9ebd55d5e3fbc6cfda3e2b3f0e77c0eed94d71e7fd
|
File details
Details for the file medialaxis3d-1.0.3-cp313-cp313-manylinux_2_39_x86_64.whl.
File metadata
- Download URL: medialaxis3d-1.0.3-cp313-cp313-manylinux_2_39_x86_64.whl
- Upload date:
- Size: 3.1 MB
- Tags: CPython 3.13, manylinux: glibc 2.39+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
593a35e7d791fd2ec1f988e1cc1cca04d0bdd15db80f206dc71e11696153caaf
|
|
| MD5 |
5ed8ca76c0f7a74432592ebcd24ef736
|
|
| BLAKE2b-256 |
566e799c5b4bab940ec277d211513bb86eb14d97758bd5d4a3ff84a6b6382b1a
|