Melanoma segmentation and classification
Project description
Melanoma Segmentation and Classification
This package/repository provides a comprehensive solution for melanoma segmentation and classification using deep learning techniques. The core of this project involves training and evaluating convolutional neural networks (CNNs) for accurately identifying and segmenting skin lesions in dermatoscopic images.
Features:
- Segmentation Models: Includes implementations of various U-Net based models like R2U-Net and TransUNet with enhancements such as attention mechanisms and transformer blocks to achieve accurate segmentation of skin lesions.
- Dataset Preparation: Automates the splitting of large dermatoscopic datasets into training, validation, and testing sets using customizable transformations for data augmentation, ensuring robust training.
- Custom Metrics and Evaluation: Implements metrics such as Dice coefficient, Intersection over Union (IoU), accuracy, and recall to measure the model’s performance in segmentation tasks.
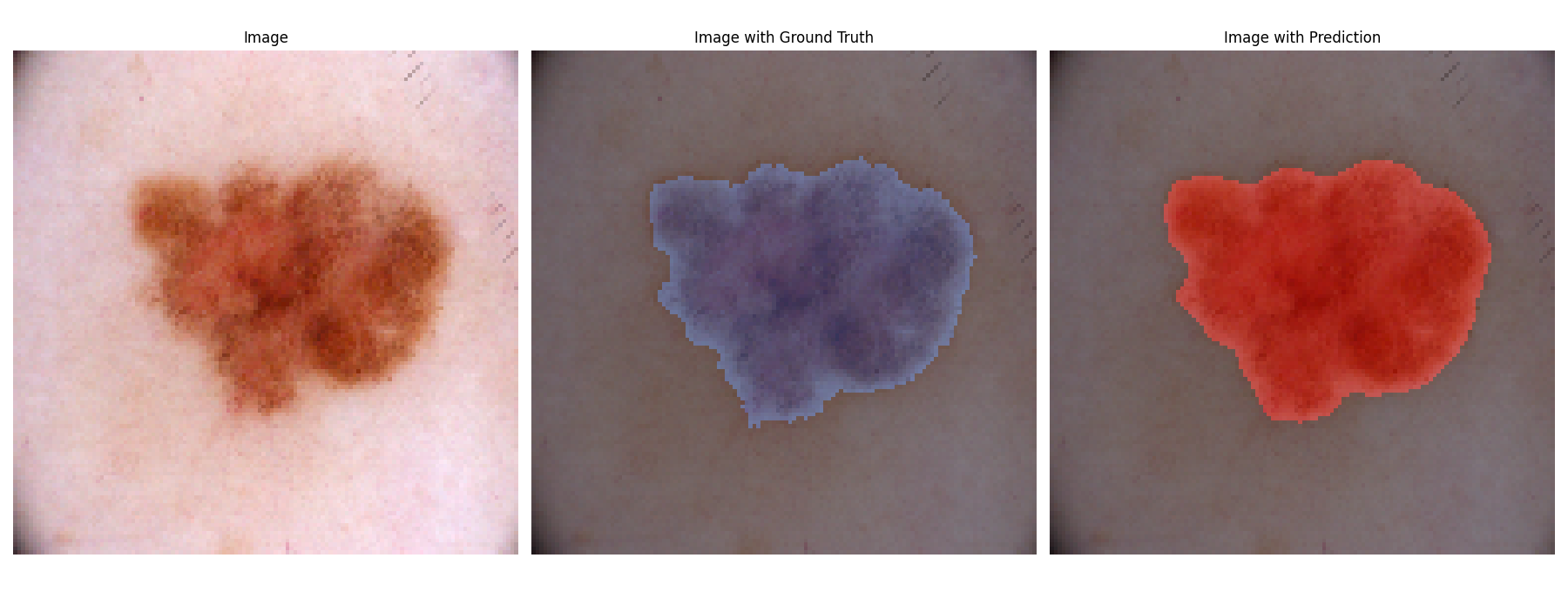
- Visualization Tools: Offers functionality to visualize images alongside ground truth and predicted segmentation masks, making it easier to analyze model performance.
- Comprehensive Data Processing Pipeline: Integrates data loading, preprocessing, model training, and evaluation in a unified pipeline using PyTorch, facilitating reproducibility and scalability.
- Trained_models: Trained models that can be used to compare metrics, there are multiple of them accesible as you can see inside
evaluate.py
Dependencies:
The project uses various dependencies such as PyTorch, torchvision, and albumentations for image augmentation and preprocessing, as well as pre-trained models for enhanced feature extraction. The package also leverages numpy, pandas, and scikit-learn for numerical operations and data handling.
Use Case:
This project is intended for researchers and developers aiming to build or extend segmentation models for medical image analysis, specifically for tasks related to skin lesion identification and segmentation. It is well-suited for experimentation with dermatoscopic datasets and contributes to research efforts in melanoma detection.
Package Link:
References:
- Visualization: Based on code by an-eve from the ISIC 2016 Lesion Segmentation Challenge GitHub link.
- TransUNet Design: Inspired by TESL-Net: A Transformer-Enhanced CNN for Accurate Skin Lesion Segmentation by Shahzaib Iqbal et al. (2024). DOI: arXiv link.
- Data Augmentation: Inspired by the top-ranked solution of the SIIM-ISIC Melanoma Classification Challenge GitHub link.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file melanoma-segmentation-1.4.0.tar.gz.
File metadata
- Download URL: melanoma-segmentation-1.4.0.tar.gz
- Upload date:
- Size: 19.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.10.11
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
ea85a99805044e3b2246f9bba448d58aee5808a51b25e3d7f4d688b74973f9c0
|
|
| MD5 |
1557b8fc4049ceffaa0b40c00eb25f98
|
|
| BLAKE2b-256 |
06108c0a718905f3177af9f8da019a8f101ed49ab2092c9bb9ef0e6878baa3c3
|
File details
Details for the file melanoma_segmentation-1.4.0-py3-none-any.whl.
File metadata
- Download URL: melanoma_segmentation-1.4.0-py3-none-any.whl
- Upload date:
- Size: 25.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.10.11
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
fa0d491a3a4300d8303c779e134183d6bbe48680e7eff677f3ad9abefb7e946a
|
|
| MD5 |
b349f23de90a7f86f0bd6f9e71533947
|
|
| BLAKE2b-256 |
b7465666f51534e4899a28a3fc805d7f80408aad31c0fecff1253992c51d5a29
|