membrane protein localization for cryo-ET
Project description
MemBrain-Stats
Membrain-Stats1 is a Python project developed by the CellArchLab for computing membrane protein statistics in 3D for cryo-electron tomography (cryo-ET). It is part of the larger MemBrain v2 package, which also includes MemBrain-pick for membrane protein detection and MemBrain-seg for membrane segmentation.
The goal of the package is to provide easy-to-use tools to analyze the distribution of membrane proteins in relation to the underlying membrane geometry. To this end, we provide the following functionalities:
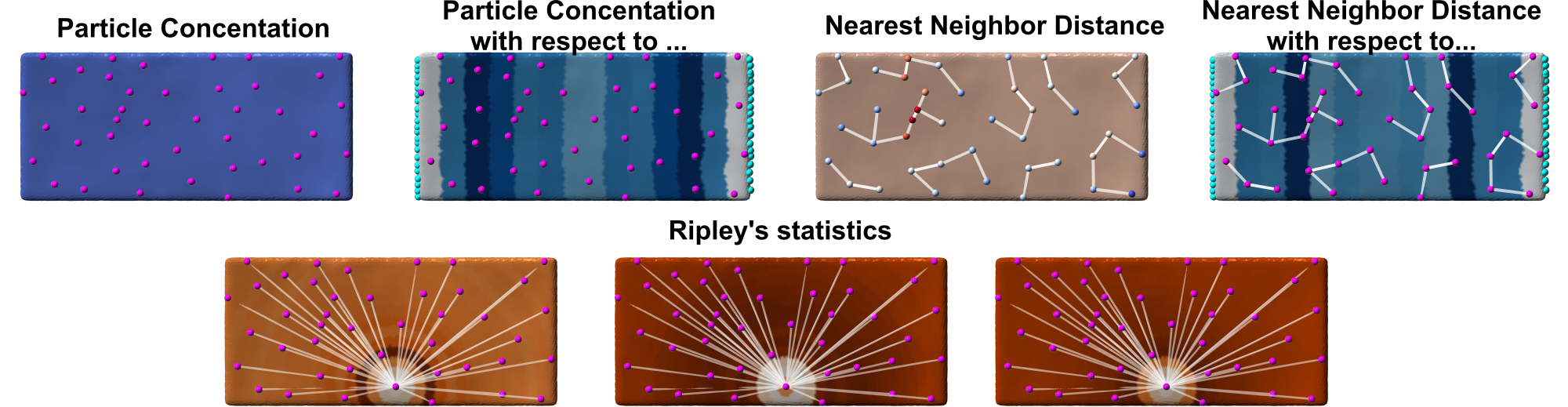
- Protein concentration computation
- Protein concentration with respect to another point class (e.g., membrane edge)
- Protein concentration with respect to a membrane property (e.g., morphometrics property)
- (Geodesic) Nearest Neighbor distances (and orientations of neighbors to each other)
- (Geodesic) Nearest Neighbor distances with respect to another point class (e.g., membrane edge)
- (Geodesic) Ripley's statistics
Publication:
Membrain-stats's functionalities are described in more detail in our preprint [1].
[1] Lamm, L., Zufferey, S., Righetto, R.D., Wietrzynski, W., Yamauchi, K.A., Burt, A., Liu, Y., Zhang, H., Martinez-Sanchez, A., Ziegler, S., Isensee, F., Schnabel, J.A., Engel, B.D., and Peng, T, 2024. MemBrain v2: an end-to-end tool for the analysis of membranes in cryo-electron tomography. bioRxiv, https://doi.org/10.1101/2024.01.05.574336
Usage:
For more details about how to use MemBrain-stats, refer to our User Instructions document.
Example Notebooks:
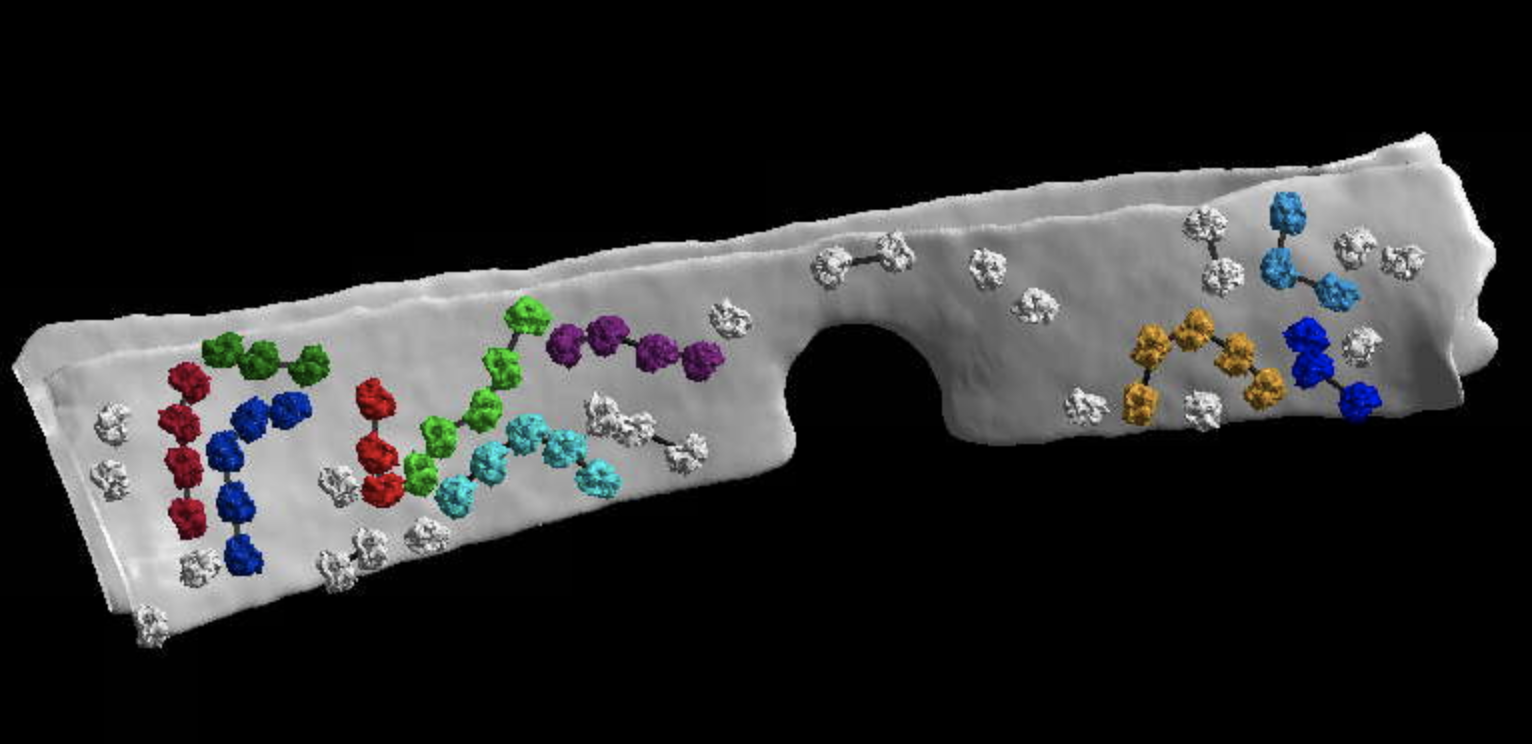
We provide an example jupyter notebook highlighting what can be done with MemBrain-stats here. This is an advanced example, using outputs from template matching and analyzing ribosome chains (this example was also shown in our preprint). We did not wrap this functionality into a single command in MemBrain-stats due to its complexity and need for user input. But if you are interested in analyzing similar data, this notebook can serve as a guide.
An example notebook (Colab tutorial) showcasing the pure functionalities of MemBrain-stats can be found here (
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file membrain_stats-0.0.3.tar.gz.
File metadata
- Download URL: membrain_stats-0.0.3.tar.gz
- Upload date:
- Size: 83.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.19
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1161f178da52b221cf465e2018879a2d891ed972306dbb185ce17c20b7a66d86
|
|
| MD5 |
5554ff11c0f43797b3032f1139cdb462
|
|
| BLAKE2b-256 |
ca76aae731c01508feb294ade474fdbbeb667e61584bf18763bc2a2df43c90f4
|
File details
Details for the file membrain_stats-0.0.3-py3-none-any.whl.
File metadata
- Download URL: membrain_stats-0.0.3-py3-none-any.whl
- Upload date:
- Size: 35.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.19
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
170cd20b5fe52e2395f31d20d6c6d55c07a1ebc5282d7ed54a565be99103a520
|
|
| MD5 |
8851e8f2b740715d7358eba374f8e22a
|
|
| BLAKE2b-256 |
1398acc5dbaeb72009c04d5fccc8c3a540ce4e28dfe05c1875e9d2d935bed713
|











