membrane protein localization for cryo-ET
Project description
MemBrain-v2
This is the main repository for MemBrain v2. It includes MemBrain-seg, MemBrain-pick, MemBrain-stats, and several Napari tools.


Installation
To install MemBrain-v2, you only need to set up a Python virtual environment and use pip:
pip install membrain
This will install all the necessary dependencies for MemBrain-v2 and give you access to all of MemBrain-v2's tools.
Usage
The main commands you will want to use are:
membrain segment
membrain_pick
membrain_stats
By typing them, you will get more information on how to use them. However, it is strongly recommended to check the respective repositories for more detailed information and documentation. You can find them in the following links:
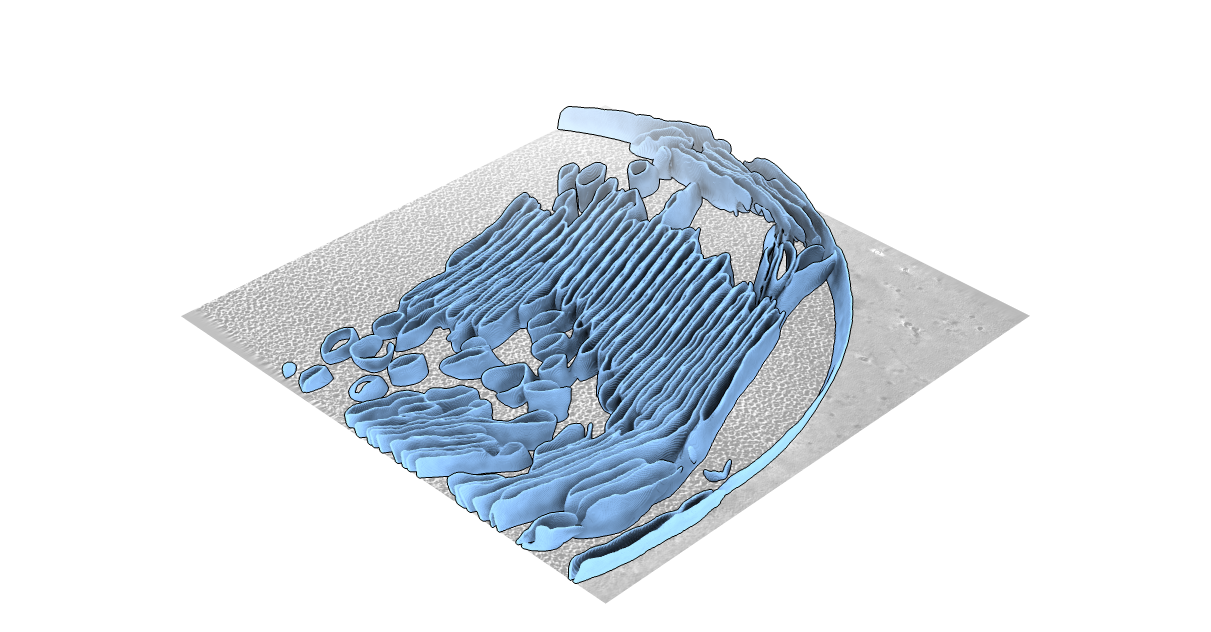
MemBrain-seg
MemBrain-seg gives you the ability to segment 3D cryo-electron tomograms. It is based on the U-Net architecture and is trained on a variety of datasets.
You can find the repository here.
You can also check out the MemBrain-seg tutoral:
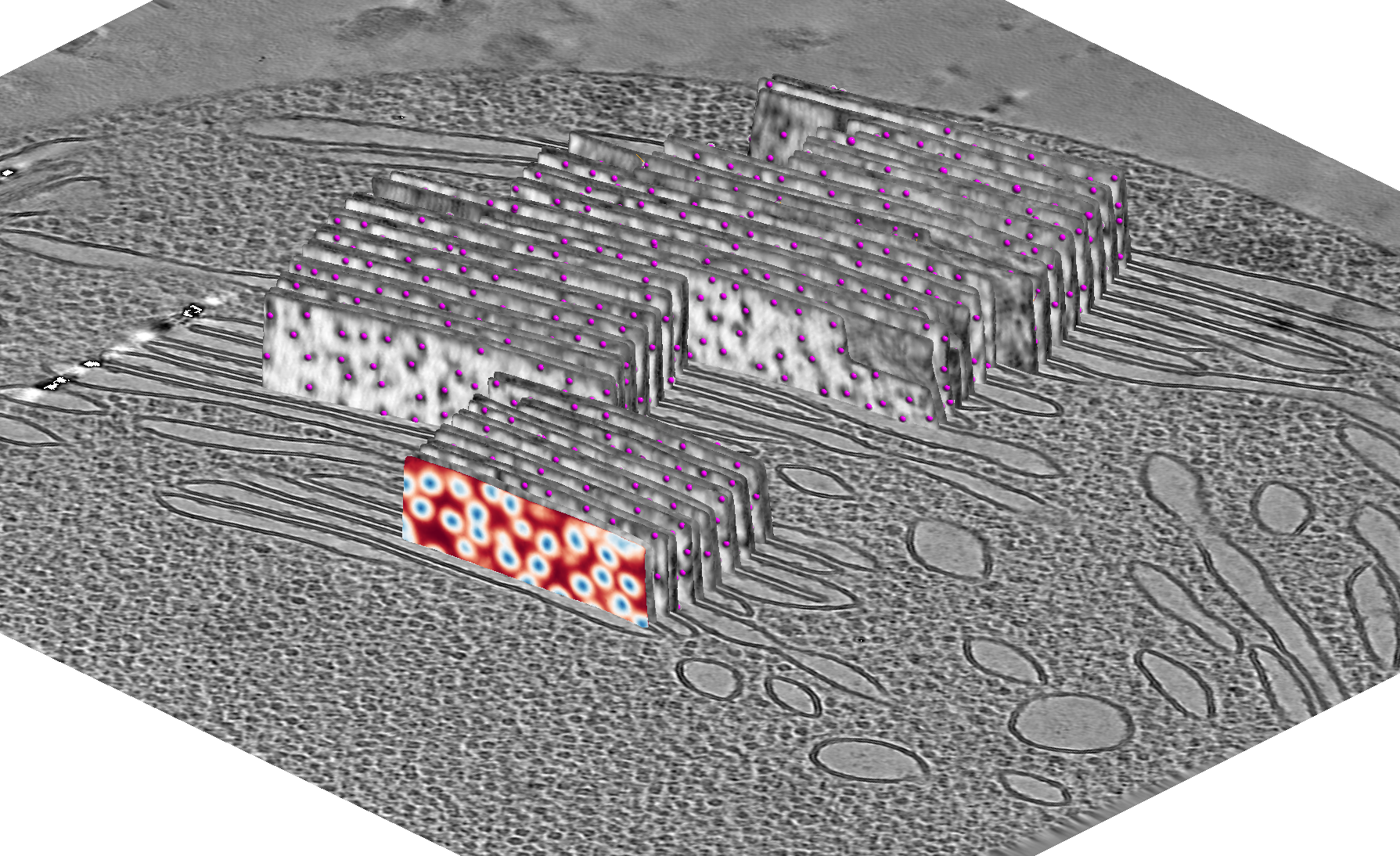
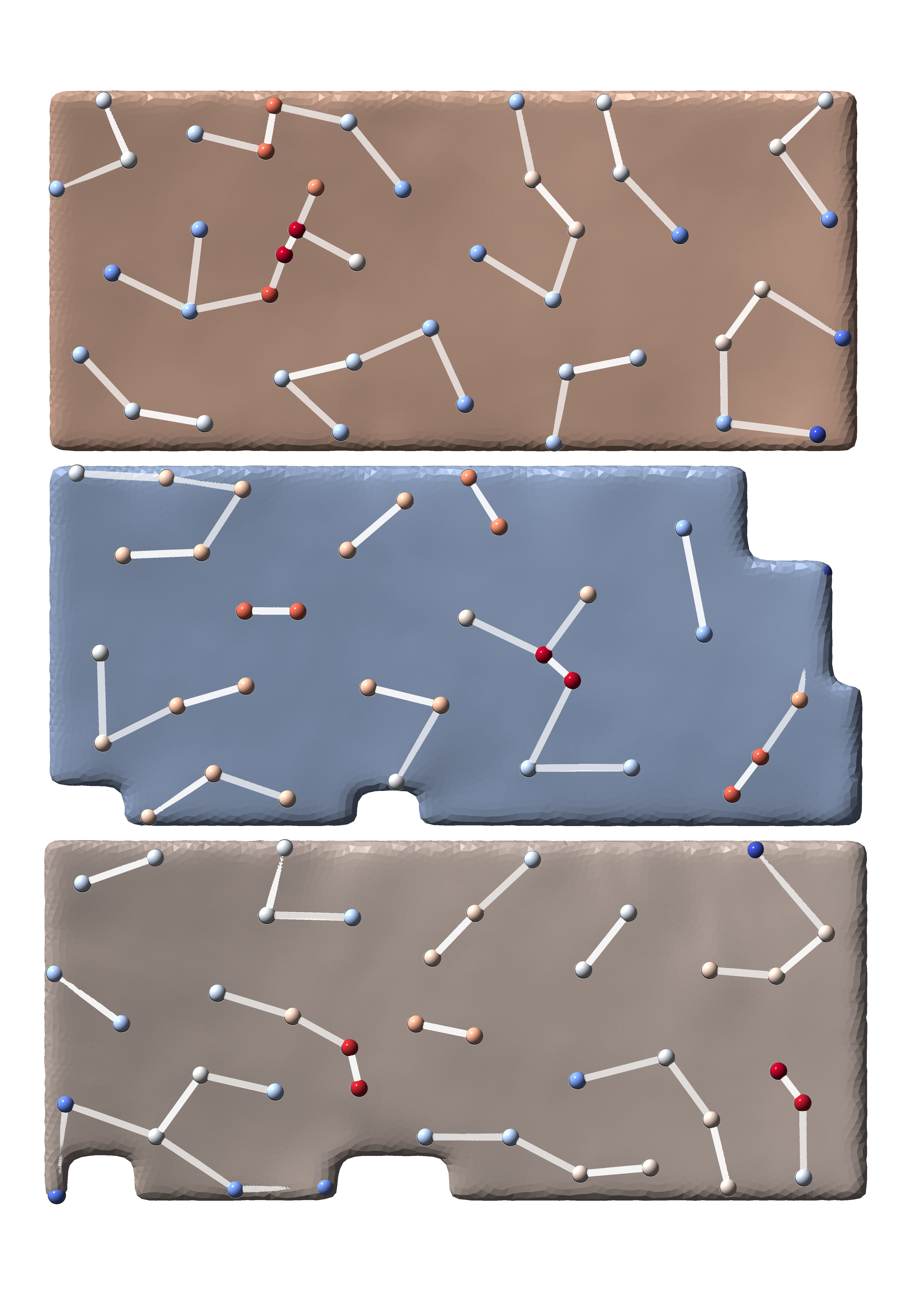
MemBrain-pick
MemBrain-pick is a tool that allows you to pick membrane particles from 3D cryo-electron tomograms. It is based on the diffusionNet architecture, which operates directly on membrane meshes.
You can find the repository here.
You can also check out the MemBrain-pick tutoral:
MemBrain-stats
MemBrain-stats is a tool that allows you to analyze the statistics of membrane particles picked from 3D cryo-electron tomograms, in combination with the membrane architecture.
You can find the repository here.
There is also an example on how to use MemBrain-stats in the above MemBrain-pick tutorial.
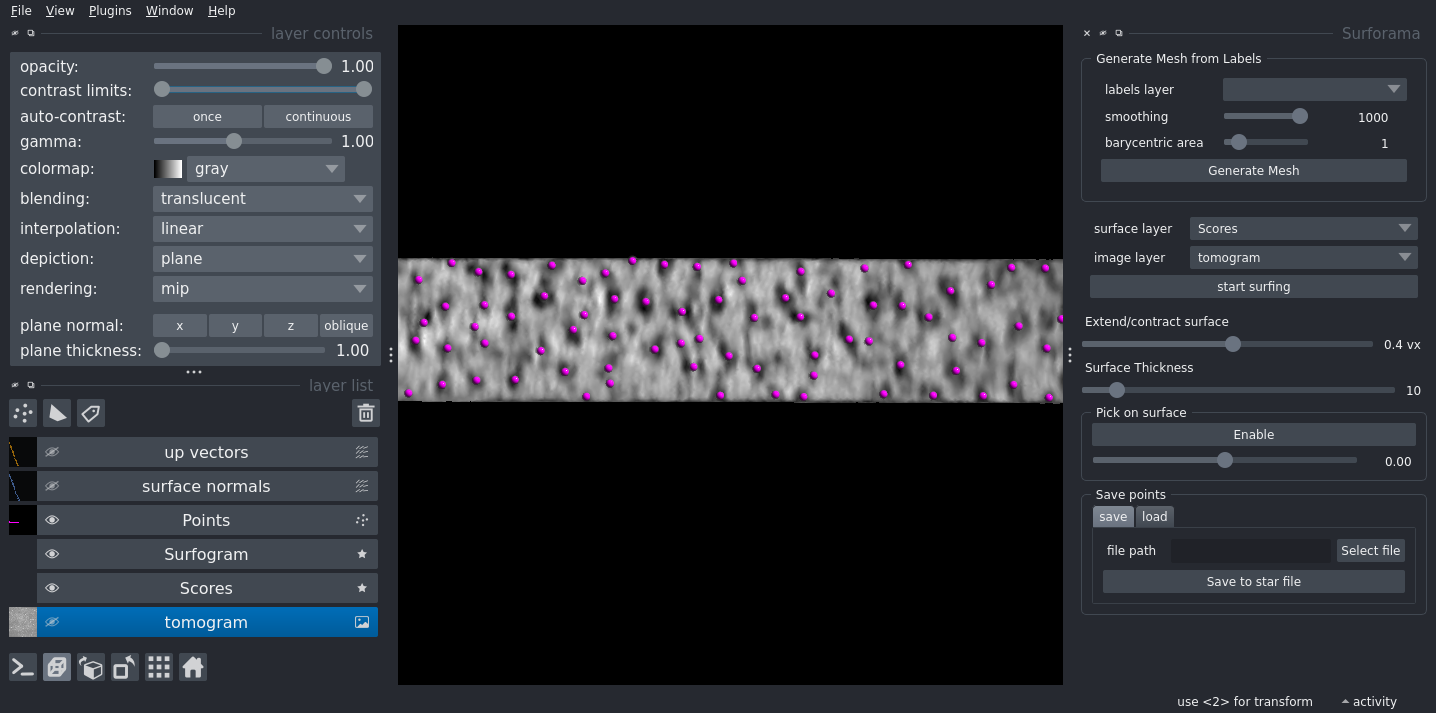
Surforama
Surforama is a Napari-based tool that allows you to visualize and analyze membranes and their embedded particles in 3D cryo-electron tomograms.
You can find the repository here.
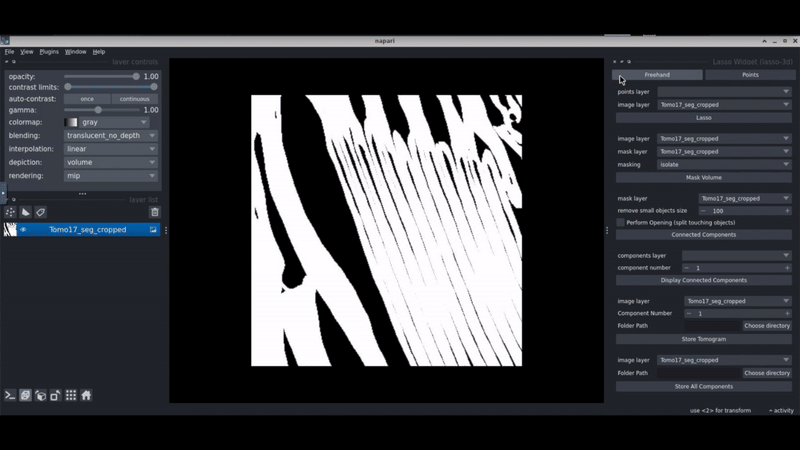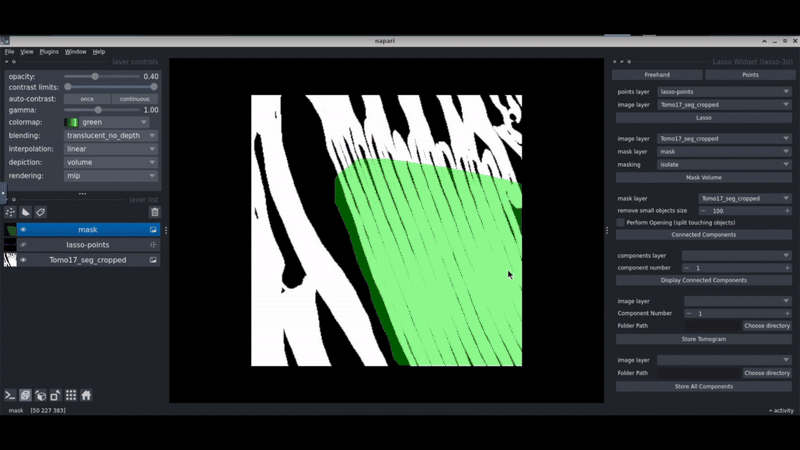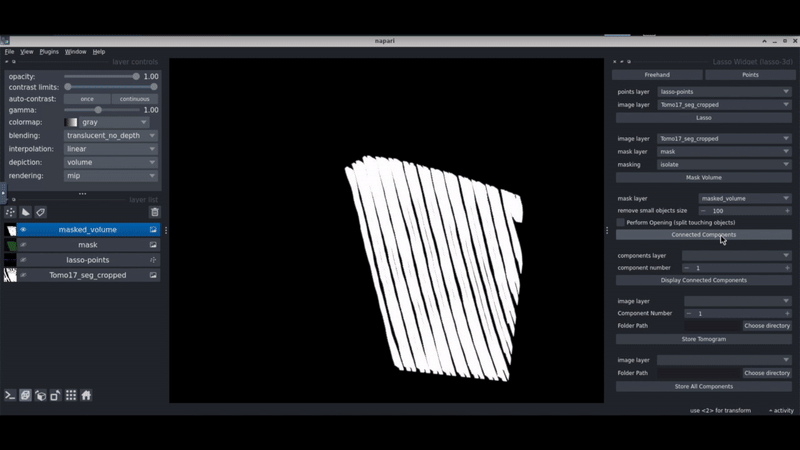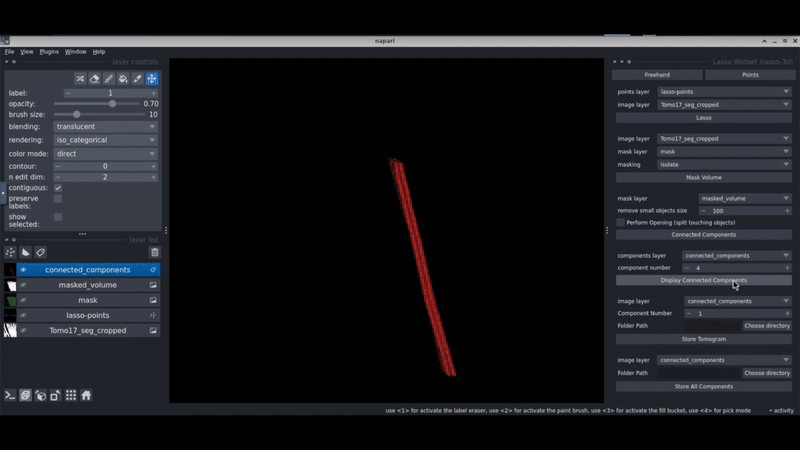
Napari-lasso-3d
Napari-lasso-3d is a Napari plugin that allows you to select regions in your segmentations by drawing a lasso in 3D. This aims to facilitate the selection of regions of interest in 3D cryo-electron tomograms.
You can find the repository here
Citation
If you use MemBrain-v2 in your research, please cite the following paper
[1] Lamm, L., Zufferey, S., Righetto, R.D., Wietrzynski, W., Yamauchi, K.A., Burt, A., Liu, Y., Zhang, H., Martinez-Sanchez, A., Ziegler, S., Isensee, F., Schnabel, J.A., Engel, B.D., and Peng, T, 2024. MemBrain v2: an end-to-end tool for the analysis of membranes in cryo-electron tomography. bioRxiv, https://doi.org/10.1101/2024.01.05.574336
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file membrain-0.0.1.tar.gz.
File metadata
- Download URL: membrain-0.0.1.tar.gz
- Upload date:
- Size: 5.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.16
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
731fbb2e3eca3eedf6be7a80cc86756585cc53fad1b23ac5378453e33689c3aa
|
|
| MD5 |
422e22ed2d0545fa14b2c06d4b2c86ca
|
|
| BLAKE2b-256 |
484d1f98381c2e2425a15134f87add262544a588463d4426b354a448d99670ac
|
File details
Details for the file membrain-0.0.1-py3-none-any.whl.
File metadata
- Download URL: membrain-0.0.1-py3-none-any.whl
- Upload date:
- Size: 2.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.16
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
900274404bedf16a21c03a87313ca1f9563e4a2cc9053c94bbb0696f08e4d448
|
|
| MD5 |
e685415cfcf9edee26b36febde7eae70
|
|
| BLAKE2b-256 |
f998fe5132d854fdbf785af0c91408f7615e45fdb123d1a4c2b687ea012e2ad6
|











