Merge Stardist Masks
Project description
This repository contains the python package for the new StarDist post-processing step StarDist OPP. StarDist OPP allows to use StarDist segmentation on non-star-convex objects. In our paper, we show that StarDist OPP outperforms other methods in instance segmentation tasks for three-dimensional microbial biofilms. Check it out for more information.
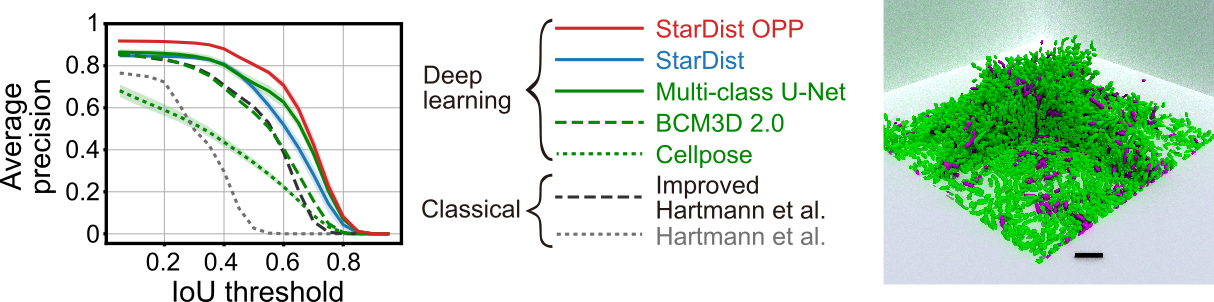
Features
StarDist OPP merges masks together - hence the repository name
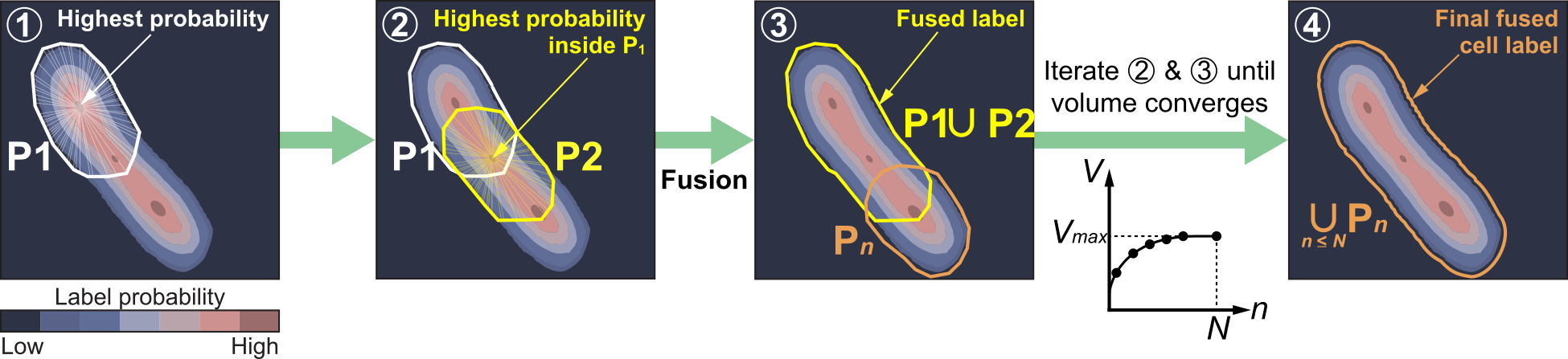
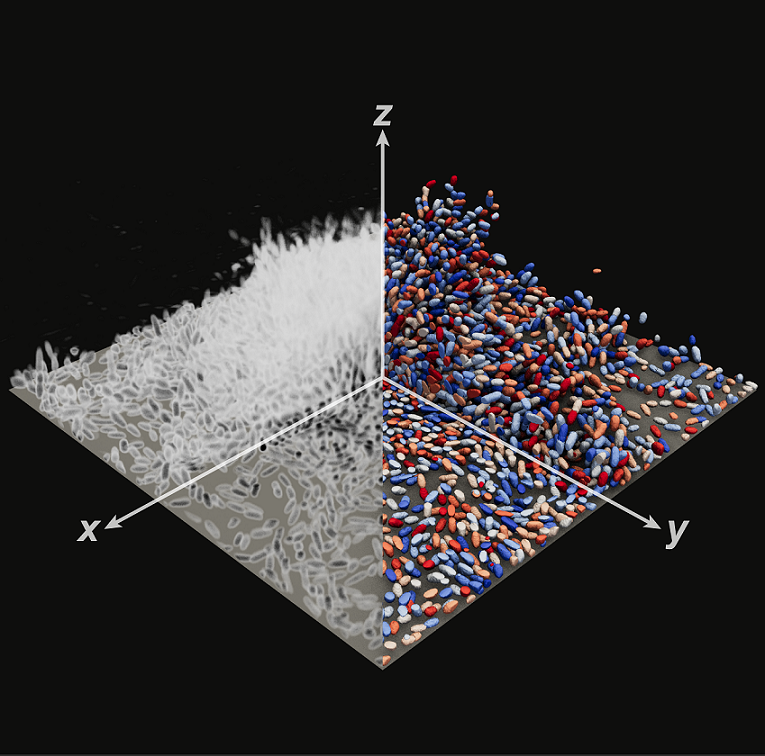
StarDist OPP works in 2D and 3D
In 2D, StarDist OPP works also on big and winding objects
Requirements
A StarDist installation.
Usage
Please see the EXAMPLE in Usage for details or check out the tutorial of our napari plugin to directly use StarDist OPP on your data.
Installation
You can install StarDist OPP via pip from PyPI:
$ pip install merge-stardist-masksContributing
Contributions are very welcome. To learn more, see the Contributor Guide.
License
Distributed under the terms of the MIT license, StarDist OPP is free and open source software.
Issues
If you encounter any problems, please file an issue along with a detailed description.
How to cite
@article{https://doi.org/10.1111/mmi.15064,
author = {Jelli, Eric and Ohmura, Takuya and Netter, Niklas and Abt, Martin and Jiménez-Siebert, Eva and Neuhaus, Konstantin and Rode, Daniel K. H. and Nadell, Carey D. and Drescher, Knut},
title = {Single-cell segmentation in bacterial biofilms with an optimized deep learning method enables tracking of cell lineages and measurements of growth rates},
journal = {Molecular Microbiology},
volume = {119},
number = {6},
pages = {659-676},
keywords = {3D segmentation, biofilm, deep learning, image analysis, image cytometry, Vibrio cholerae},
doi = {https://doi.org/10.1111/mmi.15064},
url = {https://onlinelibrary.wiley.com/doi/abs/10.1111/mmi.15064},
eprint = {https://onlinelibrary.wiley.com/doi/pdf/10.1111/mmi.15064},
abstract = {Abstract Bacteria often grow into matrix-encased three-dimensional (3D) biofilm communities, which can be imaged at cellular resolution using confocal microscopy. From these 3D images, measurements of single-cell properties with high spatiotemporal resolution are required to investigate cellular heterogeneity and dynamical processes inside biofilms. However, the required measurements rely on the automated segmentation of bacterial cells in 3D images, which is a technical challenge. To improve the accuracy of single-cell segmentation in 3D biofilms, we first evaluated recent classical and deep learning segmentation algorithms. We then extended StarDist, a state-of-the-art deep learning algorithm, by optimizing the post-processing for bacteria, which resulted in the most accurate segmentation results for biofilms among all investigated algorithms. To generate the large 3D training dataset required for deep learning, we developed an iterative process of automated segmentation followed by semi-manual correction, resulting in >18,000 annotated Vibrio cholerae cells in 3D images. We demonstrate that this large training dataset and the neural network with optimized post-processing yield accurate segmentation results for biofilms of different species and on biofilm images from different microscopes. Finally, we used the accurate single-cell segmentation results to track cell lineages in biofilms and to perform spatiotemporal measurements of single-cell growth rates during biofilm development.},
year = {2023}
}
Credits
This project was generated from @cjolowicz’s Hypermodern Python Cookiecutter template.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file merge_stardist_masks-0.2.2.tar.gz.
File metadata
- Download URL: merge_stardist_masks-0.2.2.tar.gz
- Upload date:
- Size: 28.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.12.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
0cf924449bd9b2ae1707fc4db2562913c1801311eecfb89108563215dbe48892
|
|
| MD5 |
f724a7322fbcd8a65e318c7d9371c565
|
|
| BLAKE2b-256 |
73e3067ac6a3b56dea2a728a2e1428157cf42d93605e92c27fa826fbc783942c
|
Provenance
The following attestation bundles were made for merge_stardist_masks-0.2.2.tar.gz:
Publisher:
release.yml on gatoniel/merge-stardist-masks
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
merge_stardist_masks-0.2.2.tar.gz -
Subject digest:
0cf924449bd9b2ae1707fc4db2562913c1801311eecfb89108563215dbe48892 - Sigstore transparency entry: 312120575
- Sigstore integration time:
-
Permalink:
gatoniel/merge-stardist-masks@c28105bee44c0b224b49239d71d5859e1035c45e -
Branch / Tag:
refs/heads/main - Owner: https://github.com/gatoniel
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yml@c28105bee44c0b224b49239d71d5859e1035c45e -
Trigger Event:
push
-
Statement type:
File details
Details for the file merge_stardist_masks-0.2.2-py3-none-any.whl.
File metadata
- Download URL: merge_stardist_masks-0.2.2-py3-none-any.whl
- Upload date:
- Size: 34.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.12.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
dd0761fa5fd9d92a97c6336dcb65ec9b2d7b335f07212032f42e97c3a3b02773
|
|
| MD5 |
0c47acaed342a760f0e3bd4c410f30a7
|
|
| BLAKE2b-256 |
63290c6459b5ff2d4b7f79e467efa1d51552e9dee2cef2bd7aca96e3f7a46bf2
|
Provenance
The following attestation bundles were made for merge_stardist_masks-0.2.2-py3-none-any.whl:
Publisher:
release.yml on gatoniel/merge-stardist-masks
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
merge_stardist_masks-0.2.2-py3-none-any.whl -
Subject digest:
dd0761fa5fd9d92a97c6336dcb65ec9b2d7b335f07212032f42e97c3a3b02773 - Sigstore transparency entry: 312120586
- Sigstore integration time:
-
Permalink:
gatoniel/merge-stardist-masks@c28105bee44c0b224b49239d71d5859e1035c45e -
Branch / Tag:
refs/heads/main - Owner: https://github.com/gatoniel
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yml@c28105bee44c0b224b49239d71d5859e1035c45e -
Trigger Event:
push
-
Statement type:





















