Multiomics and Ecological Spatial Analysis for Quantitative Decoding of Cellular Neighborhoods and Tissue Compartments
Project description
MESA: Multiomics and Ecological Spatial Analysis for Quantitative Decoding of Tissue Disease States
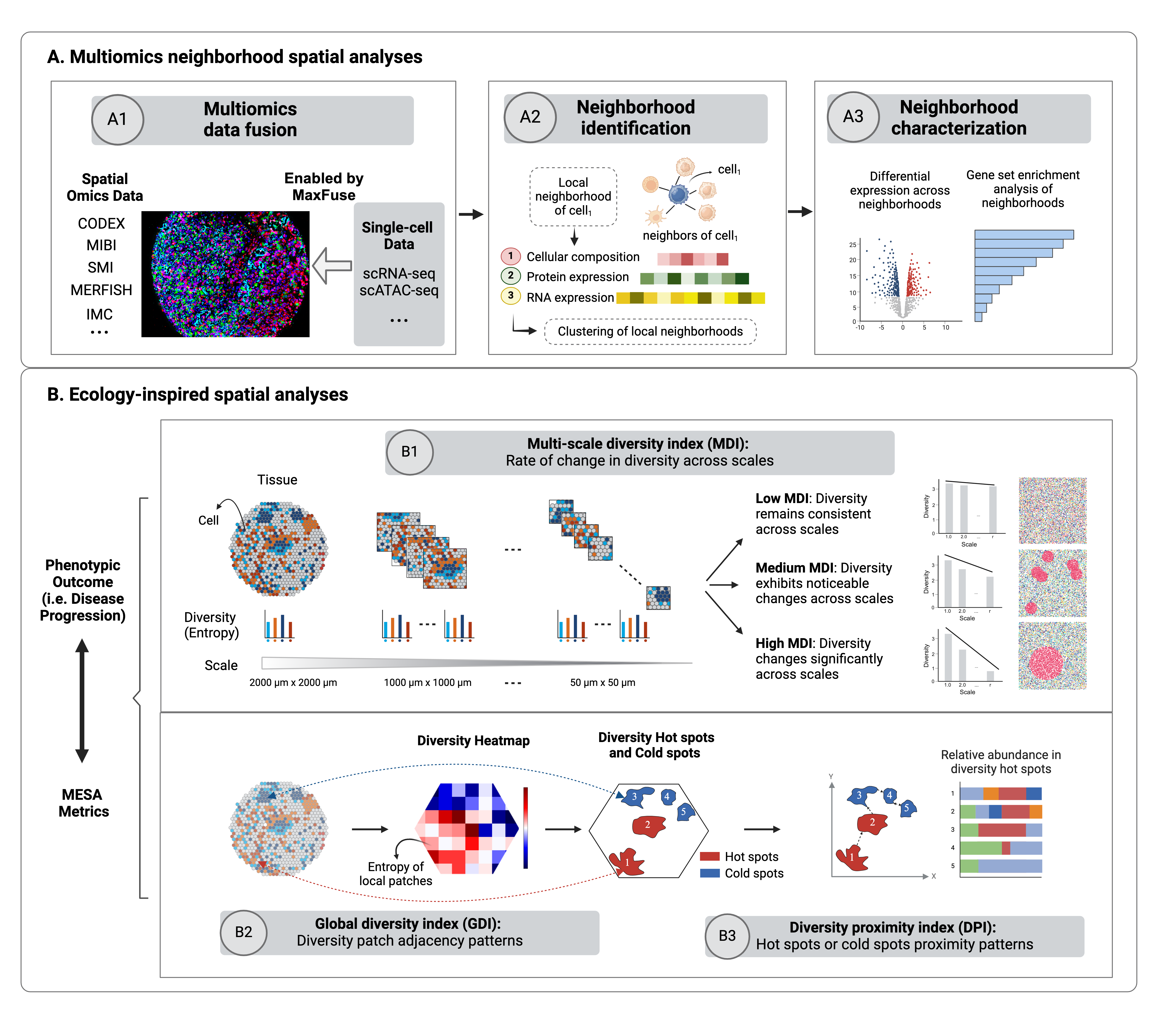
MESA is a novel pipeline that brings ecological principles together with multiomics data integration thereby enabling deeper and more quantitative decoding of the functional and spatial shifts of tissue remodeling across disease states.
- Drawing inspiration from ecological studies, MESA adapts diversity metrics traditionally used to gauge biodiversity for spatial omics data, creating tools for systematic quantification of cellular diversity. Specifically, we introduce a multi-scale diversity index, alongside global and local diversity indices, to capture not only the overarching diversity of a tissue but also the localized patterns and dependencies.
- Furthermore, MESA employs a multi-omics approach to spatial omics analyses. MESA in silico amalgamates cross-modality single-cell data to enrich the context of spatial omics observations. With the additional layers of information brought to bear by multiomics, MESA facilitates an extended view of cellular neighborhoods and their spatial interactions within tissue microenvironments. MESA's approach, incorporating differential expression, gene set enrichment, and ligand-receptor interaction analyses within these spatially defined cellular assemblies, further enhances a mechanistic understanding of tissue remodeling across disease states.
Installation
MESA is hosted on pypi and can be installed via pip.
pip install mesa-py
Visit our documentation to see examples and tutorials!
License
MESA is under the Academic Software License Agreement, please use accordingly.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file mesa_py-0.1.2.tar.gz.
File metadata
- Download URL: mesa_py-0.1.2.tar.gz
- Upload date:
- Size: 84.0 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.0.0 CPython/3.11.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
bf791d14d30cf2afa8f9a0dac8f85c1b7958be299e04d750531ea20cdc01bdb5
|
|
| MD5 |
e026478604b3287d1832f88c47046980
|
|
| BLAKE2b-256 |
2b5481986fb0abca61383363a6b1be9ab44b1dd5ea0d25f9f486fe629dc1a7cc
|
File details
Details for the file mesa_py-0.1.2-py3-none-any.whl.
File metadata
- Download URL: mesa_py-0.1.2-py3-none-any.whl
- Upload date:
- Size: 84.0 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.0.0 CPython/3.11.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f8d882e8bfc478f4902808dd47e6cadfbb20f1e116f01caeabb9e1bb92154fc5
|
|
| MD5 |
77b6d317ab30d9f83d4d456393d96f10
|
|
| BLAKE2b-256 |
8063821011c5d44f988648c7a98913f8888ac073cac3725d59ccf4a8d45fa955
|











