Spatial metabolic communication flow of cells.
Project description
MetaChat
Brief introduction
MetaChat is a Python package to screen metabolic cell communication (MCC) from spatial multi-omics data of transcriptomics and metabolomics. It contains many intuitive visualization and downstream analysis tools, provides a great practical toolbox for biomedical researchers.
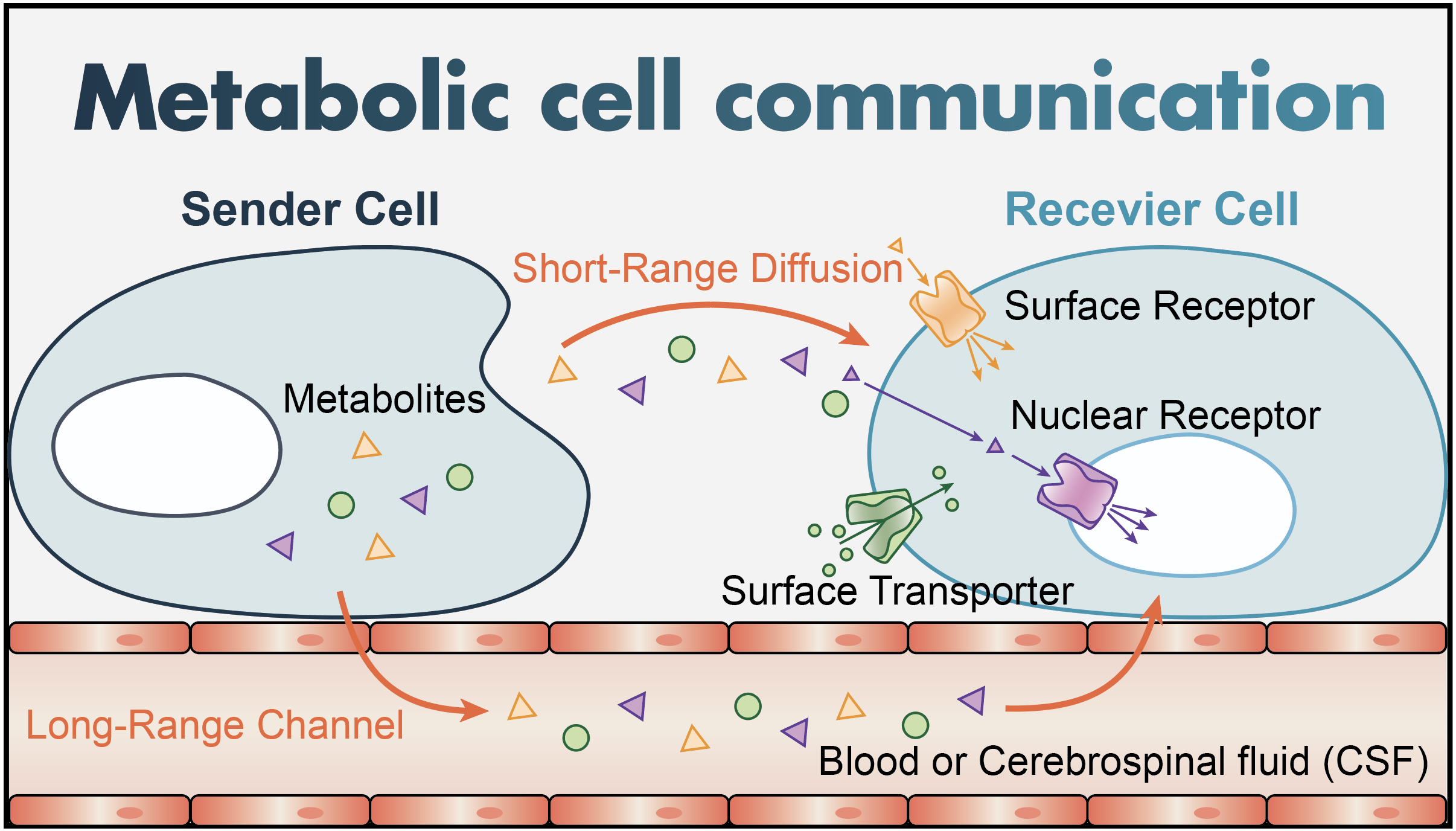
Metabolic cell communication
Metabolic cell-cell communication (MCC) occurs when sensor proteins in the receiver cells detect metabolites in their environment, activating intracellular signaling events. There are three major potential sensors of metabolites: surface receptors, nuclear receptors, and transporters. Metabolites secreted from cells are either transported over short-range distances (a few cells) via diffusion through extracellular space, or over long-range distances via the bloodstream and the cerebrospinal fluid (CSF).
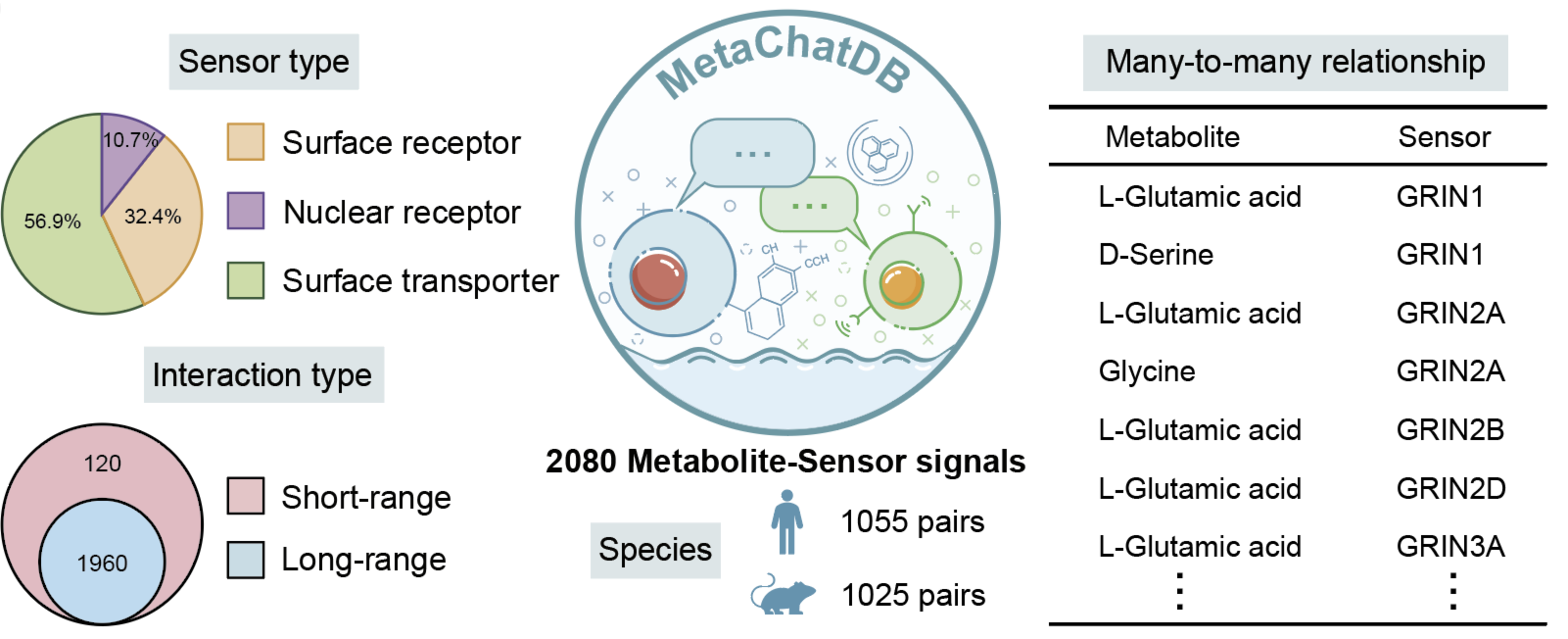
MetaChatDB
MetaChatDB is a literature-supported database for metabolite-sensor interactions for both human and mouse. All the metabolite-sensor interactions are reported based on peer-reviewed publications. Specifically, we manually build MetaChatDB by integrating three high-quality databases (PDB, HMDB, UniProt) that are being continually updated.
Documentation, and Tutorials
For more basic tutorial and real data examples, please see MetaChat documentation that is available through the link https://metachat.readthedocs.io/en/latest/.
Analysis pipeline
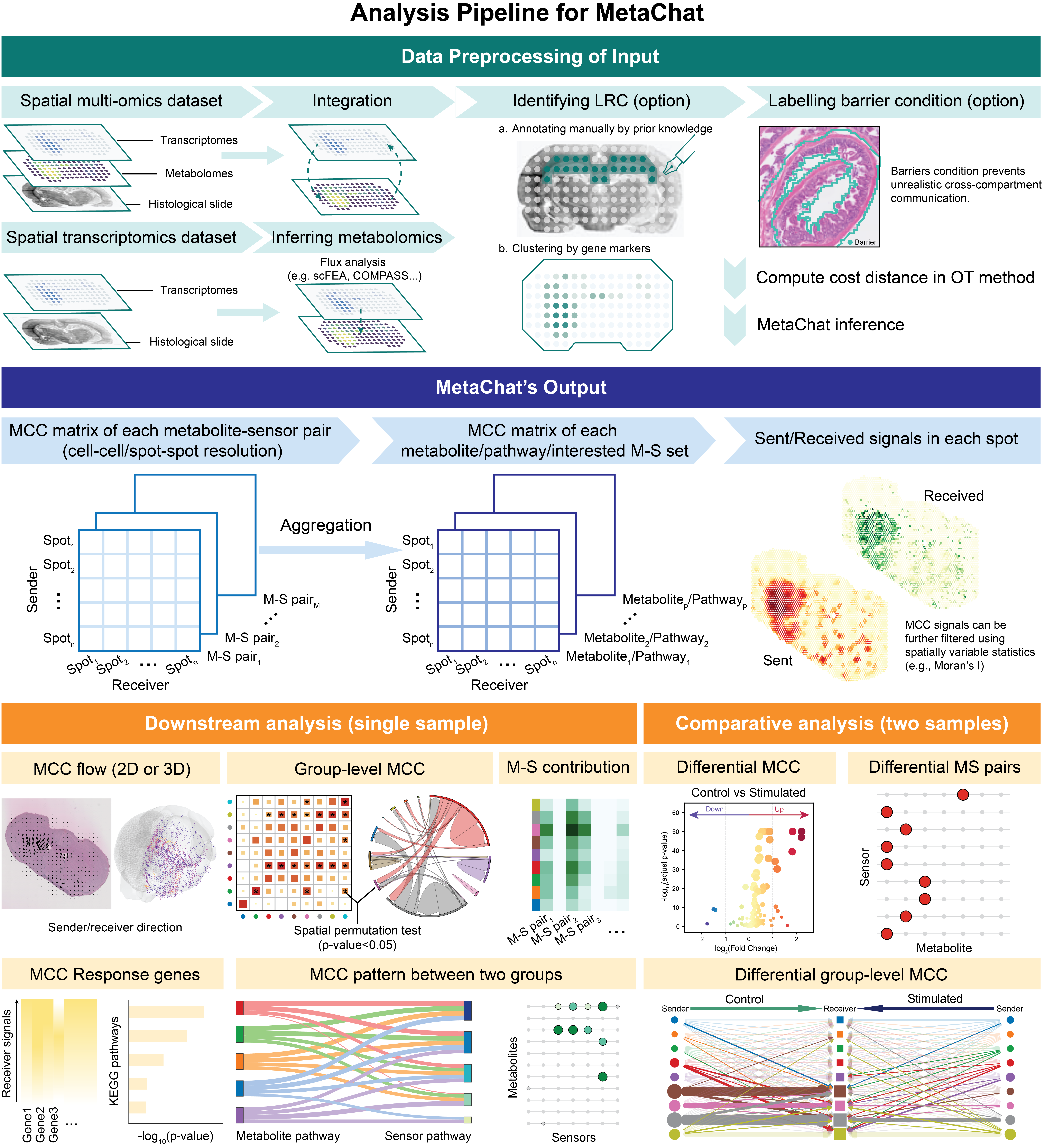
Installation
System requirements
Recommended operating systems: macOS or Linux. MetaChat was developed and tested on Linux and macOS.
Python requirements
MetaChat was developed using python 3.9.
Installation using pip
We suggest setting up MetaChat in a separate mamba or conda environment to prevent conflicts with other software dependencies. Create a new Python environment specifically for MetaChat and install the required libraries within it.
mamba create -n metachat_env python=3.9 r-base=4.3.2
mamba activate metachat_env
pip install metachat
if you use conda, r-base=4.3.2 may not included in the channels. Instead, you can r-base=4.3.1 in conda.
Reference
Luo S., Almet A.A., Zhao W., He C., Tsai Y.-C., Ozaki H., Sugita B.K., Du K., Shen X., Cao Y., Yang Q., Watanabe M., Nie Q.* Spatial metabolic communication flow of cells.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file metachat-0.0.7.tar.gz.
File metadata
- Download URL: metachat-0.0.7.tar.gz
- Upload date:
- Size: 294.4 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.9.23
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c873576e7d83639f8079ec6fa9e982df343b8222a69b87079ef9437c5754059c
|
|
| MD5 |
d3fb5661dbc7f0911da8dad42f10b86f
|
|
| BLAKE2b-256 |
1fc2dc92bd4150ef37b23336346a4a07916768d00ceb9697d46ffc26460f55e3
|
File details
Details for the file metachat-0.0.7-py3-none-any.whl.
File metadata
- Download URL: metachat-0.0.7-py3-none-any.whl
- Upload date:
- Size: 679.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.9.23
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c95ef0a6bc1aa2a858c0ef9fc1e789f21dab70513195e2ba774fd48a8fede6e8
|
|
| MD5 |
761eb2fed43940b7fc19b5f79ff8534f
|
|
| BLAKE2b-256 |
7dd88242c2ca5137dc0b5a4d80e00d76a49333d6b45c06bd27d49fbb1bcecab1
|











