A portable and modular meta-predictor for identifying Long Non-coding RNAs (lncRNAs).
Project description
metaLncRNA v1.2.3 🧬🤖

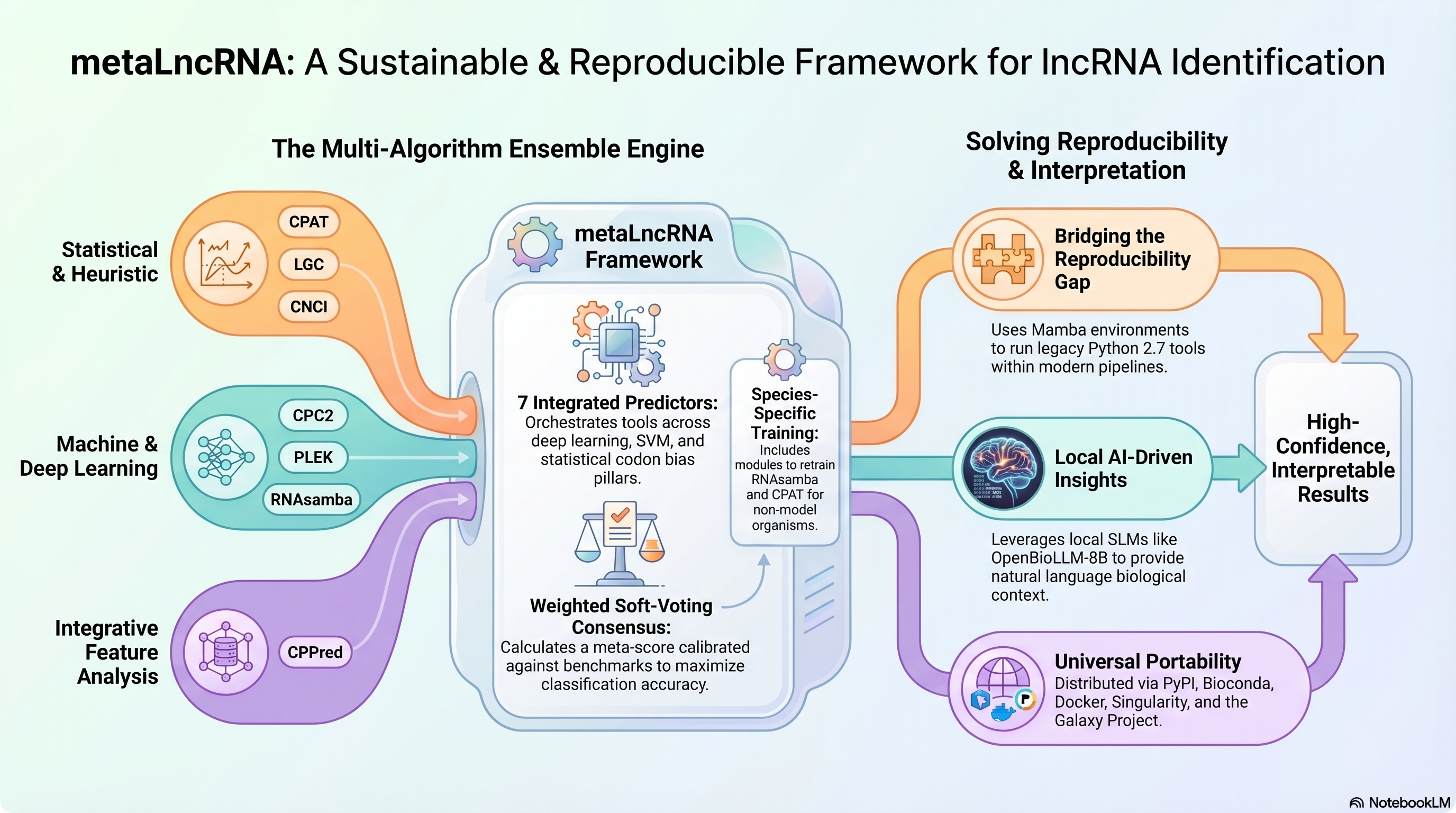
metaLncRNA is a modular, high-performance Python framework designed to identify Long Non-coding RNAs (lncRNAs) by orchestrating an ensemble of seven diverse computational tools. It resolves the "reproducibility gap" by automating environment management and providing a robust consensus prediction through weighted soft-voting.
📂 Repository Structure
.
├── conda/ # Bioconda recipe and metadata
├── deploy/ # Containerization (Dockerfile, Singularity.def)
├── docs/ # Technical documentation and user guides
├── examples/ # Quick-start samples (FASTA, config templates)
├── galaxy/ # Galaxy Tool wrapper and test data
├── INPI_Registration/ # Legal software registration assets
├── paper/ # JOSS publication manuscript and bibliography
├── scripts/ # Bash scripts for HPC/Batch processing
├── src/
│ └── metalncrna/ # Main Python Package
│ ├── cli.py # Command-line interface entry point
│ ├── adapters/ # Wrappers for 7 lncRNA predictors
│ ├── engine/ # Core logic (Consensus, Dispatcher, Trainer)
│ ├── utils/ # AI Agent, Env management, Reports, FASTA handling
│ ├── data/ # Built-in weights and default configurations
│ └── third_party/ # Bundled legacy tools (CNCI, CPPred, LGC)
├── tests/ # Comprehensive Unit and Integration tests
├── pyproject.toml # Build system and dependency definitions
└── pixi.toml # Environment management configuration
🧩 Core Components Detail
src/metalncrna/adapters/: Orchestrates external tools like RNAsamba, CPAT, CPC2, etc., providing a unified interface for prediction.src/metalncrna/engine/:consensus.py: Implements the weighted soft-voting algorithm.dispatcher.py: Manages parallel execution of the ensemble.
src/metalncrna/utils/agent.py: Integrates with local LLMs (Ollama) for automated biological interpretation of results.galaxy/: AllowsmetaLncRNAto be integrated into Galaxy instances, supporting reproducible web-based workflows.
🔧 Recent Fixes (v1.2.3)
- CNCI Stability: Fixed a critical hang in the CNCI legacy tool caused by non-canonical nucleotides (e.g., K, V, M) and a multiprocessing deadlock in the original Python 2.7 implementation.
- Improved Filtering: Implemented rigorous FASTA validation in the CNCI adapter to exclude sequences with ambiguous characters, preventing
KeyErrorcrashes. - Resource Optimization: Optimized tool dispatching by limiting CNCI threads to 4, reducing I/O overhead and improving performance for small-to-medium files.
- Debug Resiliency: The dispatcher now preserves intermediate files automatically if a tool failure occurs, facilitating troubleshooting.
🔧 Previous Fixes (v1.2.1)
- Consensus Logic: Updated
consensus_supportto reflect the number of tools that agree with the final consensus label, providing better interpretability. - CPC2 Integration: Fixed a critical parsing error where coding probability and label columns were mismatched (v1.1.8).
- Cleanup: Removed unimplemented/experimental adapters to ensure stability.
⚙️ Configuration
metaLncRNA follows a robust configuration loading order:
- Internal Defaults: Built-in weights and paths in
src/metalncrna/data/default_config.yaml. - Local Config:
metaLncRNA_config.yamlin your current working directory. - User Home:
~/.metalncrna/config.yaml. - Explicit Path: Provided via the
-cor--configflag.
🚀 Key Features
- Ensemble Prediction: Combines 7 tools (RNAsamba, CPAT, CPC2, PLEK, CNCI, CPPred, LGC).
- Interactive AI Agent: Integrated local LLM assistant (Llama-3.2 or OpenBioLLM) to interpret results and explain classification decisions.
- Reproducibility First: Built-in environment isolation via Mamba and Pixi.
- Standardized Reports: Comprehensive TSV reports with tool congruence metrics.
- Publication Ready: Formatted according to JOSS standards for scientific software.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file metalncrna-1.2.3.tar.gz.
File metadata
- Download URL: metalncrna-1.2.3.tar.gz
- Upload date:
- Size: 1.9 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5987011c6075219b0b0a61b981c40c5b45e17df562f04acff1bdc39e4ac519d5
|
|
| MD5 |
c165f888068110015cbcd7d85782765c
|
|
| BLAKE2b-256 |
396293da0815184275500847c242305eb8edb0fa1610424d061ac9d582ee299c
|
File details
Details for the file metalncrna-1.2.3-py3-none-any.whl.
File metadata
- Download URL: metalncrna-1.2.3-py3-none-any.whl
- Upload date:
- Size: 1.9 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c533ca11dc450ac9eacd13feb56d3f806b1d0010394fde16f824937ea6712a25
|
|
| MD5 |
04f238f63e03f36eac00bc46607dc3a4
|
|
| BLAKE2b-256 |
ce4ef8943953d28938750704bf1d2ed9167a6637205b06d20c43821b118d040b
|

























