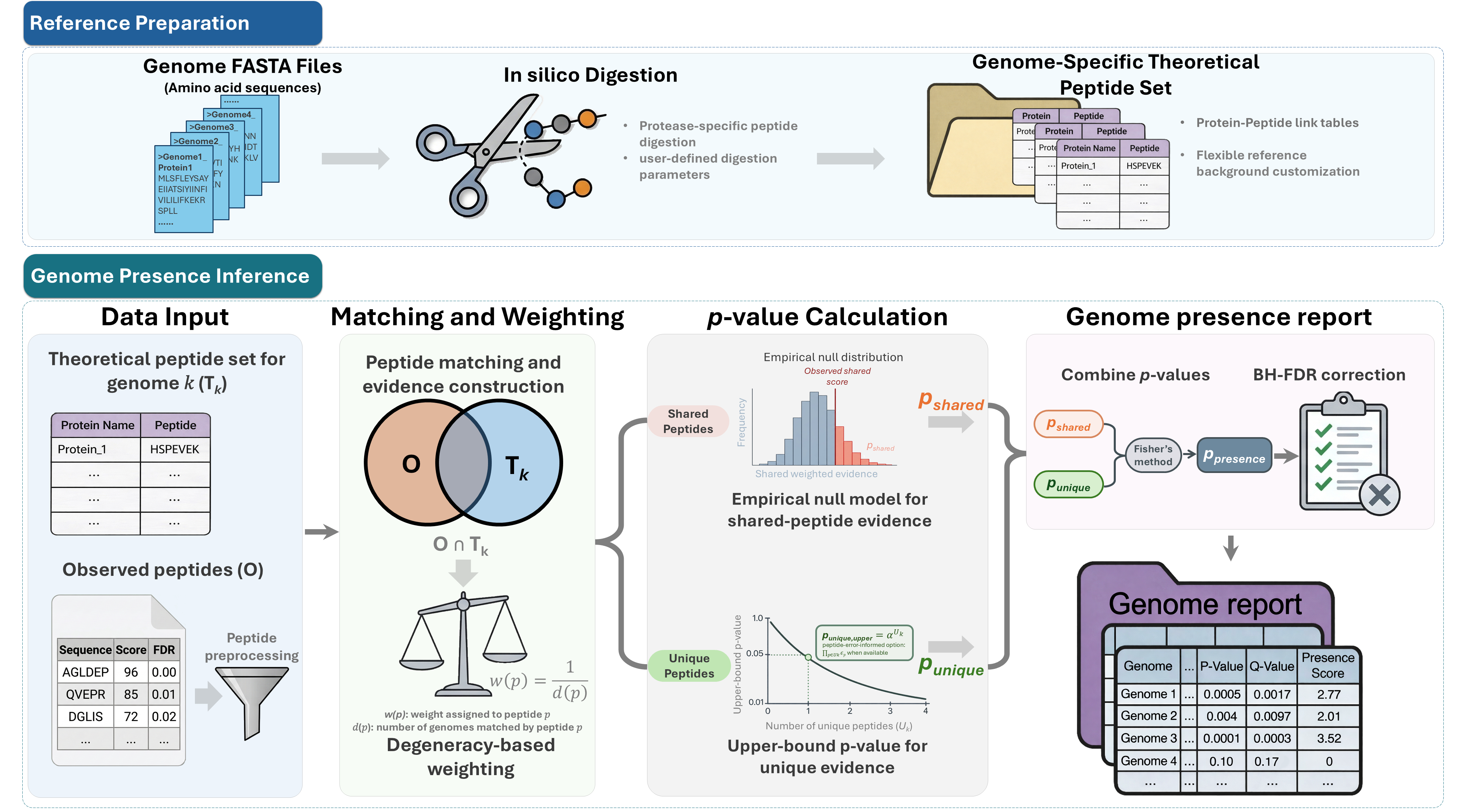
Genome-level presence inference from metaproteomic peptide lists.
Project description
MetaUmbra
Genome-level presence inference from metaproteomic peptides
MetaUmbra performs genome-level presence inference from metaproteomic peptide lists. It combines unique peptide support with weighted shared peptide evidence to identify statistically supported microbial genomes and generate interpretable presence rankings.
Main features
- Evaluate candidate genome support from metaproteomic peptide tables
- Build genome-specific theoretical peptide references from protein FASTA files
- Support user-defined genome collections, including isolate genomes, strain panels, and MAG catalogs
- Use both unique and shared peptide evidence for genome presence inference
- Report genome-level p-values, BH-adjusted q-values, and presence scores
- Provide GUI, command-line, and Python workflow support
- Support peptide tables from common metaproteomics workflows such as DIA-NN and MaxQuant
Workflow overview
Installation
MetaUmbra requires Python 3.10 or newer.
pip install metaumbra
Usage
MetaUmbra can be used through either the graphical interface or the command line.
Graphical interface
metaumbra-gui
The GUI supports FASTA digestion, peptide table loading, genome presence scoring, and result export.
Command line
MetaUmbra provides separate commands for the main workflow steps:
metaumbra digest --help
metaumbra score --help
metaumbra extract-parquet --help
A typical workflow is:
metaumbra digest ...
metaumbra score ...
Use metaumbra extract-parquet ... to convert DIA-NN parquet reports to peptide TSV files before scoring.
Input
MetaUmbra requires:
- Protein FASTA files, with one FASTA file per genome
- An observed peptide table containing peptide sequences
Optional inputs include peptide scores, peptide-level error values, decoy flags, and genome lineage annotations.
Output
The main output is a TSV table containing genome-level evidence and significance values.
Key output columns include:
| Column | Description |
|---|---|
genome_id |
Candidate genome identifier |
num_peptides_matched |
Number of observed peptides matched to the genome |
num_peptides_unique |
Number of matched peptides unique to the genome |
weighted_evidence |
Total degeneracy-weighted peptide evidence |
weighted_evidence_shared |
Weighted evidence from shared peptides |
p_presence |
Genome-level p-value |
q_presence |
BH-adjusted genome-level q-value |
presence_score |
Ranking score based on q-value |
Citation
If you use MetaUmbra, please cite:
Wu Q, Ning Z, Zhang A, Cheng K, Figeys D. MetaUmbra: Statistically Controlled Genome-Level Presence Inference from Metaproteomic Peptides.[J]. bioRxiv, 2026.
A formal citation will be added after publication.
Contact
For questions or issues, please use the GitHub issue tracker or contact the corresponding author listed in the associated manuscript.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file metaumbra-1.1.2.tar.gz.
File metadata
- Download URL: metaumbra-1.1.2.tar.gz
- Upload date:
- Size: 3.2 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
30686764ea500c281c87fa9d2969f069146d363493db03e731d67b71bf4d5dae
|
|
| MD5 |
5198fa6f48c6e38eeb9687c22cadfc1a
|
|
| BLAKE2b-256 |
eb89585a4edc769d6ca641d4860e914f0f35891e8855151199df8dd039221308
|
File details
Details for the file metaumbra-1.1.2-py3-none-any.whl.
File metadata
- Download URL: metaumbra-1.1.2-py3-none-any.whl
- Upload date:
- Size: 3.2 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
6021238e5a719d8b87856213077561aef1d337d2acee8c2184e0baa4d1d92bda
|
|
| MD5 |
8faa13eff424e3ea895fec33605345f0
|
|
| BLAKE2b-256 |
2b9ce3ba90003576d0d7d8cf3c08baebe8af9e3e47d28d26c955f8cd7cf1dcb2
|