A logistic regression model with missing data handling using the SAEM algorithm.
Project description
misaem: Logistic Regression with Missing Covariates
misaem is a Python package that implements logistic regression for data with missing covariates. It is based on the Stochastic Approximation of the Expectation-Maximization (SAEM) algorithm. The algorithm relies on Metropolis-Hastings sampling within the MCMC step to handle the imputation of missing data, under the assumption that the covariates follow a multi-variate normal distribution.
This package provides a robust method for parameter estimation and prediction under Missing Completely at Random (MCAR) and Missing At Random (MAR) assumptions. It is a direct port of the misaem R package and is designed to integrate seamlessly with the scikit-learn ecosystem, adhering to its API conventions for ease of use.
🌟 Key Features
-
Robust Handling of Missing Data: Employs the SAEM algorithm to handle missing covariate data during model fitting and prediction.
-
Predictive Power: Can be used to make predictions on new, incomplete datasets.
-
scikit-learn Compatibility: Follows scikit-learn's API, allowing for direct use in pipelines, cross-validation, and other standard workflows.
-
Statistical Foundation: Provides estimated coefficients and their standard errors, enabling statistical inference.
💻 Installation
You can install the package via pip:
pip install misaem
🚀 Getting Started
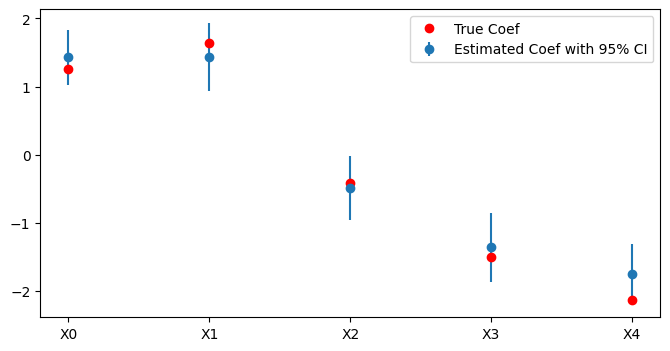
Here is a full example demonstrating how to use SAEMLogisticRegression to fit a model, make predictions, and visualize the results.
import numpy as np
from misaem import SAEMLogisticRegression
import matplotlib.pyplot as plt
# 1. Generate Data
np.random.seed(1324)
n_train = 1000
n_test = 300
n_features = 5
cov = [[1,0.75,0.5,0.25,0],
[0.75,1,0.75,0.5,0.25],
[0.5,0.75,1,0.75,0.5],
[0.25,0.5,0.75,1,0.75],
[0,0.25,0.5,0.75,1]]
X = np.random.multivariate_normal(mean=np.zeros(n_features), cov=cov, size=n_train + n_test)
true_beta = np.hstack([0.5, np.random.normal(size=n_features)])
linear_pred = np.hstack([np.ones((n_train + n_test, 1)), X]) @ true_beta
probabilities = 1 / (1 + np.exp(-linear_pred))
y = np.random.binomial(1, probabilities)
X_missing = X.copy()
missing_mask = np.random.rand(n_train + n_test, n_features) < 0.2
X_missing[missing_mask] = np.nan
X_train = X_missing[:n_train,:]
X_test = X_missing[(n_train+1):]
y_train = y[:n_train]
y_test = y[(n_train+1):]
# 2. Fit SAEMLogisticRegression
model = SAEMLogisticRegression()
model.fit(X_train, y_train)
# 3. Predict on test set
preds = model.predict_proba(X_test, method="map")[:,1]
# 4. Evaluate
accuracy = np.mean((preds > 0.5) == y_test)
print(f"Accuracy of SAEMLogisticRegression: {accuracy:.4f}")
coef = model.coef_.ravel()
se = model.std_err_.ravel()[1:] # remove the intercept
plt.figure(figsize=(8, 4))
plt.errorbar(range(len(coef)), coef, yerr=1.96*se, fmt='o', label='Estimated Coef with 95% CI')
plt.xticks(range(len(coef)), [f'X{i}' for i in range(len(coef))])
plt.plot(range(len(coef)), true_beta[1:], 'o', color='red', label='True Coef')
plt.legend()
plt.show()
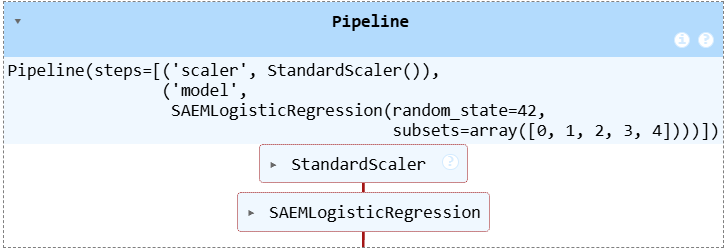
⚙️ scikit-learn Integration
The misaem package is compatible with scikit-learn's API, which allows you to seamlessly integrate it into your machine learning pipelines. For instance, you can combine it with data preprocessing steps like StandardScaler.
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from misaem import SAEMLogisticRegression
pipeline = Pipeline([
('scaler', StandardScaler()),
('model', SAEMLogisticRegression(random_state=42))
])
pipeline.fit(X_train, y_train)
y_pred = pipeline.predict(X_test)
accuracy_pipeline = np.mean(y_pred == y_test)
print(f"Accuracy of SAEMLogisticRegression: {accuracy_pipeline:.4f}")
📚 Reference
This package is based on the SAEM algorithm for logistic regression with missing covariates. For more details on the methodology, please refer to the following publication:
Jiang, W., Josse, J., Lavielle, M., & TraumaBase Group. (2020). Logistic regression with missing covariates—Parameter estimation, model selection and prediction within a joint-modeling framework. Computational Statistics & Data Analysis, 145, 106907.
🤝 Contributing
We welcome contributions! If you encounter any bugs, have feature requests, or want to contribute to the code, please open an issue or a pull request on our GitHub repository.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file misaem-1.0.0.tar.gz.
File metadata
- Download URL: misaem-1.0.0.tar.gz
- Upload date:
- Size: 44.1 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.9.23
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
3d9130fbe8fdb61ef70723833c0f01ac277cb96a4062e3bfeaef1c1ef624ebbf
|
|
| MD5 |
424165bac6a33a059a3d558ee37a757a
|
|
| BLAKE2b-256 |
e6431c88eefe3c919f025afd64e7748face7070fe3d1d2d42dadf940511335a1
|
File details
Details for the file misaem-1.0.0-py3-none-any.whl.
File metadata
- Download URL: misaem-1.0.0-py3-none-any.whl
- Upload date:
- Size: 10.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.9.23
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
61edf45ade5a5c8021221889ae232a7fb520714e3006a5b4c111b6d7afd7068f
|
|
| MD5 |
a7dc3a20d81b23b4a66e114d52942a95
|
|
| BLAKE2b-256 |
ec0d731e18c473cd13a281aa58a12aada6f1ace491f1f38db7e480cc67e6bdb7
|