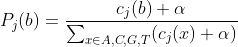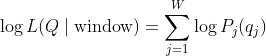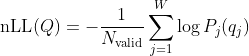Tools for consensus sequence assembly and frameshift correction (mixedAssembly)
Project description
Viral Mixed Assembly Pipeline
This repository provides tools to:
- Run a mixed assembly pipeline (
run_mixed_assembly.py) that merges IRMA and ABACAS information, applies quality control at the sliding-window level, and restores IRMA-specific insertions relative to the reference. - Build empirical priors (
build_priors.py) from large multiple-sequence alignments, used later to evaluate windows during mixed assembly. - Remove frameshifts from an alignment with the reference (
remove_frameshifts.py) - Provide supporting functions via utils scripts.
Installation
pip install mixedassembly
CLI usage
# Create priors
mixedassembly build-priors --input sequences.fasta --ref REF_ID --output priors.parquet
# Run mixed assembly
mixedassembly run-mixed-assembly --input alignment.aln --ref REF_ID --prior priors.parquet --output_dir results
# Correct frameshifts
mixedassembly remove-frameshifts --aln consensus.aln --out corrected.fasta
🚀 Main Script: run_mixed_assembly.py
This is the entrypoint of the pipeline. It creates a mixed consensus sequence by combining IRMA and ABACAS outputs, both aligned to a reference sequence, but only after performing quality control (QC) at the window level.
🔑 Inputs
-
--input→ path to an alignment file (.aln) containing at least:- the reference sequence
- the IRMA consensus sequence
- the ABACAS consensus sequence
-
--ref→ ID of the reference sequence in the alignment. -
--prior→ path to a priors table (.parquet) generated withbuild_priors.py. -
--output_dir→ directory to save the results.
🧪 Workflow
- Filter alignment by reference coordinates → all sequences are trimmed to the reference coordinate system.
- Create overlapping windows → sliding windows across the genome (size and overlap defined in the priors file).
- Window QC →
- Check if ABACAS window is more informative than IRMA, and if it passes some basic qc assumptions (<50 Ns,only 1 or 2 "fragments")
- For each valid window,compute the normalized negative log-likelihood (nLL) of the ABACAS sequence against the prior distribution. Thus, evaluating how rare is the substitution pattern in the window compared to the prior information.
- Compare to the
nLL_p95threshold in the priors table. - If ABACAS passes QC → keep ABACAS bases for that window.
- Otherwise → keep IRMA bases.
- Consensus construction → merge windows into a full consensus sequence.
- Insertion restoration → insertions that IRMA had relative to the reference (lost when filtering by reference coordinates) are reinserted into the final consensus.
- QC reporting → compute coverage, substitutions, and insertion metrics comparing the final mixed consensus to IRMA.
📦 Outputs
The script produces three files inside --output_dir:
-
Mixed consensus FASTA
- File:
<basename>-MIX_ASSEMBLY.fasta - Contains the final consensus sequence after merging and reinserting insertions.
- File:
-
Window QC trace (CSV)
- File:
windows_trace.csv - One row per window, recording:
start,end→ genomic coordinates.MISSING_IRMA,MISSING_ABACAS→ counts of missing bases.ABACAS_MORE_INFO→ whether ABACAS has fewer missing bases than IRMA.ABACAS_FRAGMENTS→ fragmentation level of ABACAS in this window (keep: 0 < n fragments < 3 ).WINDOW_PRIOR_nLL_p95→ threshold from priors.WINDOW_SCORE_nLL→ score of ABACAS in this window.WINDOW_QC_PASSED→ True/False decision.
- File:
-
Consensus QC summary (JSON)
- File:
qc.json - Provides overall metrics comparing the IRMA consensus and the mixed consensus:
IRMA_COVERAGE→ % of genome covered in IRMA.MIXED_COVERAGE→ % of genome covered in mixed consensus.IRMA_SUBSTITUTIONS→ substitutions vs. reference in IRMA.MIXED_SUBSTITUTIONS→ substitutions vs. reference in mixed consensus.EXPECTED_SUBSTITUTIONS→ expected number of substitutions, extrapolated from IRMA.OBS-EXP_SUBSTITUTIONS→ difference between observed and expected substitutions.N_INSERTIONS→ number of insertions added back.TOTAL_INSERTIONS_LENGTH→ total inserted length.INSERTIONS→ list of insertions with their coordinates.
- File:
▶️ Example run
python run_mixed_assembly.py
--input /path/to/250694-RSVWGS.aln
--ref RSV_BD
--prior /path/to/RSVBD_win100_ovlp50_priors.parquet
--output_dir results
This will generate:
results/250694-RSVWGS-MIX_ASSEMBLY.fastaresults/windows_trace.csvresults/qc.json
🛠 Script: build_priors.py
This script creates empirical priors (overlapped windows) from a large multiple sequence alignment.
These priors are later used by run_mixed_assembly.py to evaluate windows.
🔑 Inputs
-i / --input→ aligned FASTA file with multiple sequences.-r / --ref→ ID of the reference sequence.-o / --output→ output file (.parquet).--win→ window size (default: 100).--overlap→ overlap size (default: 10).
▶️ Example run
python build_priors.py
-i alignment.fasta
-r ReferenceID
-o priors.parquet
--win 100
--overlap 10
📦 Output
A .parquet file with one row per window, containing:
start,end→ window coordinates.nLL_p95,nLL_p99→ empirical thresholds.profile→ base probability distributions for each position in the window.
🧮 Methodology (build_priors.py)
1. Probability distributions per position
For each window of size W bases (e.g., W = 100), and for each position j within that window, we compute the probability of observing each nucleotide:
Where:
= number of sequences with base
at position
.
= pseudocount (Laplace smoothing, default
) to avoid zero probabilities.
- Bases
Nare ignored in the counts.
This gives a per-position categorical distribution.
2. Log-likelihood of a sequence in a window
Given a query sequence 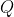
For each valid (non-N) position 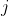
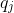
The normalized negative log-likelihood (nLL) is:
Where:
= number of positions in the window where
has a non-
Nbase.
Smaller nLL values indicate sequences more likely under the empirical profile.
3. Empirical priors
To characterize "normal variation" for each window:
- Score all sequences from the alignment against the window profile.
- Collect the distribution of nLL values.
- Extract percentiles (e.g., 95th and 99th) to serve as thresholds.
Thus, for each window we store:
- The distribution (profile).
- Empirical thresholds:
nLL_p95andnLL_p99.
A new sequence can later be compared:
- If
nLL < nLL_p95→ typical. - If
nLL > nLL_p99→ unusually variable, possibly unreliable region.
� Supporting utils
Several utility scripts provide reusable functions for both processes:
-
utils.py → basic alignment and scoring functions:
load_alignment,extract_ref_positions,sliding_windows,score_window.
-
utils_mixed_assembly.py → additional helpers for mixed assembly:
- missingness and fragmentation counts,
- insertion handling,
- QC calculations,
- consensus merging,
- window evaluation wrapper.
These modular functions keep the pipeline clean and reusable.
🧹 Script: remove_frameshifts.py
This script is used to detect and mask problematic gaps in a consensus sequence aligned against a reference.
Frameshifts appear when deletions are not multiples of 3 or when gaps are too long, which can shift the reading frame.
The algorithm replaces those regions with Ns (and optionally adds padding on the sides), preventing errors in downstream analyses.
🔑 Inputs
--aln→ MSA FASTA file (must include reference and consensus).--ref-index→ index of the reference in the alignment (default:0).--cons-index→ index of the consensus in the alignment (default:1).--min-gap→ minimum gap length (multiple of 3) that triggers aggressive masking (default:15).--pad→ number of additional bases to mask around the gap (default:0).--out→ output FASTA file with the corrected consensus.
📦 Output
- FASTA file with the masked consensus → long gaps or gaps not multiple of 3 are replaced by Ns.
▶️ Example run
Assume we have an alignment example.aln.fasta with two sequences:
- index
0: reference - index
1: preliminary consensus
python remove_frameshifts.py \
--aln example.aln.fasta \
--ref-index 0 \
--cons-index 1 \
--min-gap 12 \
--pad 0 \
--out consensus_fixed.fasta
🔍 Example input (simplified):
Reference
ATGGCTTACG CTGGA CTG
Preliminary consensus
ATGGCTTACG-TGGA------G
Here we see:
- A deletion of 1 bases → not a multiple of 3 → potential frameshift.
- A deletion of 6 bases → a multiple of 3 and < min gap → leave gap.
📤 Output (consensus_fixed.fasta):
consensus_masked ATGGCTTACGNTGGA---G
Thus, frameshifts are corrected by masking with Ns, preserving the reading frame validity.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file mixedassembly-0.1.3.tar.gz.
File metadata
- Download URL: mixedassembly-0.1.3.tar.gz
- Upload date:
- Size: 20.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.18
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
869cdd75f69eaa25f16e1a225fad301187fd2ab25cfa0a4563b03feffd5014cf
|
|
| MD5 |
0c482e9cb87ee2aeb66e1aed3b60132e
|
|
| BLAKE2b-256 |
dd7647d757f63a331bc7eee3e98de1748661b3d752c2ff04d53af04b25132d70
|
File details
Details for the file mixedassembly-0.1.3-py3-none-any.whl.
File metadata
- Download URL: mixedassembly-0.1.3-py3-none-any.whl
- Upload date:
- Size: 19.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.18
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
786df26f85f3b2e6b43500023ba12d93951809cf757b03dbd73afd31baf986ed
|
|
| MD5 |
1e63f2213908403f0ff70f8826cc137b
|
|
| BLAKE2b-256 |
ac4d8b5f8f86e14feb4cb74fc620ebfc55c357cea63d2afe50859667ec088741
|