MNE-BIDS: Organizing MEG, EEG, and iEEG data according to the BIDS specification and facilitating their analysis with MNE-Python
Project description
MNE-BIDS
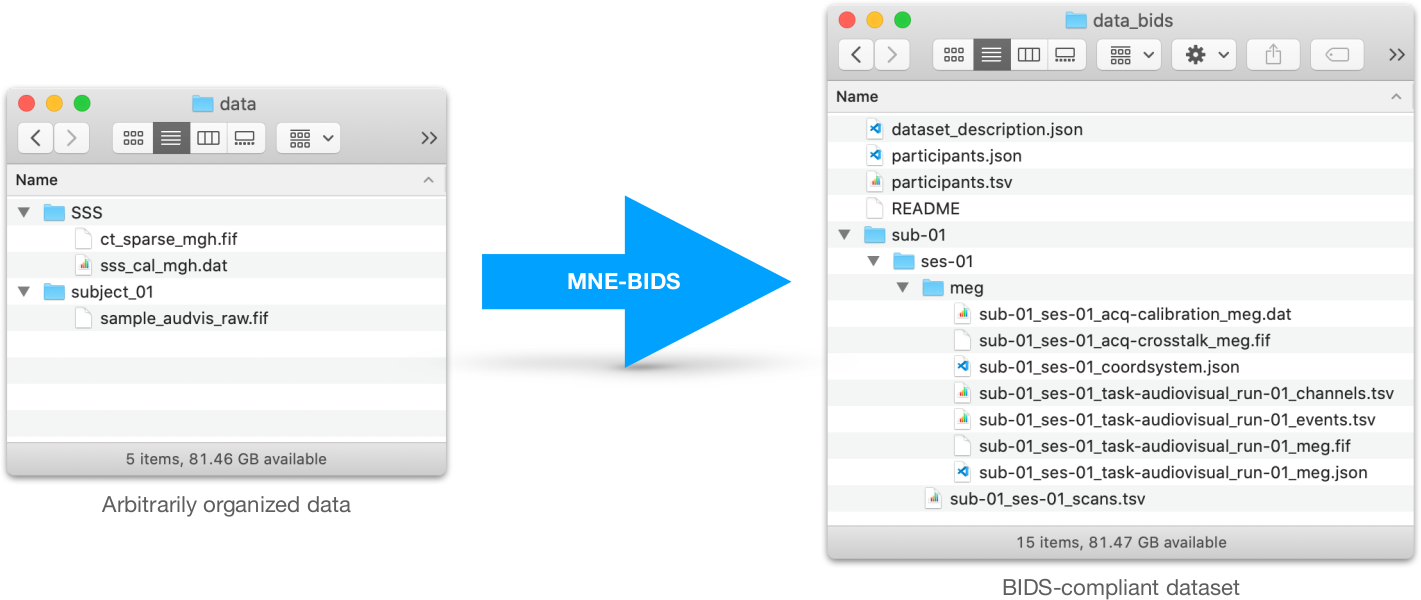
MNE-BIDS is a Python package that allows you to read and write BIDS-compatible datasets with the help of MNE-Python.
Why?
MNE-BIDS links BIDS and MNE-Python with the goal to make your analyses faster to code and more robust, and to facilitate data and code sharing with co-workers and collaborators.
Having your data in BIDS format also allows the use of automated pipelines, for example mne-bids-pipeline.
How?
The documentation can be found under the following links:
- for the stable release
- for the latest (development) version
Getting Help
For any usage questions, please post to the MNE Forum.
Be sure to add the mne-bids tag to your question.
Citing
If you use MNE-BIDS in your work, please cite our publication in JOSS:
Appelhoff, S., Sanderson, M., Brooks, T., Vliet, M., Quentin, R., Holdgraf, C., Chaumon, M., Mikulan, E., Tavabi, K., Höchenberger, R., Welke, D., Brunner, C., Rockhill, A., Larson, E., Gramfort, A., & Jas, M. (2019): MNE-BIDS: Organizing electrophysiological data into the BIDS format and facilitating their analysis. Journal of Open Source Software, 4:1896. DOI: 10.21105/joss.01896
Please also cite one of the following papers to credit BIDS, depending on which data type you used:
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file mne_bids-0.18.0.tar.gz.
File metadata
- Download URL: mne_bids-0.18.0.tar.gz
- Upload date:
- Size: 140.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9db6bbeabc8b465ce17c2356ad11606b871a0116054866a2413a9b313421134a
|
|
| MD5 |
3ad768f9511b80ef3e4b20b1352c820e
|
|
| BLAKE2b-256 |
b69dbe59dca5fdfa720bb12bc0d3457ba576f7f48f41f70becb1d26aa23d50d0
|
Provenance
The following attestation bundles were made for mne_bids-0.18.0.tar.gz:
Publisher:
release.yml on mne-tools/mne-bids
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
mne_bids-0.18.0.tar.gz -
Subject digest:
9db6bbeabc8b465ce17c2356ad11606b871a0116054866a2413a9b313421134a - Sigstore transparency entry: 777297069
- Sigstore integration time:
-
Permalink:
mne-tools/mne-bids@9139214b9377c2d079f9a45f93049ff3b6c7ae51 -
Branch / Tag:
refs/tags/v0.18.0 - Owner: https://github.com/mne-tools
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yml@9139214b9377c2d079f9a45f93049ff3b6c7ae51 -
Trigger Event:
release
-
Statement type:
File details
Details for the file mne_bids-0.18.0-py3-none-any.whl.
File metadata
- Download URL: mne_bids-0.18.0-py3-none-any.whl
- Upload date:
- Size: 178.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
2c3ef03b99f92717cdf426a1a68e4279644ff9e34576a52db14529a0abc539bc
|
|
| MD5 |
01a1a6a7d21e99870f33240399100e52
|
|
| BLAKE2b-256 |
d72f61cdb5a0e3fdbca99ee7d4275ba0e43b1e84dd866bca7a7de8cff30dece6
|
Provenance
The following attestation bundles were made for mne_bids-0.18.0-py3-none-any.whl:
Publisher:
release.yml on mne-tools/mne-bids
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
mne_bids-0.18.0-py3-none-any.whl -
Subject digest:
2c3ef03b99f92717cdf426a1a68e4279644ff9e34576a52db14529a0abc539bc - Sigstore transparency entry: 777297088
- Sigstore integration time:
-
Permalink:
mne-tools/mne-bids@9139214b9377c2d079f9a45f93049ff3b6c7ae51 -
Branch / Tag:
refs/tags/v0.18.0 - Owner: https://github.com/mne-tools
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yml@9139214b9377c2d079f9a45f93049ff3b6c7ae51 -
Trigger Event:
release
-
Statement type: