python package for in silico peptide design and QSAR studies
Project description





modlAMP
This is a Python package that is designed for working with peptides, proteins or any amino acid sequence of natural amino acids. It incorporates several modules, like descriptor calculation (module descriptors) or sequence generation (module sequences). For basic instructions how to use the package, see Usage section of this README or the documentation.
Installation
Quick note: modlAMP supports Python 3 since version 4. Use with Python 2.7 is deprecated.
For the installation to work properly, pip needs to be installed. If you’re not sure whether you already have pip, type pip --version in your terminal. If you don’t have pip installed, install it via sudo easy_install pip.
There is no need to download the package manually to install modlAMP. In your terminal, just type the following command:
pip install modlamp
To update modlamp to the latest version, run the following:
pip install --upgrade modlamp
Usage
This section gives a quick overview of different capabilities of modlAMP. For a detailed description of all modules see the module documentation.
Importing modules
After installation, you should be able to import and use the different modules like shown below. Type python or ipython in your terminal to begin, then the following import statements:
>>> from modlamp.sequences import Helices >>> from modlamp.descriptors import PeptideDescriptor >>> from modlamp.database import query_database
Generating Sequences
The following example shows how to generate a library of 1000 sequences out of all available sequence generation methods:
>>> from modlamp.sequences import MixedLibrary >>> lib = MixedLibrary(1000) >>> lib.generate_sequences() >>> lib.sequences[:10] ['VIVRVLKIAA','VGAKALRGIGPVVK','QTGKAKIKLVKLRAGPYANGKLF','RLIKGALKLVRIVGPGLRVIVRGAR','DGQTNRFCGI','ILRVGKLAAKV',...]
These commands generated a mixed peptide library comprising of 1000 sequences.
The module sequences incorporates different sequence generation classes (random, helices etc.). For documentation thereof, consider the docs for the module modlamp.sequences.
Calculating Descriptor Values
Now, different descriptor values can be calculated for the generated sequences: (see Generating Sequences)
How to calculate the Eisenberg hydrophobic moment for given sequences:
>>> from modlamp.descriptors import PeptideDescriptor, GlobalDescriptor >>> desc = PeptideDescriptor(lib.sequences,'eisenberg') >>> desc.calculate_moment() >>> desc.descriptor[:10] array([[ 0.60138255],[ 0.61232763],[ 0.01474009],[ 0.72333858],[ 0.20390763],[ 0.68818279],...]
Global descriptor features like charge, hydrophobicity or isoelectric point can be calculated as well:
>>> glob = GlobalDescriptor(lib.sequences) >>> glob.isoelectric_point() >>> glob.descriptor[:10] array([ 10.09735107, 8.75006104, 12.30743408, 11.26385498, ...]
Auto- and cross-correlation type functions with different window sizes can be applied on all available amino acid scales. Here an example for the pepCATS scale:
>>> pepCATS = PeptideDescriptor('sequence/file/to/be/loaded.fasta', 'pepcats')
>>> pepCATS.calculate_crosscorr(7)
>>> pepCATS.descriptor
array([[ 0.6875 , 0.46666667, 0.42857143, 0.61538462, 0.58333333,
Many more amino acid scales are available for descriptor calculation. The complete list can be found in the documentation for the modlamp.descriptors module.
Plotting Features
We can also plot the calculated values as a boxplot, for example the hydrophobic moment:
>>> from modlamp.plot import plot_feature
>>> D = PeptideDescriptor('sequence/file/to/be/loaded.fasta', 'eisenberg') # Eisenberg hyrophobicity scale
>>> D.calculate_moment()
>>> plot_feature(D.descriptor,y_label='uH Eisenberg')
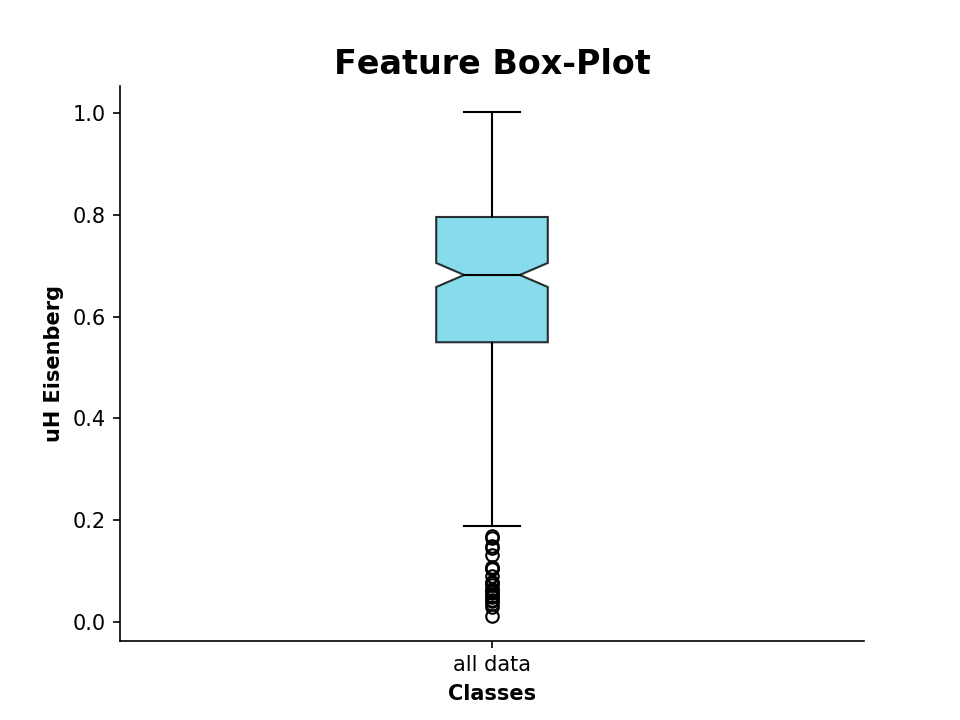
We can additionally compare these descriptor values to known AMP sequences. For that, we import sequences from the APD3, which are stored in the FASTA formatted file APD3.fasta:
>>> APD = PeptideDescriptor('/Path/to/file/APD3.fasta', 'eisenberg')
>>> APD.calculate_moment()
Now lets compare the values by plotting:
>>> plot_feature([D.descriptor, APD.descriptor], y_label='uH Eisenberg', x_tick_labels=['Library', 'APD3'])
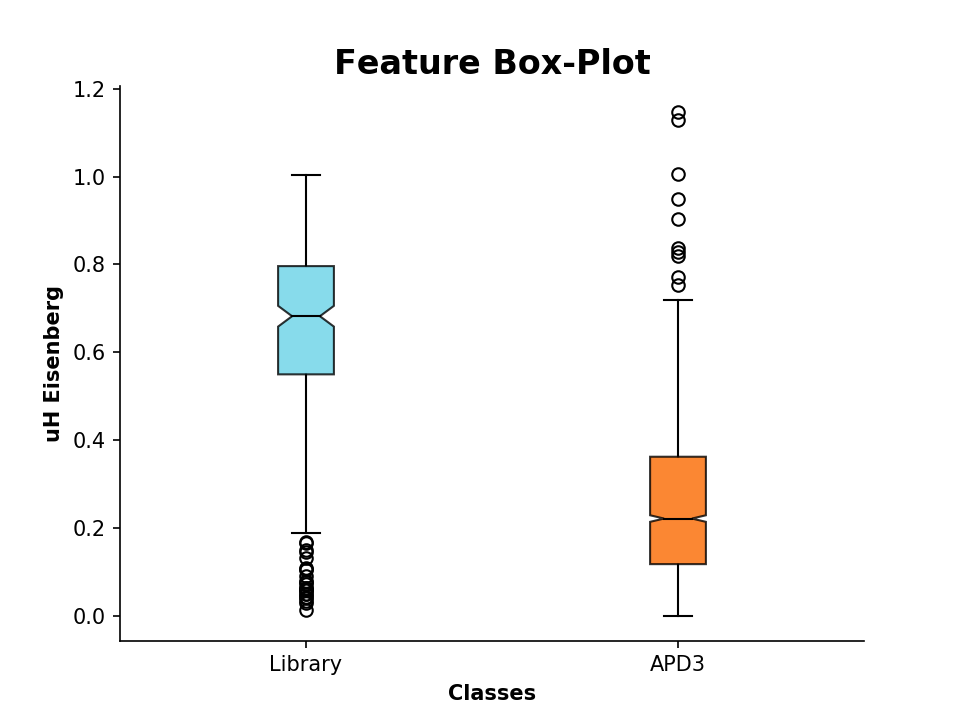
It is also possible to plot 2 or 3 different features in a scatter plot:
- Example:
2D Scatter Plot
>>> from modlamp.plot import plot_2_features
>>> A = PeptideDescriptor('/Path/to/file/class1&2.fasta', 'eisenberg')
>>> A.calculate_moment()
>>> B = GlobalDescriptor('/Path/to/file/class1&2.fasta')
>>> B.isoelectric_point()
>>> target = [1] * (len(A.sequences) / 2) + [0] * (len(A.sequences) / 2)
>>> plot_2_features(A.descriptor, B.descriptor, x_label='uH', y_label='pI', targets=target)
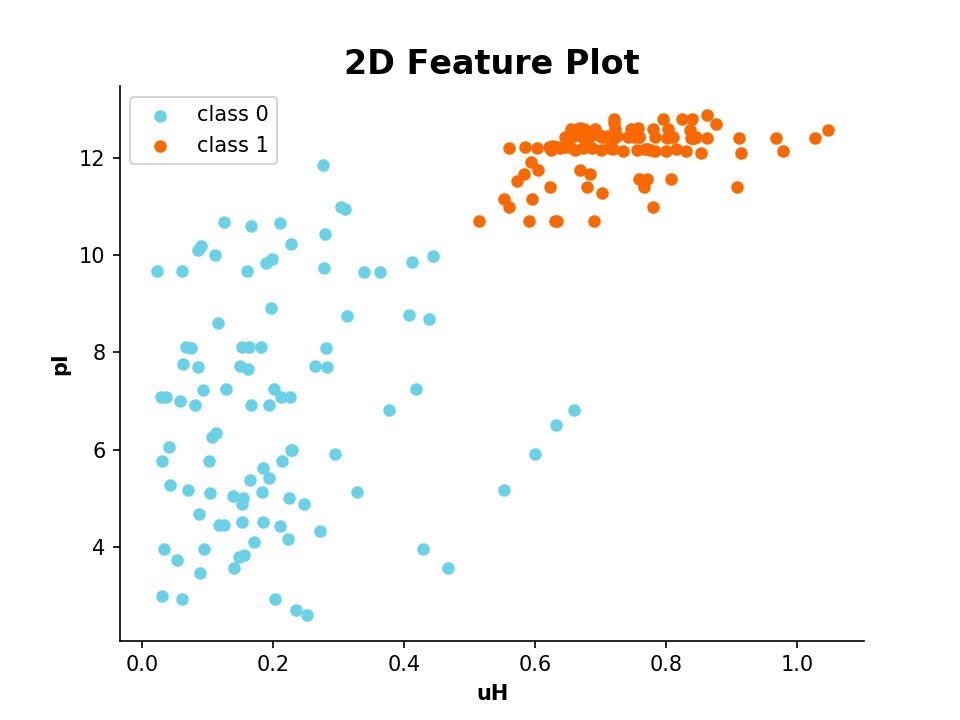
- Example:
3D Scatter Plot
>>> from modlamp.plot import plot_3_features >>> B = GlobalDescriptor(APD.sequences) >>> B.isoelectric_point() >>> B.length(append=True) # append descriptor values to afore calculated >>> plot_3_features(APD.descriptor, B.descriptor[:, 0], B.descriptor[:, 1], x_label='uH', y_label='pI', z_label='len')
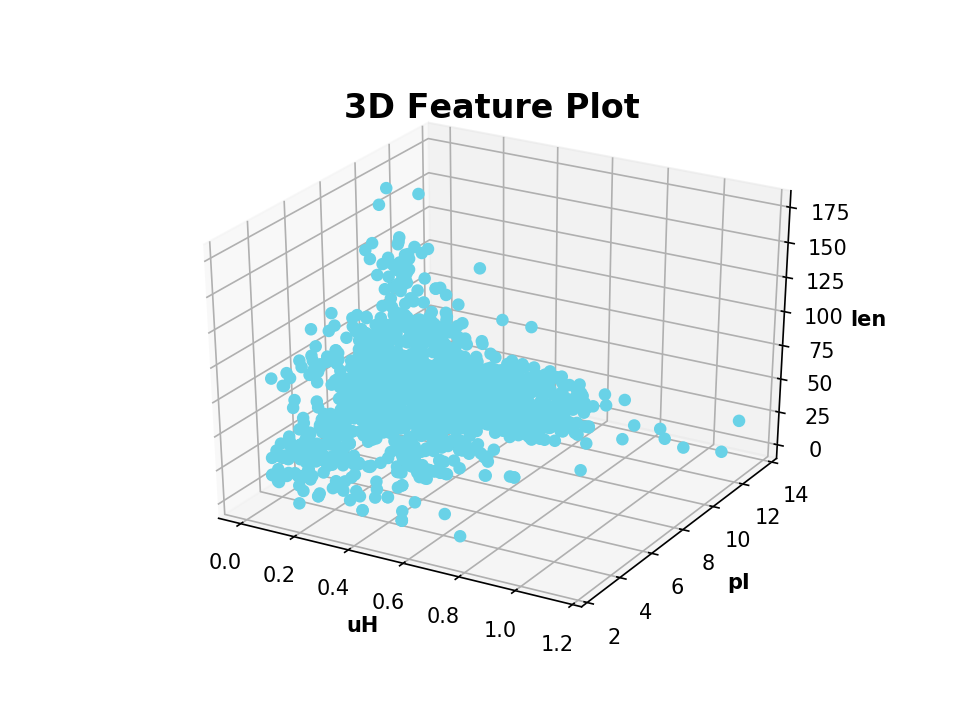
- Example:
Helical Wheel Plot
>>> from modlamp.plot import helical_wheel
>>> helical_wheel('GLFDIVKKVVGALGSL', moment=True)

Further plotting methods are available. See the documentation for the modlamp.plot module.
Database Connection
Peptides from the two most prominent AMP databases APD and CAMP can be directly scraped with the modlamp.database module.
For downloading a set of sequences from the APD database, first get the IDs of the sequences you want to query from the APD website. Then proceed as follows:
>>> query_apd([15, 16, 17, 18, 19, 20]) # download sequences with APD IDs 15 to 20 ['GLFDIVKKVVGALGSL','GLFDIVKKVVGAIGSL','GLFDIVKKVVGTLAGL','GLFDIVKKVVGAFGSL','GLFDIAKKVIGVIGSL','GLFDIVKKIAGHIAGSI']
The same holds true for the CAMP database:
>>> query_camp([2705, 2706]) # download sequences with CAMP IDs 2705 & 2706 ['GLFDIVKKVVGALGSL','GLFDIVKKVVGTLAGL']
modlAMP also hosts a module for connecting to your own database on a private server. Peptide sequences included in any table in the database can be downloaded.
Sequences (stored in a column named sequence) from the personal database can then be queried as follows:
>>> from modlamp.database import query_database
>>> query_database('my_experiments', ['sequence'], configfile='./modlamp/data/db_config.json')
Password: >? ***********
Connecting to MySQL database...
connection established!
['ILDSSWQRTFLLS','IKLLHIF','ACFDDGLFRIIKFLLASDRFFT', ...]
Loading Prepared Datasets
For AMP QSAR models, different options exist of choosing negative / inactive peptide examples. We assembled several datasets for classification tasks, that can be read by the modlamp.datasets module. The available datasets can be found in the documentation of the modlamp.datasets module.
- Example:
AMPs vs. transmembrane regions of proteins
>>> from modlamp.datasets import load_AMPvsTM >>> data = load_AMPvsTM() >>> data.keys() ['target_names', 'target', 'feature_names', 'sequences']
The variable data holds four different keys, which can also be called as its attributes. The available attributes for load_helicalAMPset() are target_names (target names), target (the target class vector), feature_names (the name of the data features, here: ‘Sequence’) and sequences (the loaded sequences).
- Example:
>>> data.target_names # class names
array(['TM', 'AMP'], dtype='|S3')
>>> data.sequences[:5] # sequences
[array(['AAGAATVLLVIVLLAGSYLAVLA', 'LWIVIACLACVGSAAALTLRA', 'FYRFYMLREGTAVPAVWFSIELIFGLFA', 'GTLELGVDYGRAN',
'KLFWRAVVAEFLATTLFVFISIGSALGFK'], dtype='|S100')
>>> data.target # corresponding target classes
array([0, 0, 0, 0, 0 .... 1, 1, 1, 1])
Analysing Wetlab Circular Dichroism Data
The modlule modlamp.wetlab includes the class modlamp.wetlab.CD to analyse raw circular dichroism data from wetlab experiments. The following example shows how to load a raw datafile and calculate secondary structure contents:
>>> cd = CD('/path/to/your/folder', 185, 260) # load all files in a specified folder
>>> cd.names # peptide names read from the file headers
['Pep 10', 'Pep 10', 'Pep 11', 'Pep 11', ... ]
>>> cd.calc_meanres_ellipticity() # calculate the mean residue ellipticity values
>>> cd.meanres_ellipticity
array([[ 260. , -266.95804196],
[ 259. , -338.13286713],
[ 258. , -387.25174825], ...])
>>> cd.helicity(temperature=24., k=3.492185008, induction=True) # calculate helical content
>>> cd.helicity_values
Name Solvent Helicity Induction
Peptide1 T 100.0 3.823
Peptide1 W 26.16 0.000
Peptide2 T 76.38 3.048
Peptide2 W 25.06 0.000 ...
The read and calculated values can finally be plotted as follows:
>>> cd.plot(data='mean residue ellipticity', combine=True)
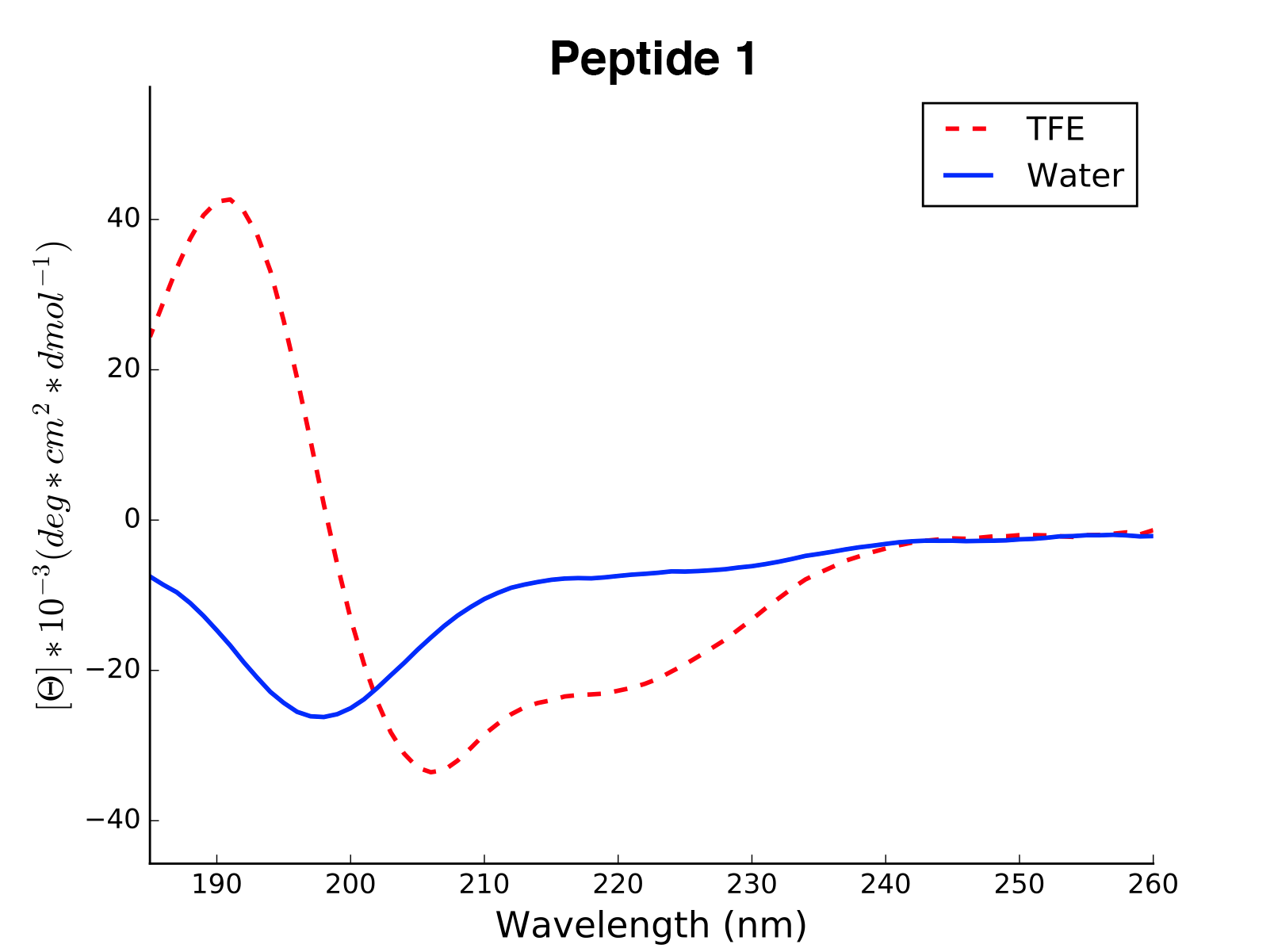
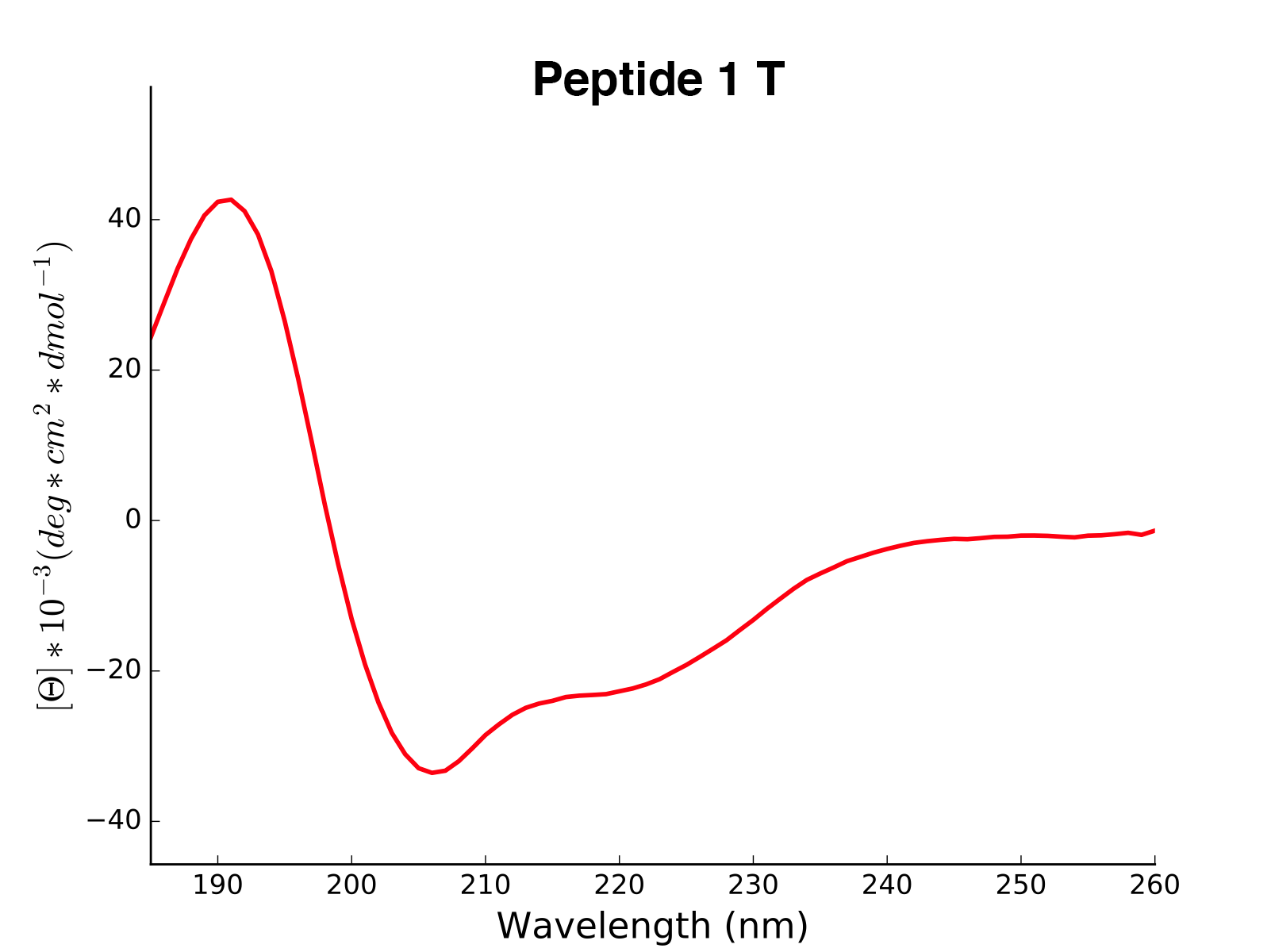
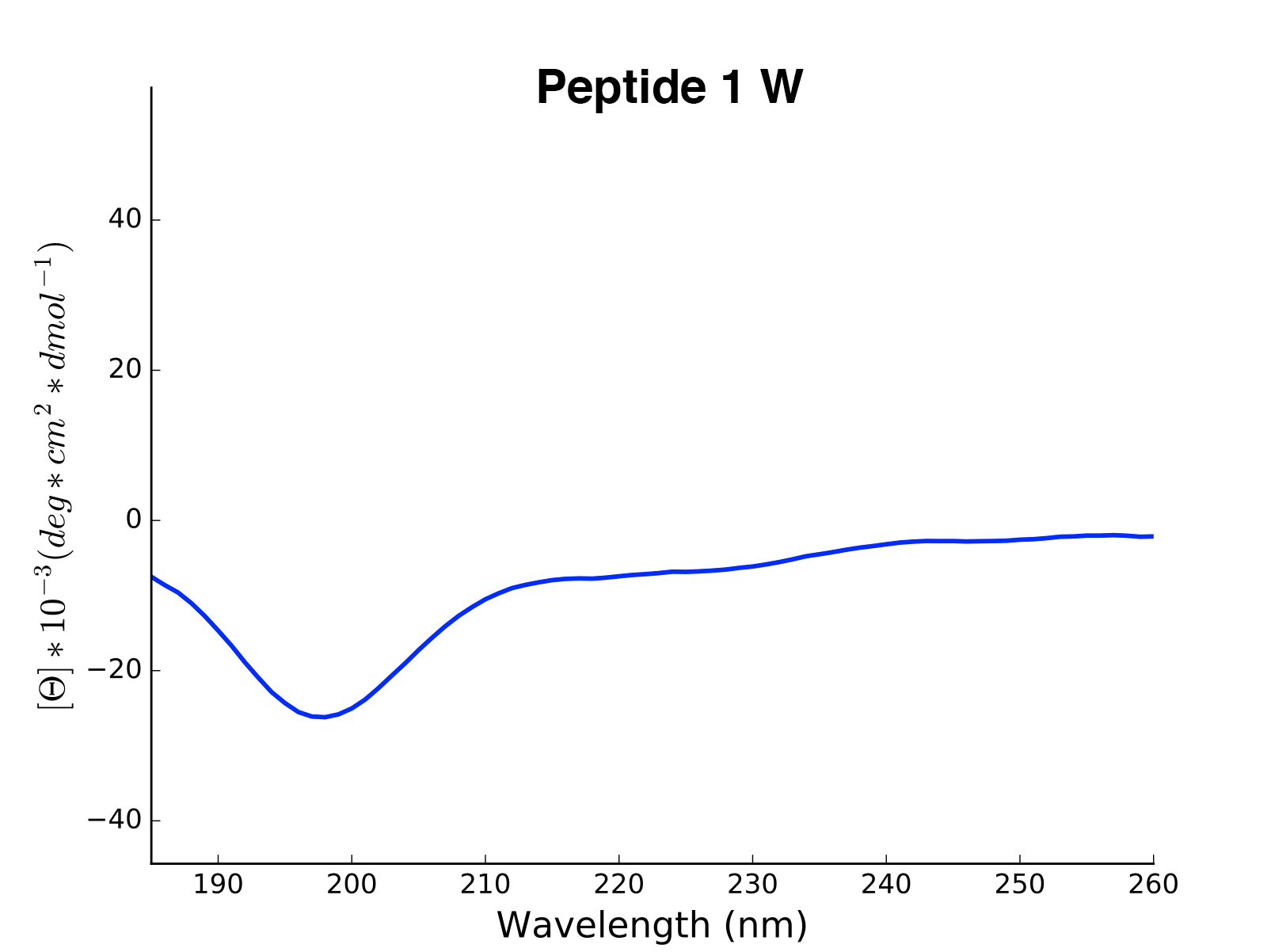
Analysis of Different Sequence Libraries
The modlule modlamp.analysis includes the class modlamp.analysis.GlobalAnalysis to compare different sequence libraries. Learn how to use it with the following example:
>>> lib # sequence library with 3 sub-libraries
array([['ARVFVRAVRIYIRVLKAFAKL', 'IRVYVRIVRGFGRVVRAYARV', 'IRIFIRIARGFGRAIRVFVRI', ..., 'RGPCFLQVVD'],
['EYKIGGKA', 'RAVKGGGRLLAG', 'KLLRIILRGARIIIRGLR', ..., 'AKCLVDKK', 'VGGAFALVSV'],
['GVHLKFKPAVSRKGVKGIT', 'RILRIGARVGKVLIK', 'MKGIIGHTWKLKPTIPSGKSAKC', ..., 'GRIIRLAIKAGL']], dtype='|S28')
>>> lib.shape
(3, 2000)
>>> from modlamp.analysis import GlobalAnalysis
>>> analysis = GlobalAnalysis(lib, names=['Lib 1', 'Lib 2', 'Lib 3'])
>>> analysis.plot_summary()
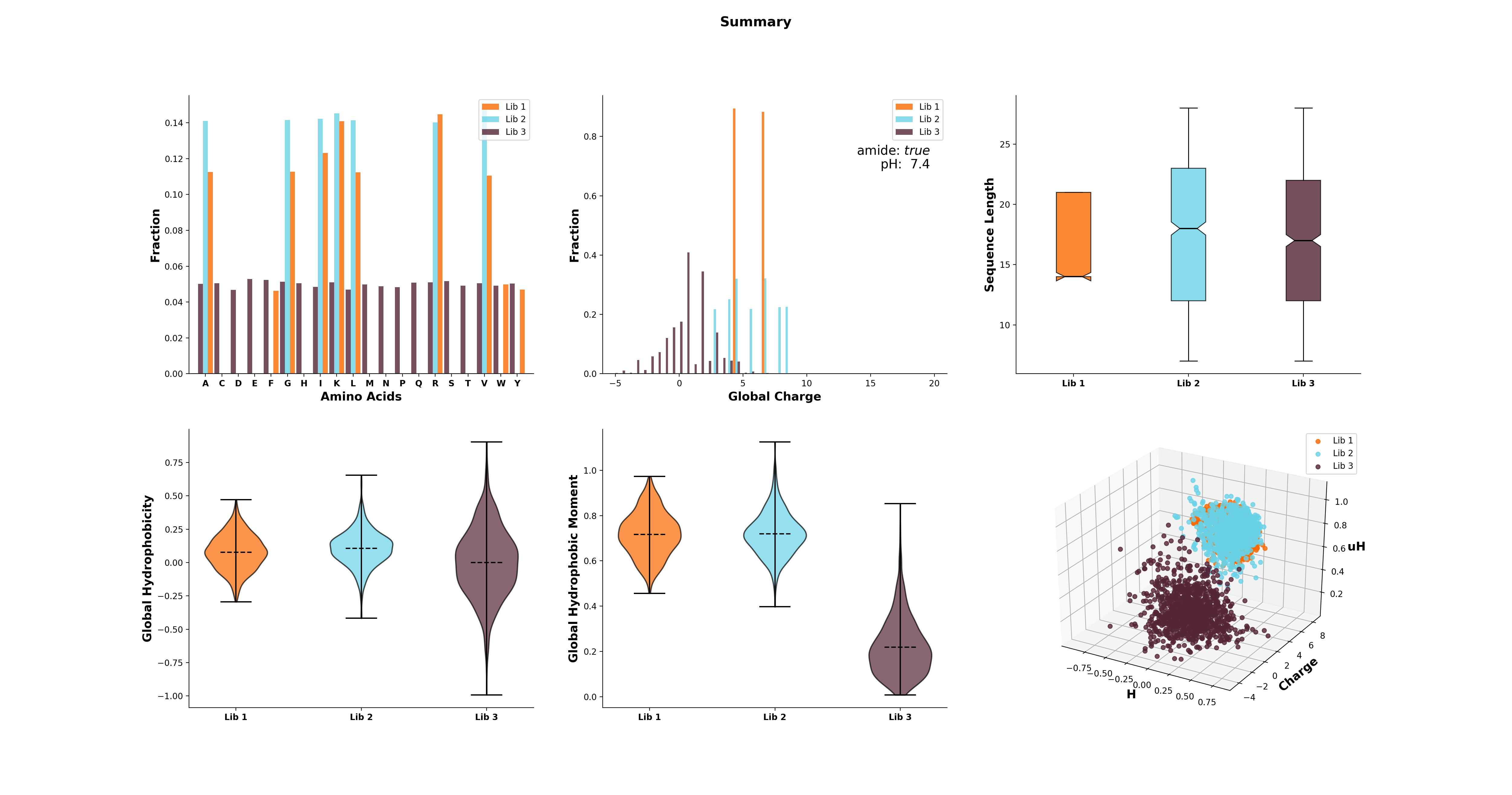
Documentation
A detailed documentation of all modules is available from the modlAMP documentation website.
Contributing
We are open to contributions! Please clone the repo, create your own branch and, once you’re done, create a pull request.
Also, please use the test cases in the tests folder to ensure that your code is working properly. To run tests, use the following command in the root directory of the package:
python -m unittest discover tests
Citing modlAMP
If you are using modlAMP for a scientific publication, please cite the following paper:
Müller A. T. et al. (2017) modlAMP: Python for anitmicrobial peptides, Bioinformatics 33, (17), 2753-2755, DOI:10.1093/bioinformatics/btx285.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file modlamp-4.3.2.tar.gz.
File metadata
- Download URL: modlamp-4.3.2.tar.gz
- Upload date:
- Size: 2.7 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.9.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cc36c29ef154a1d3b454dd452206d0261489118ec612cf4747bf0c9d0289bedd
|
|
| MD5 |
e7bd6a9122699ff43bab1b3df1a71293
|
|
| BLAKE2b-256 |
6233ffee2c8e9470a2747e7872f1a749684b11024e0308f413fc6f6541baf58c
|
File details
Details for the file modlamp-4.3.2-py3-none-any.whl.
File metadata
- Download URL: modlamp-4.3.2-py3-none-any.whl
- Upload date:
- Size: 176.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.9.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
404ce261e5a2e312dd6a5c52c1087f7d91f1c4b2f4227700160731926a841c19
|
|
| MD5 |
df6a416ed772a1aa2b3450e9101b1a74
|
|
| BLAKE2b-256 |
e83c0432b145158117de34544159527c3c6cb2be29cd44a072b95b0b87604387
|











