MolCompass Visualization Tool: Visualize your Chemical Space
Project description
The MolCompass View application
Introduction

MolCompassViewer is a part of the MolCompass project. It is a tool that provides a pretrained parametric t-SNE model for chemical space visualization and the visual validation of QSAR/QSPR models.
Installation
The package can be installed: pip install molcompview
Run
Demo
To run the endocrine receptor example described in the manuscript (coming soon), you may do so by executing the following steps.
molcompview --demo
Then, wait some time for the calculation of coordinates. After that stage, the browser will open, and you can view the interactive visualization of the chemical space.
Your own dataset
To run on your own dataset, just use:
molcompview <input.csv>
Usage
MolCompassViewer intelligently identifies the types of columns within the CSV file, selecting an operational mode based on the presence of specific column types, primarily the molecular structures encoded as SMILES strings.
Operational Modes
- STRUCTURE ONLY: Activated when only the SMILES column is identified. Focuses on visualizing molecular structures, omitting additional features like color layers and QSAR/QSPR model analyses.
- ALTERNATIVE: Triggered when additional categorical or numerical columns are found alongside the SMILES column, excluding the Ground Truth and Probabilities columns. Focuses on exploring the chemical space of compounds, which does not apply to the analysis of models. Users can customize point colors in the visualization based on selected properties.
- FULL: Dedicated to the visual analysis of binary QSAR/QSPR models. Available when the CSV file comprises SMILES strings along with Ground Truth and predicted probabilities columns, unlocking access to exclusive features for visualizing binary QSAR/QSPR models.

Applicability domain analysis
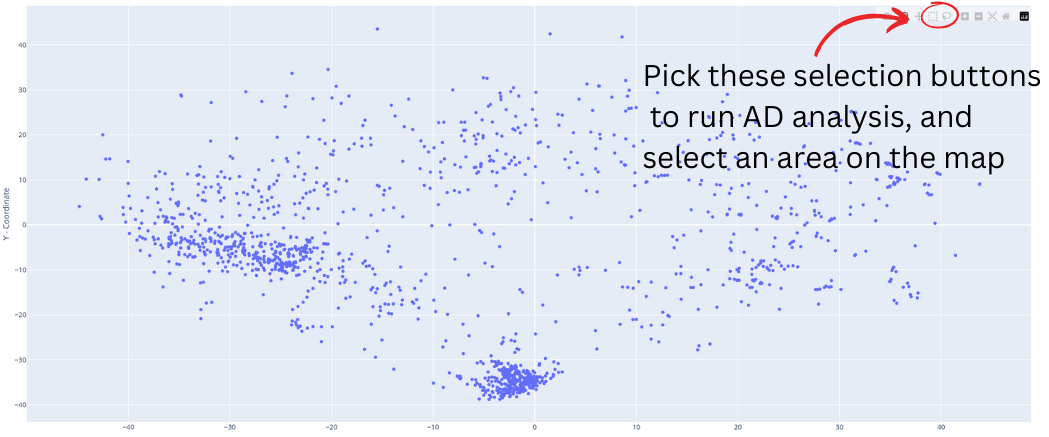
The applicability domain analysis is possible only in FULL mode for binary classification models. To run this tool, first select a tool from the top left corner of the chemical map, then select an area of interest and release. A new frame will open on the right, displaying statistical parameters and charts calculated exclusively for the selected compounds.
Screenshots

DOI for the package version
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file molcompview-0.1.7.tar.gz.
File metadata
- Download URL: molcompview-0.1.7.tar.gz
- Upload date:
- Size: 39.4 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
381cdd96ecbb0458f23fdfd75e4d05514f7f71bb127eb434acdb734ca54f31e6
|
|
| MD5 |
3c67dcee909cc81baede8933199c54b0
|
|
| BLAKE2b-256 |
4f1037b1fa40aadaf0fc1b5e383a55fac75c4f4bc94b8e39c42c82199c3d3a03
|
File details
Details for the file molcompview-0.1.7-py3-none-any.whl.
File metadata
- Download URL: molcompview-0.1.7-py3-none-any.whl
- Upload date:
- Size: 38.9 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
09a11cba2e656a1151d7369c75f22bd63263de041bcd93e78e9d88a3ee6efafa
|
|
| MD5 |
ac949a468bced1f1db894f8976e1554b
|
|
| BLAKE2b-256 |
0d60cc0d38babd716bba51659f5aa60eed9745bbd19f9ad885efd531756ef3a1
|