MOLPIPx: Permutationally Invariant Polynomials in JAX
Project description
Differentiable version of Permutationally Invariant Polynomial (PIP) models in JAX and Rust.
MOLPIPx is a JAX-based library that provides an implementation of PIP models compatible with,
- FLAX: Neural network library.
- GPU friendly.
- Fully differentiable.
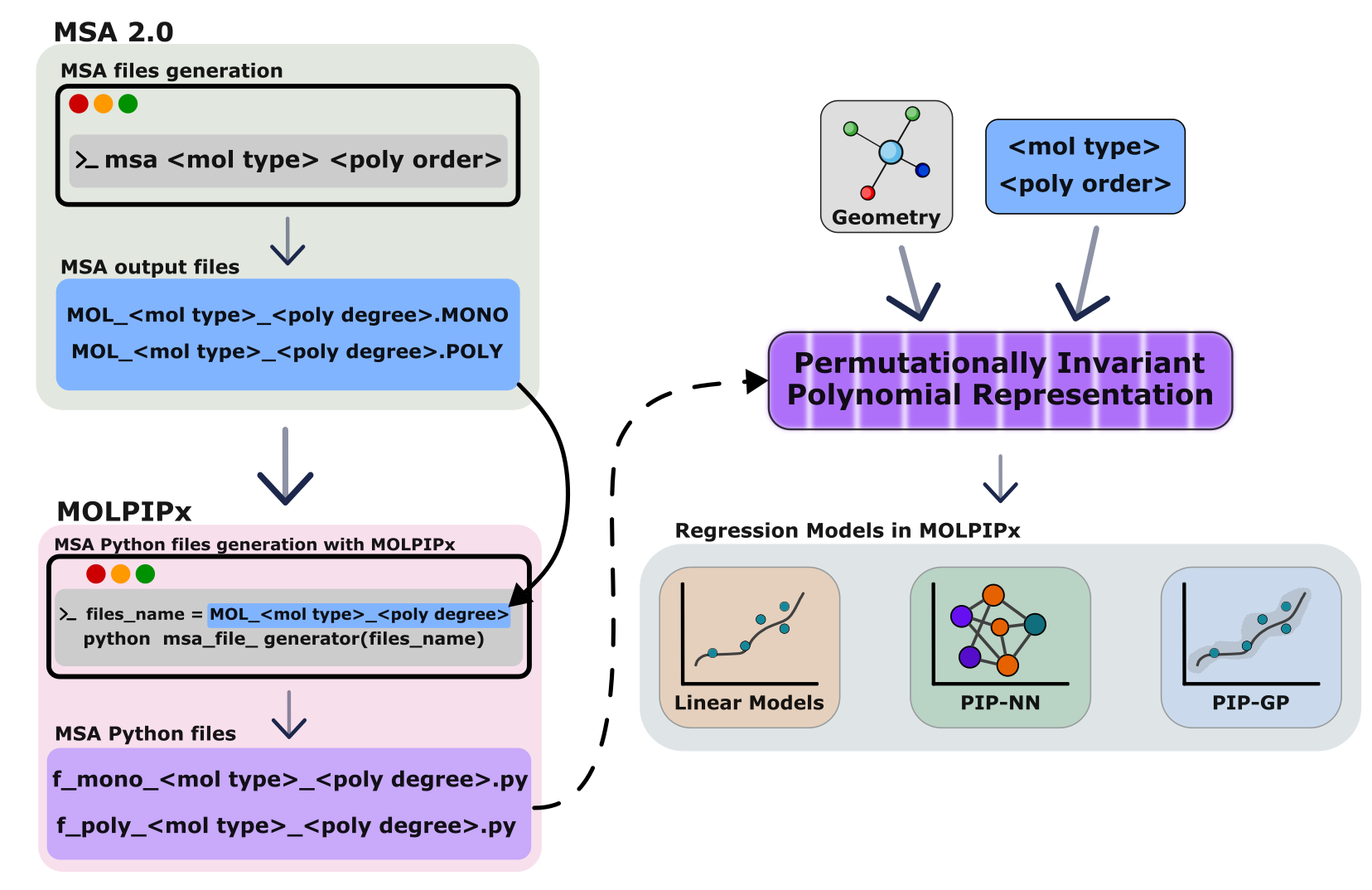
MSA files to JAX
This library translates the MSA files, specifically the _file_.MONO and _file_.POLY files to the corresponding JAX version, _file_mono.py and _file_poly.py.
The MSA files must be generated before, for more information please see https://github.com/szquchen/MSA-2.0
MSA References:
- Xie, Z.; Bowman, J.M. Permutationally Invariant Polynomial Basis for Molecular Energy Surface Fitting via Monomial Symmetrization. J. Chem. Theory Comput. 2010, 6, 26-34.
Installation
MOLPIPx requires Python 3.10 or later. We recommend installing it in a virtual environment to avoid conflicts.
python -m venv molpipx
source molpipx/bin/activate
The latest stable release is available through PyPI:
pip install molpipx
To install the latest development version from source, clone the source code from the GitHub and install with pip from local file:
git clone https://github.com/ChemAI-Lab/molpipx.git
cd molpipx
pip install .
MSA-JAX files generation
MOLPIPx package includes msa_file_generator, which translates monomial and polynomial files from MSA to JAX and Rust for molecules.
Check out an example on generating msa files
from molpipx import msa_file_generator
head_files = 'MOL_<info>_<deg>'
path = '<path_to_the_files>'
label = '<file_label>'
msa_file_generator(head_files, path, label)
The structure of the library is kept simple, as each molecular system could need individual elements.
Models
MOLPIPx incorporated PIPs with three main regression models, i.e., linear regression, neural networks and Gaussian processes. This library leverages two main automatic differentiation engines, JAX for The Python version and Enzyme-AD for the Rust version improve the simulation of a wide range of chemical systems.
Rust Version
The Rust version makes use of std::autodiff, an experimental feature of Rust which is currently in the process of upstreaming. While upstreaming is in progress, you will need to build our custom fork of Rust which already includes autodiff. Instruction for how to do so are available here. Once upstreaming completed, you will be able to use any nightly Rust version. This tracking issue shows the progress in upstreaming the remaining autodiff pieces.
Tutorials
Check out our tutorials to get started with MOLPIPx. These tutorials define inputs for different regression approaches, train machine learning models with or without forces, and make predictions.
- Linear regression with permutationally invariant polynomials (Linear PIP)
- Anisotropic linear regression with permutationally invariant polynomials (Anisotropic Linear PIP)
- Permutationally Invariant Polynomial Neural Networks (PIP-NN)
- Permutationally Invariant Polynomial Gaussian Process (PIP-GP)
📖 Documentation
Full documentation, including installation instructions, API reference, and examples, is available at:
👉 https://chemai-lab.github.io/molpipx/
Reference
@article{molpipx,
author = {Drehwald, Manuel S. and Jamali, Asma and Vargas-Hernández, Rodrigo A.},
title = {MOLPIPx: An end-to-end differentiable package for permutationally invariant polynomials in Python and Rust},
journal = {The Journal of Chemical Physics},
volume = {162},
number = {8},
pages = {084115},
year = {2025},
month = {02},
issn = {0021-9606},
doi = {10.1063/5.0250837},
url = {https://doi.org/10.1063/5.0250837},
eprint = {https://pubs.aip.org/aip/jcp/article-pdf/doi/10.1063/5.0250837/20416060/084115_1_5.0250837.pdf},
}
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file molpipx-0.1.1.tar.gz.
File metadata
- Download URL: molpipx-0.1.1.tar.gz
- Upload date:
- Size: 54.1 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1619686db4e3b4154889eb1de71093a3093d073e8df7c1513468821b6ba46f97
|
|
| MD5 |
e7d5cfbf75a9cd08789722b6ee5f4430
|
|
| BLAKE2b-256 |
030f4a6ec96756864a077563e1acb7d841a72132518deb3f2526f9be0ff0a849
|
File details
Details for the file molpipx-0.1.1-py3-none-any.whl.
File metadata
- Download URL: molpipx-0.1.1-py3-none-any.whl
- Upload date:
- Size: 54.4 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a7c8e7d3398c9b40e7baf68103415cb600b4a33a8200e293b40498e1be0eeede
|
|
| MD5 |
989ab1da279940c4993ab39e333fa778
|
|
| BLAKE2b-256 |
416158e7cba5311e137a260775d47ee1279ee92a4191a83a8367ff0875a106cd
|