Cross-platform desktop ROI editor for microscopy, histology, and fluorescence imaging
Project description

Montaris-X
Cross-platform ROI editor for scientific microscopy images
About
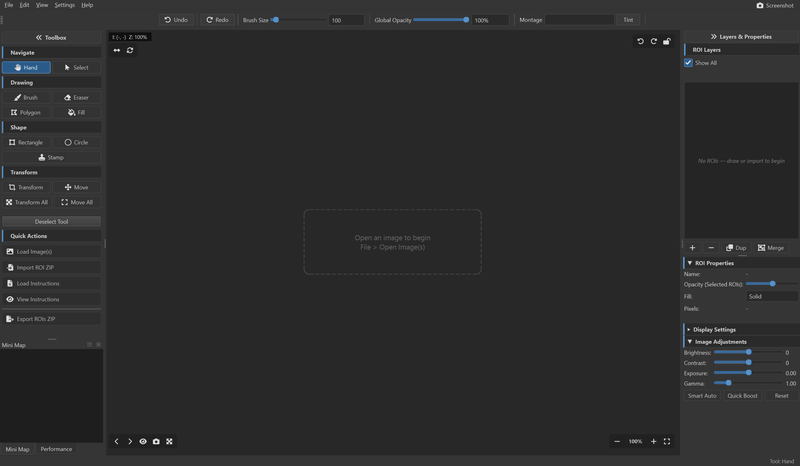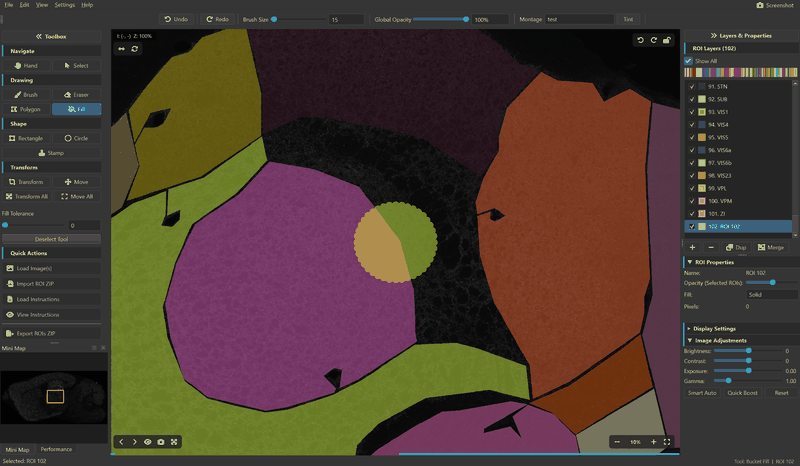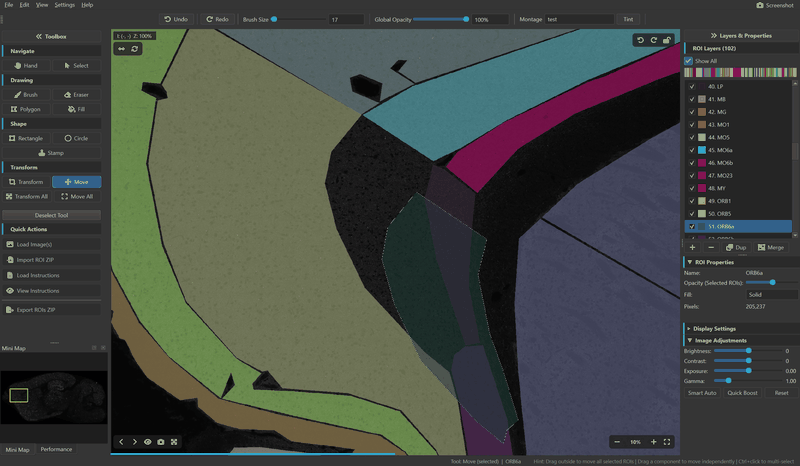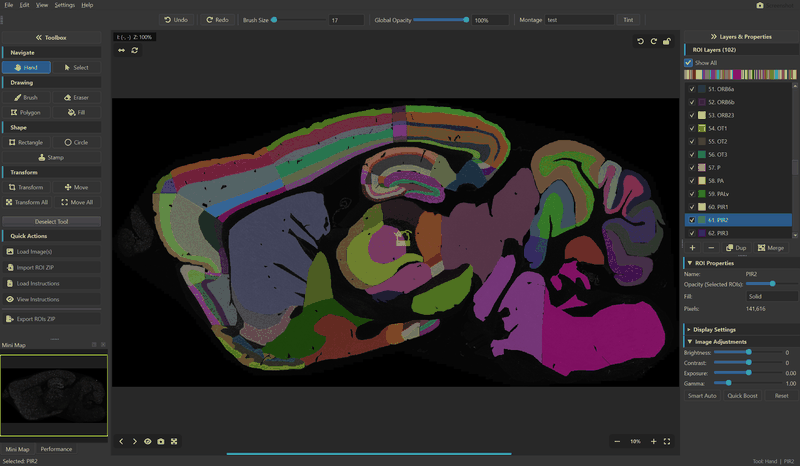
Montaris-X is a native desktop application for drawing, editing, and managing Regions of Interest (ROIs) on scientific microscopy images. It is built for researchers who need precise, pixel-level annotation of histological sections, brain atlases, fluorescence micrographs, and other large-format scientific imagery.
Why Montaris-X?
- Purpose-built for science — handles 16/32-bit TIFFs, multi-channel composites, and images with thousands of ROIs without breaking a sweat.
- ImageJ-compatible — import and export
.roi/.zipfiles directly from ImageJ/FIJI workflows. - No cloud, no account — runs entirely offline on your machine. Your data never leaves your computer.
- Cross-platform — native support for Windows, macOS, and Linux.
Installation
Option 1: Download Pre-built Executable (Recommended)
Download the latest release for your platform from the Releases page:
| Platform | File | Notes |
|---|---|---|
| Windows | montaris-x-windows.exe |
Double-click to run. Signed binary. |
| macOS | montaris-x-macos |
Run chmod +x montaris-x-macos && ./montaris-x-macos |
| Linux | montaris-x-linux |
Run chmod +x montaris-x-linux && ./montaris-x-linux |
macOS note: If macOS shows "unidentified developer", right-click the file and select Open, then click Open in the dialog.
Linux note: You may need to install EGL and D-Bus libraries:
sudo apt install libegl1 libxkbcommon0 libdbus-1-3
Option 2: Install from PyPI
pip install montaris-x
Then launch:
montaris
Option 3: Install from Source
git clone https://github.com/digin1/Montaris-X.git
cd Montaris-X
pip install -e .
Then launch:
python main.py
# or
python -m montaris
# or
montaris
Optional: GPU Acceleration
Install Numba for JIT-compiled rendering on large images:
pip install montaris-x[accel]
# or, if installing from source:
pip install numba
Features
Image Support
- Scientific formats — TIFF (8/16/32-bit, multi-channel), PNG, JPEG, BMP
- Multi-channel composites — split and view individual channels with false-color display
- Image adjustments — brightness, contrast, exposure, gamma with Smart Auto and Quick Boost
- Tiled rendering — LOD-based rendering for smooth navigation on large images
- Flip & Rotate — non-destructive horizontal flip and 90-degree rotation
ROI Editing
- Drawing tools — Brush, Eraser, Rectangle, Circle, Polygon, Stamp, Bucket Fill
- Transform & Move — scale, rotate, and reposition ROIs with connected-component awareness
- Multiple ROI layers — 20+ distinct colors with per-ROI opacity and fill modes (Solid, Outline, Both)
- Multi-selection — Ctrl+click to select and edit multiple ROIs simultaneously
- Undo/Redo — full global history plus per-layer undo
- Overlap handling — auto-overlap brush mode and fix-overlaps utility
Import / Export
- Native format — save/load ROI sets as
.npzarchives - ImageJ compatible — import/export
.roifiles and ZIP bundles - PNG masks — import/export binary mask images
- Composited overlay — export the ROI overlay as a single PNG
- Batch instructions — JSON/TXT instruction files for automated workflows
- Drag-and-drop — drop images and ROI ZIPs directly onto the window
Grid Canvas
- Multi-image workspace — define an NxM grid (up to 4x4) to work with multiple images side-by-side
- Independent cells — each cell has its own image, ROIs, undo history, adjustments, and downsample factor
- Click to switch — click any cell to make it the active workspace; all tools and panels follow
- Double-click to maximize — expand a single cell to fill the entire grid; double-click again to restore
- Save All — save session progress for every cell in one action (
Ctrl+Alt+S) - Export All as ZIP — export each cell's ROIs as separate ZIP files named by source image (
Ctrl+Alt+E) - Per-cell downsampling — each cell can have its own downsample factor with correct upscaling on export
Interface
- Dark & Light themes — switch between themes on the fly
- Collapsible sidebars — minimize panels for a distraction-free canvas
- Fullscreen mode — F11 for immersive editing
- Minimap — always-visible overview of the full image
- Performance monitor — real-time FPS and memory usage
- Global opacity slider — adjust overlay transparency from the toolbar
- Session save — auto-save and restore progress across sessions
- Persistent settings — Flip on Load and Rotate on Load are remembered across launches
Keyboard Shortcuts
Click to expand full shortcut reference
File
| Shortcut | Action |
|---|---|
Ctrl+O |
Open Image(s) |
Ctrl+Shift+O |
Load ROI Set |
Ctrl+S |
Save ROI Set |
Ctrl+Shift+S |
Save Progress (Session) |
Ctrl+Alt+S |
Save All Grid Sessions |
Ctrl+E |
Export ROI(s) as PNG |
Ctrl+Alt+E |
Export All Grid Cells as ZIP |
Ctrl+W |
Close Image(s) |
Ctrl+Q |
Quit |
Edit
| Shortcut | Action |
|---|---|
Ctrl+Z |
Undo |
Ctrl+Y / Ctrl+Shift+Z |
Redo |
Ctrl+Alt+Z |
Layer Undo |
Ctrl+Alt+Y |
Layer Redo |
Ctrl+A |
Select All ROIs |
Delete |
Delete Active ROI(s) |
View
| Shortcut | Action |
|---|---|
Ctrl+0 |
Fit to Window |
Ctrl+1 |
Reset Zoom (1:1) |
Ctrl+= |
Zoom In |
Ctrl+- |
Zoom Out |
Ctrl+H |
Flip Horizontal |
Ctrl+R |
Rotate 90 CW |
Ctrl+[ |
Collapse Left Sidebar |
Ctrl+] |
Collapse Right Sidebar |
Ctrl+Shift+M |
Maximize / Restore Grid Cell |
F11 |
Fullscreen |
Ctrl+Shift+P |
Screenshot |
Tools
| Shortcut | Action |
|---|---|
H |
Hand (pan) |
Q |
Select ROI |
B |
Brush |
E |
Eraser |
P |
Polygon |
G |
Bucket Fill |
R |
Rectangle |
C |
Circle |
S |
Stamp |
T |
Transform (selected) |
Shift+T |
Transform All |
V |
Move (selected) |
Shift+V |
Move All |
[ / ] |
Adjust brush size (adaptive steps) |
Space |
Pan (hold) |
Enter |
Finish Polygon |
Escape |
Cancel / Clear selection |
Navigation
| Shortcut | Action |
|---|---|
Scroll wheel |
Zoom in/out |
Middle-click drag |
Pan |
Ctrl+Click |
Toggle ROI selection |
Supported Formats
| Type | Formats |
|---|---|
| Images | TIFF (.tif, .tiff) including 16/32-bit, PNG, JPEG, BMP |
| ROI Sets | NumPy Archive (.npz), ImageJ ROI (.roi), ZIP bundles, PNG masks |
| Instructions | JSON (.json), Text (.txt) |
Building from Source
Running Tests
pip install -e ".[test]"
pytest
Building a Standalone Executable
pip install pyinstaller
pyinstaller --name montaris-x --onefile --windowed --hidden-import montaris --collect-all montaris montaris/__main__.py
The executable will be in the dist/ folder.
Building a Python Package
pip install build
python -m build
This produces .whl and .tar.gz files in dist/ for distribution.
System Requirements
- Python 3.10 or higher (for source/PyPI installation)
- OS: Windows 10+, macOS 11+, or Linux (X11/Wayland with EGL)
- RAM: 4 GB minimum, 8 GB+ recommended for large images
- Dependencies: PySide6, NumPy, SciPy, scikit-image, tifffile, Pillow, psutil
Linux System Dependencies
On Debian/Ubuntu, install the required Qt runtime libraries:
sudo apt install libegl1 libxkbcommon0 libdbus-1-3
Code Signing
Code signing via SignPath.io (certificate by SignPath Foundation) has been applied for and is pending approval.
- Privacy policy: This program does not transfer any information to other networked systems unless specifically requested by the user or the person installing or operating it.
Contributing
Contributions are welcome! Please open an issue or submit a pull request on GitHub.
- Fork the repository
- Create a feature branch (
git checkout -b feature/my-feature) - Commit your changes
- Push to the branch and open a Pull Request
Acknowledgments
Montaris-X is developed in the Centre for Clinical Brain Sciences, University of Edinburgh, with support from the Grant Lab.
Special thanks to:
- Prof Seth Grant — Principal Investigator, Professor of Molecular Neuroscience
- Nicolas Martinez-Wise — Feature design, testing, and bug reports
- Jessica Griffiths — Feature design and testing
- Colin Yuan — Testing and feedback
And everyone in the Grant Lab for their continued feedback and support.
License
Montaris-X is released under the MIT License.
Copyright (c) 2026 Digin Dominic and Montaris-X Contributors
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file montaris_x-2.2.2.tar.gz.
File metadata
- Download URL: montaris_x-2.2.2.tar.gz
- Upload date:
- Size: 795.1 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
45e3b2fb04f9813f4a7a6b01ad3de96408bbdbf514095abf92de155cdc0ef40a
|
|
| MD5 |
b1c835f7a9b495420976302b52ae4729
|
|
| BLAKE2b-256 |
15a8961abbbd19753c86fb337c925af8818e889db394b63db733096be4a96918
|
Provenance
The following attestation bundles were made for montaris_x-2.2.2.tar.gz:
Publisher:
publish.yml on digin1/Montaris-X
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
montaris_x-2.2.2.tar.gz -
Subject digest:
45e3b2fb04f9813f4a7a6b01ad3de96408bbdbf514095abf92de155cdc0ef40a - Sigstore transparency entry: 1235846101
- Sigstore integration time:
-
Permalink:
digin1/Montaris-X@d21c9178ab59d1eb321725c6f8ff622a6e52b29d -
Branch / Tag:
refs/tags/v2.2.2 - Owner: https://github.com/digin1
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@d21c9178ab59d1eb321725c6f8ff622a6e52b29d -
Trigger Event:
push
-
Statement type:
File details
Details for the file montaris_x-2.2.2-py3-none-any.whl.
File metadata
- Download URL: montaris_x-2.2.2-py3-none-any.whl
- Upload date:
- Size: 690.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f4da2d3d9a19423f0d55f1580a6885a638b5fb63fca329f678ff264b74e3f508
|
|
| MD5 |
497e616bf90308e4aef90b9532e5b32a
|
|
| BLAKE2b-256 |
67daeb6802e46f707722462fbc06d4c985f33c8941f9dad9829443f7ea37f36b
|
Provenance
The following attestation bundles were made for montaris_x-2.2.2-py3-none-any.whl:
Publisher:
publish.yml on digin1/Montaris-X
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
montaris_x-2.2.2-py3-none-any.whl -
Subject digest:
f4da2d3d9a19423f0d55f1580a6885a638b5fb63fca329f678ff264b74e3f508 - Sigstore transparency entry: 1235846157
- Sigstore integration time:
-
Permalink:
digin1/Montaris-X@d21c9178ab59d1eb321725c6f8ff622a6e52b29d -
Branch / Tag:
refs/tags/v2.2.2 - Owner: https://github.com/digin1
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@d21c9178ab59d1eb321725c6f8ff622a6e52b29d -
Trigger Event:
push
-
Statement type:















