A package to convert a multiplex image in OME-TIFF file format to a virtual brightfield image such as H&E or IHC.
Project description
multiplex2brightfield
multiplex2brightfield is a Python package designed to convert multiplex imaging data (such as Imaging Mass Cytometry - IMC, CODEX, etc.) into virtual brightfield images, simulating traditional histological stains like Hematoxylin & Eosin (H&E), Immunohistochemistry (IHC), Masson's Trichrome, PAS, and others. The library utilizes the OME-TIFF file format for both input and output, ensuring compatibility with standard bioimaging workflows and preservation of metadata.
Features
- Multiplex to Virtual Brightfield Conversion: Transform multiplex image data into various simulated brightfield stains.
- Multiple Stain Simulations: Supports built-in configurations for H&E, IHC, Masson Trichrome, PAS, Jones Silver, Toluidine Blue, and configurations matching specific scanner outputs (e.g., Aperio CS2, Hamamatsu XR).
- OME-TIFF Input/Output: Reads and writes OME-TIFF files, preserving essential metadata like pixel sizes and channel names.
- Pyramid Generation: Optionally creates multi-resolution OME-TIFF pyramids for efficient viewing of large images.
- Flexible Channel Mapping:
- Uses predefined marker lists for common stains.
- Leverages Large Language Models (ChatGPT, Gemini, Claude) to automatically map markers to stain components based on channel names and stain type.
- Image Processing: Offers options for image enhancement and artifact reduction, including:
- Median filtering.
- Gaussian smoothing.
- Image sharpening.
- Intensity adjustments.
- Per-channel histogram normalization or clipping.
- Reference-based histogram matching.
- AI Enhancement: Includes an optional AI model to enhance the visual quality of the generated brightfield image.
- Large Image Handling: Supports tiled processing and memory mapping (
memmap) for converting very large images that may not fit into RAM. - Resampling: Allows adjusting the output pixel size.
Installation
Install the package using pip:
pip install multiplex2brightfield
Usage
Basic Conversion (Virtual H&E)
from multiplex2brightfield import convert_from_file
# Input and output file paths
input_filename = "path/to/your/input_image.ome.tiff"
output_filename = "path/to/your/output_virtual_HE.ome.tiff"
# This assumes your input OME-TIFF has channel names identifiable
# by the default H&E configuration in configuration_presets.py
convert_from_file(
input_filename=input_filename,
output_filename=output_filename,
stain="H&E", # Specify the desired stain preset
create_pyramid=True # Generate OME-TIFF pyramid
)
print(f"Virtual H&E image saved to {output_filename}")
Advanced Conversion (Virtual Masson Trichrome with AI Enhancement & LLM Mapping)
from multiplex2brightfield import convert_from_file
from matplotlib import pyplot as plt
# Input and output file paths
input_filename = "path/to/your/input_image.ome.tiff"
output_filename = "path/to/your/output_virtual_trichrome.ome.tiff"
# Requires API key for the chosen LLM
# Ensure the corresponding library (openai, google-generativeai, anthropic) is installed
api_key = "YOUR_LLM_API_KEY"
# The configuations file of the stain that the LLM will complete to add the targets
config = {
"name": "Masson Trichrome",
"components": {
"nuclei": {
"color": {"R": 30, "G": 114, "B": 201},
"description": "Uniform nuclear staining using markers that label all nuclei."
},
"muscle": {
"color": {"R": 220, "G": 67, "B": 51},
"description": "Muscle fibers along with their associated cytoplasmic components showing the structure of smooth muscle and myocytes."
},
"collagen": {
"color": {"R": 93, "G": 209, "B": 225},
"description": "Specifically highlights collagen and connective tissue."
},
"erythrocytes": {
"color": {"R": 229, "G": 128, "B": 56},
"description": "Red blood cells."
}
},
"background": {
"color": { "R": 255, "G": 255, "B": 255},
},
}
result = convert_from_file(
input_filename=input_filename,
output_filename=output_filename,
config = config, # Specify config for LLM to complete
# Use an LLM (e.g., Gemini) to determine channel mapping for Trichrome
use_gemini=True,
api_key=api_key,
# Apply AI enhancement
AI_enhancement=True,
# Create pyramid output
create_pyramid=True,
# Optional: Specify output pixel size if different from input
output_pixel_size_x=0.5,
output_pixel_size_y=0.5,
# Optional: Process large images in tiles
process_tiled=True,
tile_size=4096 # Adjust tile size based on memory
)
print(f"Virtual Masson Trichrome image saved to {output_filename}")
plt.figure(figsize=(10,10))
plt.imshow(result[0].transpose(1, 2, 0))
plt.axis('off')
plt.title(f'Brightfield Image')
plt.show()
Key Parameters (convert function)
input_filename(str): Path to the input OME-TIFF multiplex image.output_filename(str): Path for the output virtual brightfield OME-TIFF.stain(str): Name of the stain preset to use (e.g., "H&E", "IHC", "Masson Trichrome"). See configuration_presets.py or use as context for LLMs.use_chatgpt, use_gemini, use_claude(bool): Flags to enable specific LLMs for automatic channel mapping. Requires corresponding API key and library installation.api_key(str): API key for the selected LLM service.config(dict): Optionally provide a custom configuration dictionary instead of using presets or LLMs.AI_enhancement(bool): Apply the AI-based enhancement model. Downloads the model on first use.create_pyramid(bool): Generate a multi-resolution pyramid in the output OME-TIFF.downsample_count(int): Number of pyramid levels to generate.output_pixel_size_x, output_pixel_size_y(float): Specify the desired output pixel size (in units from input metadata, typically µm) for resampling.process_tiled(bool): Process the image in tiles to handle large files.tile_size(int): Size of tiles (e.g., 4096, 8192) for tiled processing.use_memmap(bool): Use memory mapping for temporary storage during tiled processing (useful for very large images).Filter Parameters(median_filter_size, gaussian_filter_sigma, sharpen_filter_amount, etc.): Override default filter settings from the configuration.NormalizationParameters (histogram_normalisation, clip, normalize_percentage_min, normalize_percentage_max, intensity): Override default normalization/intensity settings.
Configuration File Format
The virtual staining conversion uses a configuration dictionary that defines how each stain component is simulated. The configuration is based on the following template:
template = {
"name": "<stain_name>", # e.g., "H&E"
"components": {
"<component_name>": { # e.g., "haematoxylin"
"color": {
"R": 0, # Integer between 0-255, representing the red channel value.
"G": 0, # Integer between 0-255, representing the green channel value.
"B": 0, # Integer between 0-255, representing the blue channel value.
},
"description": "string", # A brief description of the component's role.
"targets": [], # A list of marker or channel names to assign to this component (e.g., markers for nuclei, cytoplasm, etc.).
"intensity": 1.0, # A scaling factor for the expression intensity of this component.
"median_filter_size": 0, # Size of the median filter kernel to reduce noise (0 means no filtering).
"gaussian_filter_sigma": 0, # The standard deviation for Gaussian smoothing (0 means no smoothing).
"sharpen_filter_amount": 0, # Amount of sharpening to apply (0 means no sharpening).
"histogram_normalisation": False, # Whether to apply histogram normalization to the component.
"normalize_percentage_min": 10, # Lower percentile bound for intensity normalization.
"normalize_percentage_max": 90, # Upper percentile bound for intensity normalization.
"clip": None, # Optional value to clip intensity values to a specific range.
},
# Additional components can be added here (e.g., "eosinophilic", "epithelial", "erythrocytes").
},
"background": {
"color": {
"R": 255, # Red channel value for the background.
"G": 255, # Green channel value for the background.
"B": 255, # Blue channel value for the background.
}
},
}
Explanation of the Configuration Fields
name(str): A string that identifies the stain configuration (e.g., "H&E"). This determines the overall naming and may be used to select preset configurations.components(dictionary): A dictionary where each key is the name of a stain component (e.g., "haematoxylin", "eosinophilic", etc.). Each component defines:component_name(str): The name of the component.color(dictionary): A dictionary with keys"R","G", and"B"specifying the color to be used for that component in the output image. Values should be in the range 0–255.description(str): A textual explanation of what the component represents (e.g., nuclear staining, cytoplasmic staining).targets(list of str): An array of strings listing marker or channel names that the component should represent. The conversion function will map the channels from your multiplex image to these targets.intensity(float): A scaling factor (typically 1.0 by default) that controls how strongly the component is rendered.median_filter_size(int): Defines the size of the median filter (if >0) for reducing noise.gaussian_filter_sigma(float): Specifies the sigma for Gaussian filtering, used to smooth the image.sharpen_filter_radius(int): Defines the size of the sharpen filter (if >0).sharpen_filter_amount(float): Determines how much to sharpen the image.histogram_normalisation(bool): A Boolean flag that, when set to True, applies histogram normalization to enhance image contrast.normalize_percentage_minandnormalize_percentage_max(float): Define the lower and upper percentile values used during normalization to adjust the dynamic range.clip(tuple of float, float or None): An optional parameter to clip intensity values within a specified range; if not needed, it remainsNone.
background:color(dictionary): A dictionary with keys"R","G", and"B"defining the background color (typically white: 255, 255, 255).
This configuration template provides a flexible way to customize how different image components are processed and visualized, making it easy to simulate various staining protocols by simply adjusting these parameters.
Examples of the Configuration Values
Below are visual examples of how individual configuration parameters affect the rendered brightfield output. All examples show the same cropped region of the image for direct comparison.
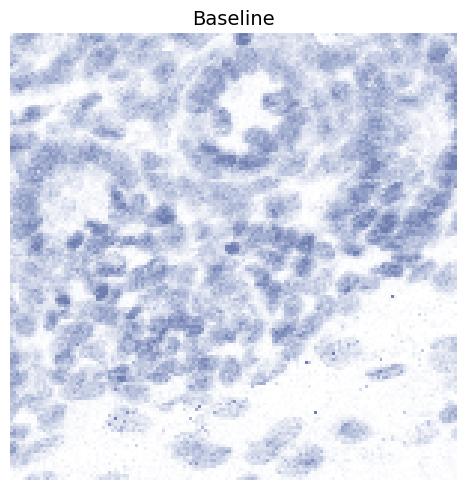
Baseline
This is the baseline configuration where all processing parameters are disabled
(i.e. set to 0, full-range, or False). The only exception is
normalize_percentage_max, which is set to 99. If normalize_percentage_max is set to 100, extreme high-intensity hotspots
dominate the scaling and cause most structural detail to disappear.
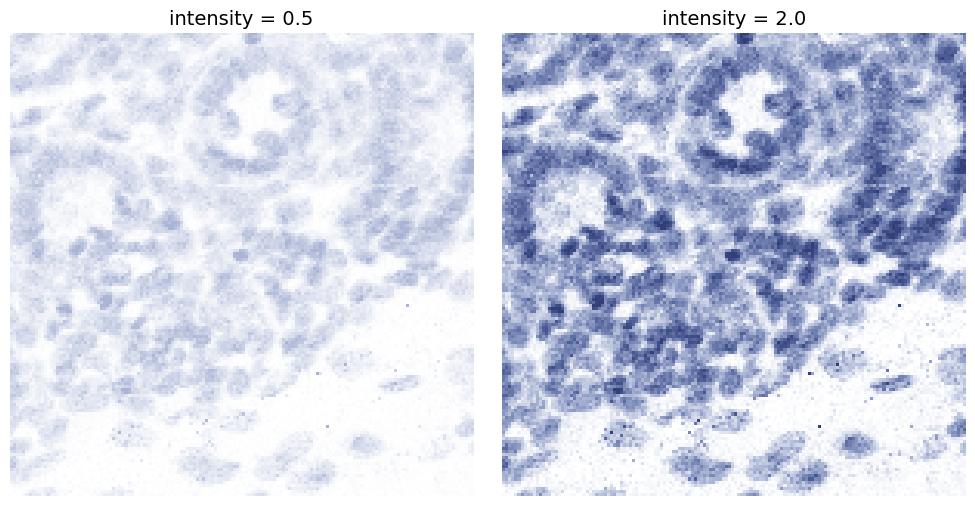
intensity
Controls the overall strength of the stain component. Higher values make the component color stronger, while lower values make it lighter.
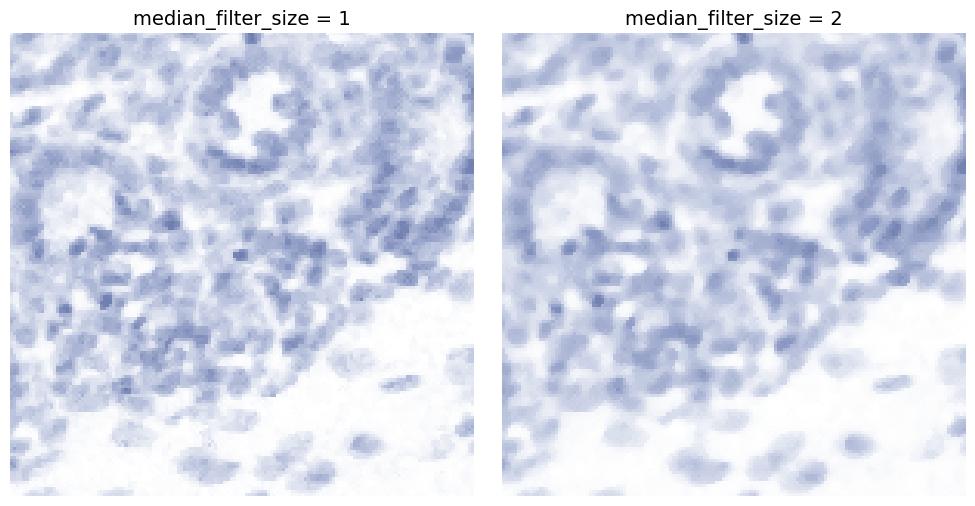
median_filter_size
Applies median filtering, typically useful for removing salt-and-pepper noise.
Higher values increase smoothing while preserving edges better than Gaussian blur in some cases.
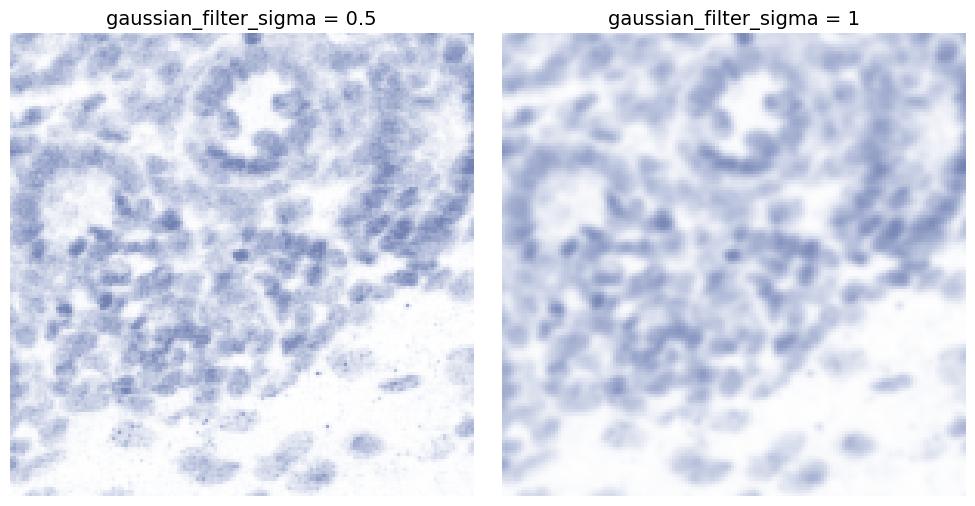
gaussian_filter_sigma
Applies Gaussian smoothing to reduce high-frequency noise.
Higher values produce a more blurred, smoother appearance.
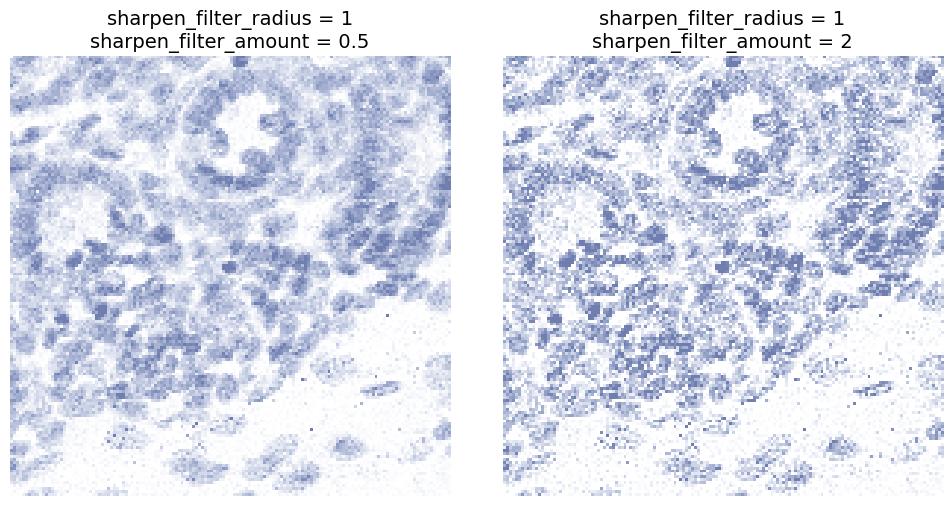
sharpen_filter_radius and sharpen_filter_amount
Sharpens edges and local contrast.
sharpen_filter_radiuscontrols the spatial scale of sharpening.sharpen_filter_amountcontrols how strong the sharpening effect is.
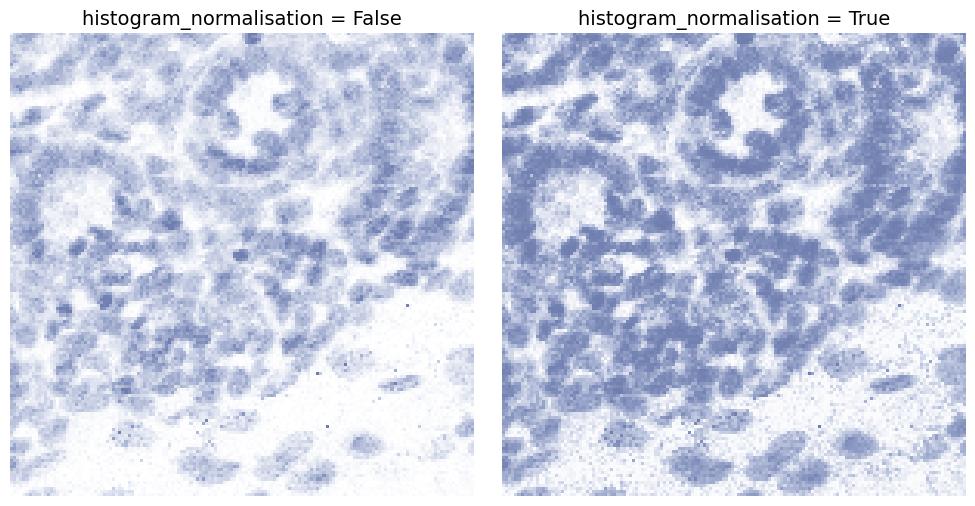
histogram_normalisation
If enabled, normalizes intensities to increase contrast and spread values across a usable range.
This can make faint structures more visible but may also amplify noise.
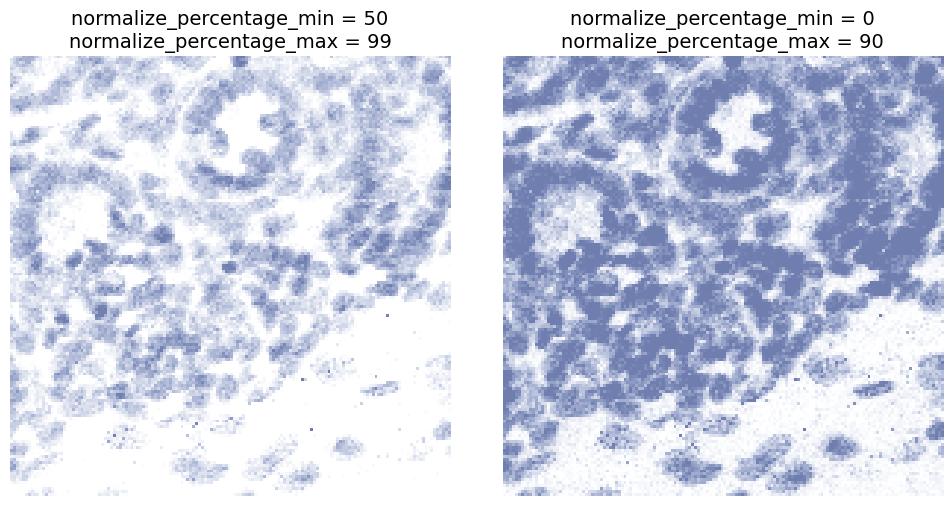
normalize_percentage_min and normalize_percentage_max
Defines the percentile range used for intensity normalization.
- Increasing
normalize_percentage_minsuppresses dim signal. - Decreasing
normalize_percentage_maxcan prevent high value pixles, such as hotspots, from dominating.
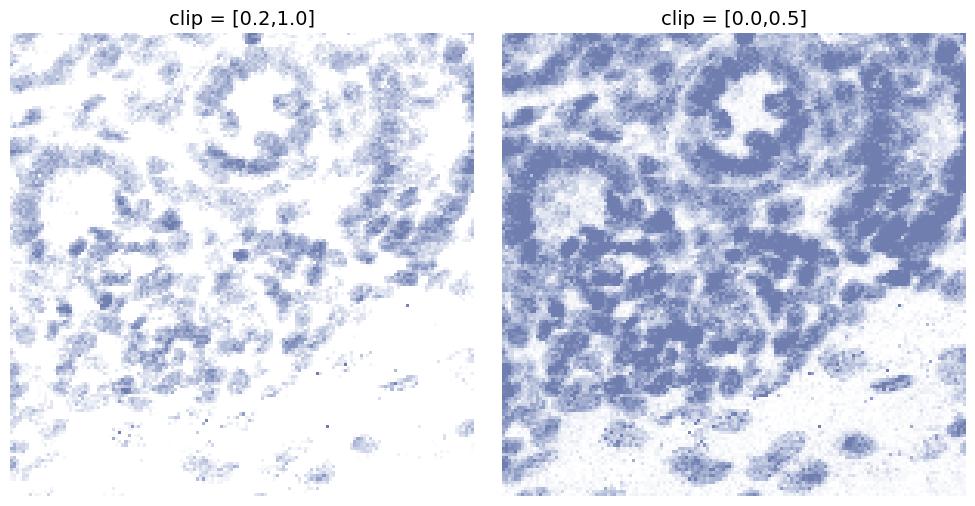
clip
Clips the intensity range to a fixed interval [min, max].
This can be used to:
- suppress very low intensity background signal
- prevent extremely bright pixels from saturating the final output
Dependencies
Key dependencies include:
- numpy
- tifffile
- scikit-image
- SimpleITK
- keras (for AI enhancement)
- csbdeep
- numpy2ometiff
- lxml
- requests (for model download)
- tqdm (for progress bars)
- Optional: openai, google-generativeai, anthropic (for LLM usage)
Contributing
Contributions to multiplex2brightfield are welcome! Feel free to fork the repository, make your changes, and submit a pull request. For major changes, please open an issue first to discuss what you would like to change.
License
This project is licensed under the BSD 3-Clause License - see the LICENSE file for details.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file multiplex2brightfield-0.2.7.tar.gz.
File metadata
- Download URL: multiplex2brightfield-0.2.7.tar.gz
- Upload date:
- Size: 40.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.10.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e98941a1769a1cb0f353dcca063b02d6b4f6eef1f0a45054f4027c8451581531
|
|
| MD5 |
21098f6f0a6edd547b4724c3cdc5cbbe
|
|
| BLAKE2b-256 |
539c59b0929e7eaa192e41d196da8d7bd63c24acd74a750372412823349d9b0b
|
File details
Details for the file multiplex2brightfield-0.2.7-py3-none-any.whl.
File metadata
- Download URL: multiplex2brightfield-0.2.7-py3-none-any.whl
- Upload date:
- Size: 37.9 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.10.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
065b395266e3cef8c99a321869018c24dd2f9acd6aed3c200fd4e1321ba7de01
|
|
| MD5 |
d829d59fb948a42f03421c734bf839d7
|
|
| BLAKE2b-256 |
f4f95822fc8b3de411fb57b633b16ea64f27abbe2fd0ea1216595c2eab048fa5
|