MyeVAE: a multi-modal variational autoencoder for risk profiling of newly diagnosed multiple myeloma
Project description
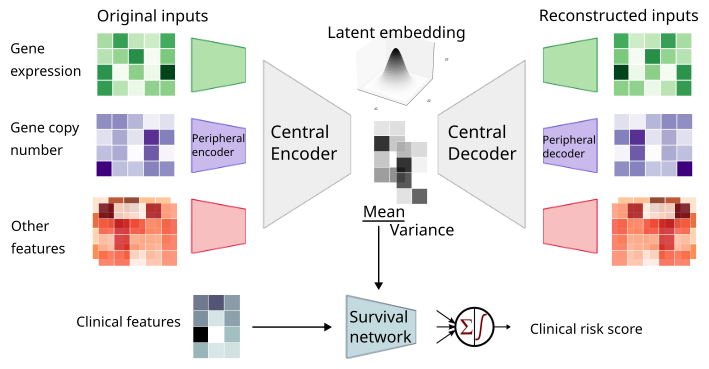
MyeVAE
A variational autoencoder leveraging multi-omics for risk prediction in newly diagnosed multiple myeloma patients.
This Python package contains the code to train and run SHAP on MyeVAE. No GPU is required.
Install
Install through PyPI (requires Python 3.9 or later)
pip install myevae
Step-by-step guide
1. Download data
Download the raw multi-omics dataset and example outputs from the following link:
-
Input datasets are stored at https://myevae.s3.us-east-1.amazonaws.com/example_inputs.tar.gz
-
Example output files are stored at https://myevae.s3.us-east-1.amazonaws.com/example_outputs.tar.gz
Please use the download links sparingly.
If you use the AWS S3 CLI, the bucket ARN is arn:aws:s3:::myevae - you can list its contents and download specific files.
2. (Optional) Feature preprocessing
As there are over 50,000 raw features and contains missing values, this step performs supervised-feature selection, scaling, and imputation.
myevae preprocess \
-e [endpoint: os | pfs] \
-i [/inputs/folder]
Processed files will be stored in train_features_[os|pfs]_processed.csv and valid_features_[os|pfs]_processed.csv.
Preprocessing of features is resource-intensive. Alternatively, proceed to step 3 using the processed features in the example input data (https://myevae.s3.us-east-1.amazonaws.com/example_inputs.tar.gz).
train_features_os_processed.csv
train_features_pfs_processed.csv
valid_features_os_processed.csv
valid_features_pfs_processed.csv
3. Fit model
Place the hyperparameter grid (param_grid.py) in /params/folder.
myevae train \
-e [endpoint: os | pfs] \
-n [model_name] \
-i [/inputs/folder] \
-p [/params/folder] \
-t [nthreads] \
-o [/output/folder]
4. Score model
myevae score \
-e [endpoint: os | pfs] \
-n [model_name] \
-i [/inputs/folder] \
-o [/output/folder]
5. Generate SHAP plots
myevae shap \
-e [endpoint: os | pfs] \
-n [model_name] \
-i [/inputs/folder] \
-o [/output/folder]
Requirements
Files
Input csv files
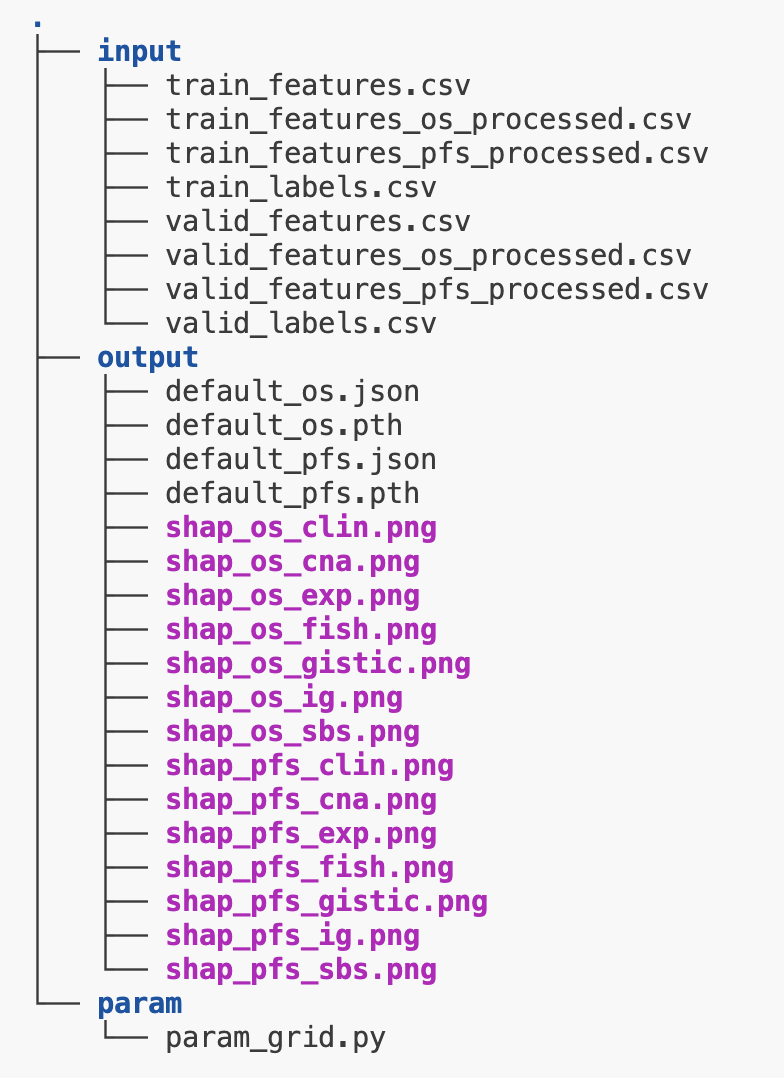
Place the following .csv files inside your inputs folder (/inputs/folder), and the param_grid.py inside your preferred folder. Also create an empty folder for the outputs.
|-----/inputs
| |----train_features.csv [ feature matrix of training samples ]
| |----valid_features.csv [ feature matrix of validation samples ]
| |----train_labels.csv [ survival files of training samples ]
| |____valid_labels.csv [ survival files of validation samples ]
|-----/params
| |____param_grid.py
|_____/output
|____[output files created here]
For features*.csv and labels*.csv, column 0 is read in as the index, which should be the patient IDs.
Example input csv files can be downloaded from AWS S3 link above.
Hyperparameter grid python file
Place a python file named param_grid.py containing the hyperparameter grid dictionary in the params folder (/params/folder). This contains the set of hyperparameters that grid search will be performed on. Otherwise, use the default provided in src/param_grid.py.
The example parameter grid provides 3 options for z_dim (dimension of latent space): 8, 16, or 32:
from torch.nn import LeakyReLU, Tanh
# the default hyperparameter grid for debugging uses
# this is not meant to be used for real training, as the search space is only on z_dim
param_grid = {
'z_dim': [8,16,32],
'lr': [5e-4],
'batch_size': [1024],
'input_types': [['exp','cna','gistic','fish','sbs','ig']],
'input_types_subtask': [['clin']],
'layer_dims': [[[32, 4],[16,4],[4,1],[4,1],[4,1],[4,1]]],
'layer_dims_subtask' : [[4,1]],
'kl_weight': [1],
'activation': [LeakyReLU()],
'subtask_activation': [Tanh()],
'epochs': [100],
'burn_in': [20],
'patience': [5],
'dropout': [0.3],
'dropout_subtask': [0.3]
}
Software
These dependencies will be automatically installed.
python >= 3.9
torch >= 1.9.0
scikit-learn >= 0.24.1
scikit-survival >= 0.23.1
importlib_resources
matplotlib
shap-0.47.3.dev8-offline-fork-for-myevae==0.0.1
The last dependency is a offline fork of shap (https://pypi.org/project/shap/) which has been modified to work MyeVAE.
Recommended hardware
-
Minimum 4 CPU cores (8 cores is recommended)
-
Minimum 16 GB RAM (64GB is required for feature preprocessing)
No GPU is required.
Total CPU time for feature preprocessing: ~24 hours
Total CPU time for training: ~2 minutes per model (no GPU required)
Citation
If you use MyeVAE in your research, please cite the authors:
[Manuscript under review. Contact author at changjiageng@u.nus.edu for citation.]
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file myevae-0.0.2.tar.gz.
File metadata
- Download URL: myevae-0.0.2.tar.gz
- Upload date:
- Size: 892.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f1871493835216238d4da61e237c97b7d40d6d08546139614622db45d7f3a549
|
|
| MD5 |
0484fbd5f6d0186e9581ede297e2f488
|
|
| BLAKE2b-256 |
10ac001fb4c59c9c8d05ab702d8e1b4be99fc1816fe3e513726991a75c29b30e
|
File details
Details for the file myevae-0.0.2-py3-none-any.whl.
File metadata
- Download URL: myevae-0.0.2-py3-none-any.whl
- Upload date:
- Size: 913.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
424c7e3c64dfed6e1acfe74e8e9a6f58ea3f09beed474e86f38bb46d8b4c8759
|
|
| MD5 |
9e10fee4cee61a7db0c7c131a82e558a
|
|
| BLAKE2b-256 |
2c79e69b27ee6f501f41bc9249e96040e538ea1ba5fc3fec6b94dc898cd2da2f
|