No project description provided
Project description
napari-cell-gater
A plugin to perform cell marker gating for multiplexed immunofluorescent imaging
Installation
Step 1. Install napari (see https://napari.org/stable/tutorials/fundamentals/installation)
Step 2.
Install napari-cell-gater via pip:
pip install git+https://github.com/melonora/napari-cell-gater.git
How to use
-
Users will select the necesary directories: images, masks, and quantification directories.
Assumptions for inputs:
1.1 Files inside these directories are named according to the samples names.
1.2 The image for sample 1, should be "1.ome.tif" or "1.tif"; the mask file "1.tif"; and the quantification file "1.csv".
1.3 Each set of files should all be inside each of the three folders.
1.4 Any extra files in those folders can make code fail. -
Select the lowerbound and upperbound channels to gate. These are all the columns from the quantification csv file that you want to threshold. You must pick the same channels if you plan to save and reload the gates.
-
Select a sample, and a marker from dropdown menus. 3 layers will load:
(a.) the reference channel (default: first channel, changeable by dropdown menu)
(b.) the segmentation mask (for large images this might be a problem)
(c.) the channel_to_be_gated
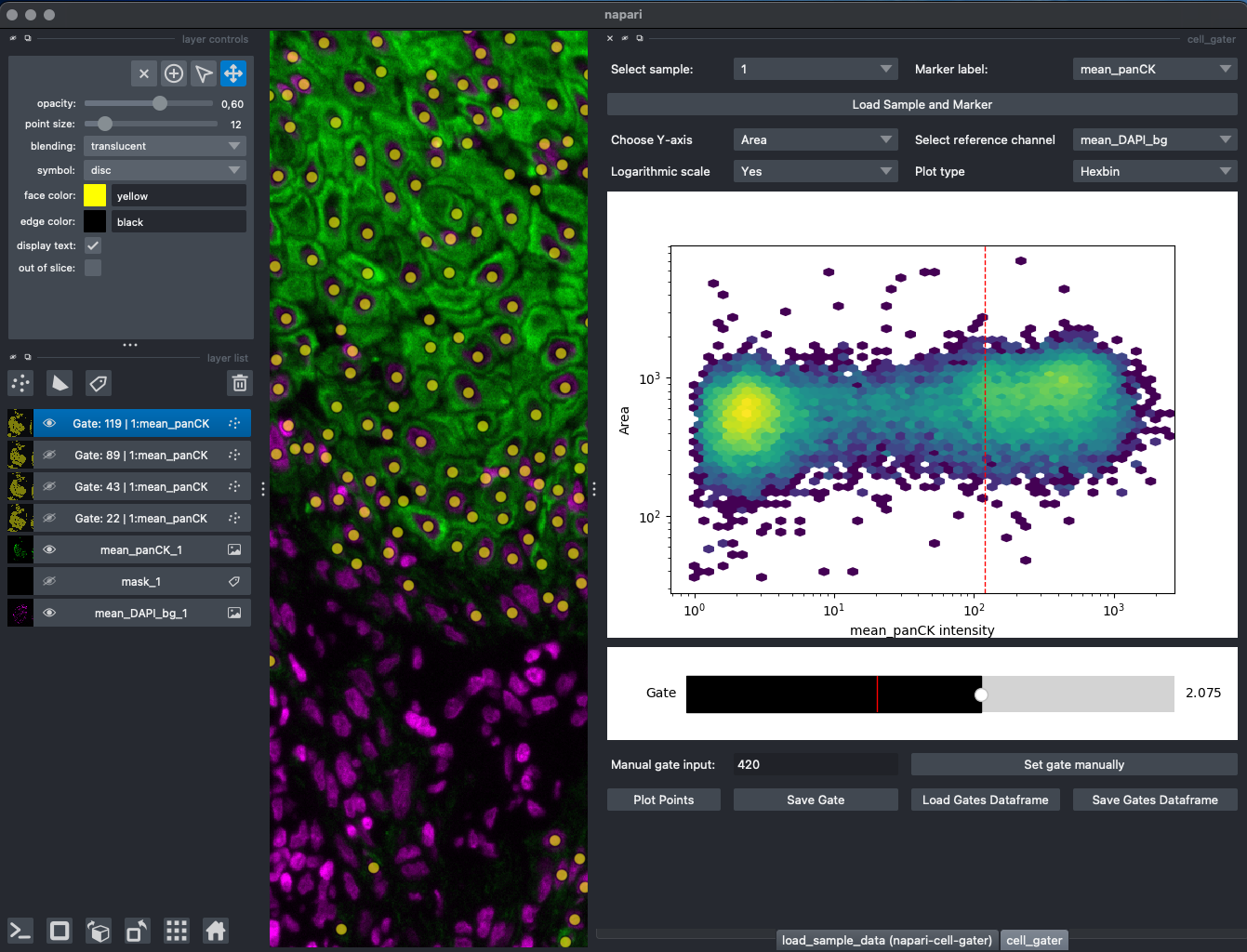
A scatter plot (default: x-axis=channel_to_be_gated intensity, y-axis=Area) (y-axis can be changed by dropdown)
Underneath the scatterplot a slider will appear, the position of the slider will show up as a vertical line in the scatter plot. The scatter plot can also be changed to a hexbin plot, which really helps with dense clusters of cells. Plotting the data in log10 space is also possible by dropdown. Most of the times it helps. Gates would still be saved in linear space. -
Adjust the contrast with the Napari layer menu (top left)
-
Drag the slider to what they think is correct
-
Click "Plot Points" to plot points on top of positive cells.
-
Repeat steps 5 and 6 until satisfied.
-
Click "Save Gate" to save the gate for the current marker and sample. Go to step 4 and repeat.
Contributing
Contributions are very welcome. Tests can be run with tox, please ensure the coverage at least stays the same before you submit a pull request.
License
Distributed under the terms of the BSD-3 license, "napari-cell-gater" is free and open source software
Issues
If you encounter any problems, please [file an issue] along with a detailed description.
Assumptions of usage
(1) Images should have the same number of channels as columns in the
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file napari_cell_gater-0.5.0.tar.gz.
File metadata
- Download URL: napari_cell_gater-0.5.0.tar.gz
- Upload date:
- Size: 24.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
19701a6ca73954c0c2c168ef31dff688e08a852319903be5f923b6c04b8f84fe
|
|
| MD5 |
5fbb60183871d0c1dc9795bc760d7769
|
|
| BLAKE2b-256 |
0fa67566ba0cad91d033eeb642e4e81862b06c128dc432d5c86db511a2f0d88f
|
File details
Details for the file napari_cell_gater-0.5.0-py3-none-any.whl.
File metadata
- Download URL: napari_cell_gater-0.5.0-py3-none-any.whl
- Upload date:
- Size: 19.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
284a694872fbd391c62cad1742c621f3dba2c9558e1a84760128654597b11854
|
|
| MD5 |
776cb25b4edf70436ccc1164074051c7
|
|
| BLAKE2b-256 |
b34245b15d8064d0898eb2b9ad24c17ebb97dac1967ed49e5056c1577086e805
|