A small package with Numba-accelerated morphological operations.
Project description
nbmorph
A small, Numba-accelerated Python package for morphological operations on 3D labeled images.
nbmorph provides a set of common morphological operations optimized for performance using Numba. It is designed to work with 3D NumPy arrays representing labeled image data, where different integer labels correspond to different objects.
Features
-
Numba-accelerated and multithreaded: Operations are just-in-time compiled with Numba for high performance on CPUs.
-
3D Label Image Support: All operations are designed for 3D labeled images (integer NumPy arrays).
-
Quasi-Spherical Structuring Elements: Approximates spherical structuring elements by alternating between box and diamond kernels for dilation and erosion.
-
Core Morphological Operations:
-
dilate_labels_spherical: Expands the boundaries of labeled regions by assigning the mode of the neighborhood to background voxels. -
erode_labels_spherical: Shrinks the boundaries of labeled regions. -
open_labels_spherical: Removes small noise and thin protrusions (erosion followed by dilation). -
close_labels_spherical: Fills small holes within objects (dilation followed by erosion). -
smooth_labels_spherical: Smoothes object boundaries by performing an opening followed by a closing.
-
-
Mode (majority) filters: Stencil-based mode filters built on top of compile-time sorting networks for fast, fixed-size neighborhoods. Each comes in two variants:
-
onlyzero_mode_box/onlyzero_mode_diamond: Fill only background (zero) voxels with the mode of their 3x3x3 box or 6-connected diamond neighborhood, leaving labeled voxels unchanged. -
mode_box/mode_diamond: Replace every voxel with the mode of its neighborhood (including the center), useful for denoising labeled volumes.
-
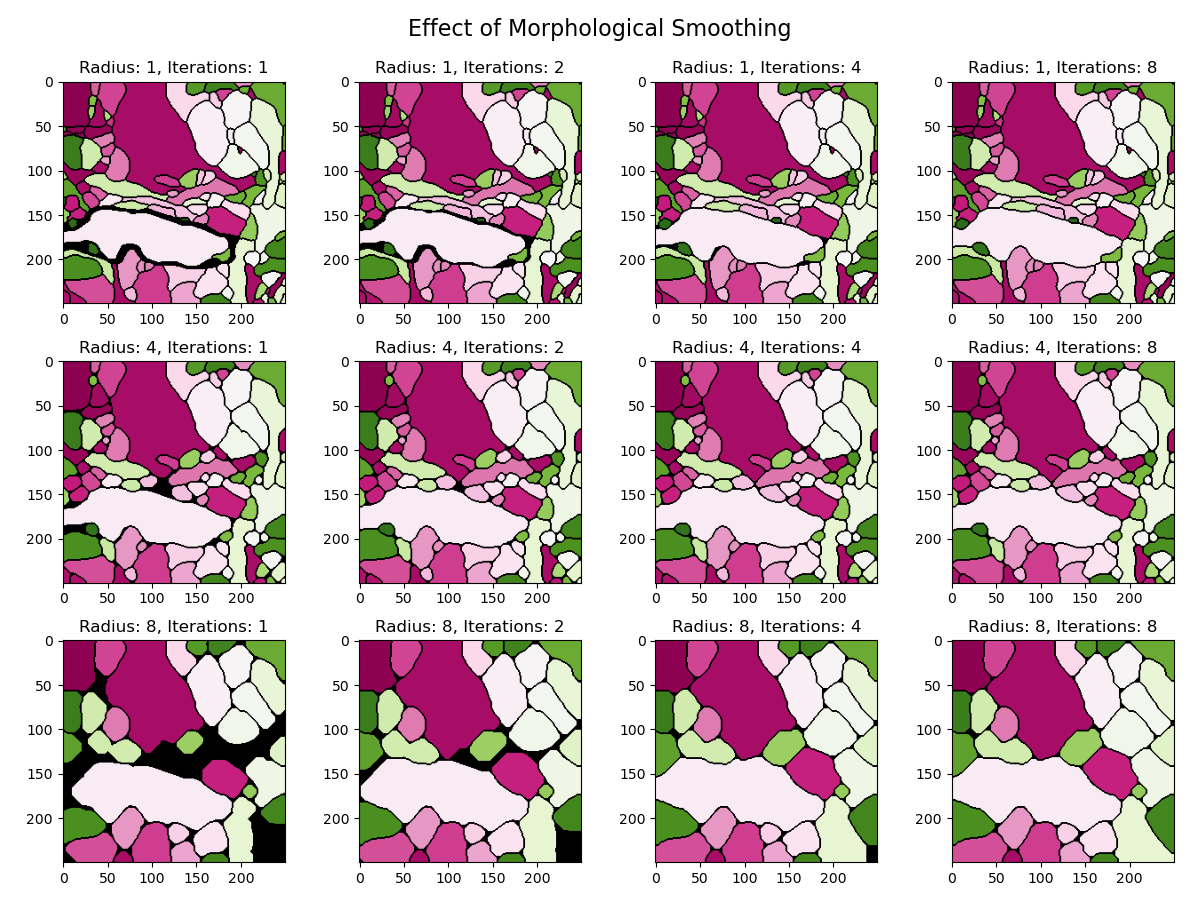
Installation
You can install nbmorph directly from pypi using pip:
pip install nbmorph
Usage
Here is a basic example of how to use nbmorph to apply operations to a 3D labeled image.
import numpy as np
import nbmorph
import numba
numba.set_num_threads(4)
# Create a sample 3D labeled image
# For example, a 5x5x5 cube of two different labels in a 10x10x10 volume
labels = np.zeros((10, 10, 10), dtype=np.uint16)
labels[2:7, 2:7, 2:5] = 1
labels[2:7, 2:7, 5:7] = 2
# First execution may take a while due to numba compilation
# Apply morphological erosion with a radius of 1
eroded_labels = nbmorph.erode_labels_spherical(labels, radius=1)
# Apply morphological dilation with a radius of 1
dilated_labels = nbmorph.dilate_labels_spherical(labels, radius=1)
# Apply morphological opening with a radius of 1
opened_labels = nbmorph.open_labels_spherical(labels, radius=1)
# Apply morphological closing with a radius of 1
closed_labels = nbmorph.close_labels_spherical(labels, radius=1)
# Apply morphological smoothing with a radius of 1
smoothed_labels = nbmorph.smooth_labels_spherical(labels, radius=1)
Testing
Tests are written using pytest. To run the tests, first install the test dependencies and then run pytest:
pip install .[test]
pytest
Benchmarking
A benchmark script is included in the scripts directory:
python scripts/benchmark.py
License
This project is licensed under the MIT License - see the LICENSE file for details.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file nbmorph-0.3.0.tar.gz.
File metadata
- Download URL: nbmorph-0.3.0.tar.gz
- Upload date:
- Size: 18.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
419d2e2888296d3bdb56c2b92f4d2fc6a92696c2c0220e5baa52b878f725344a
|
|
| MD5 |
b485f93a73816a3c3c1d6704650bc646
|
|
| BLAKE2b-256 |
16ce0dab43c1486c9af01cf1a86bb960ac0e951588f45b9ffce36f980c1f1a39
|
Provenance
The following attestation bundles were made for nbmorph-0.3.0.tar.gz:
Publisher:
release.yaml on MariusCausemann/nbmorph
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
nbmorph-0.3.0.tar.gz -
Subject digest:
419d2e2888296d3bdb56c2b92f4d2fc6a92696c2c0220e5baa52b878f725344a - Sigstore transparency entry: 1392739686
- Sigstore integration time:
-
Permalink:
MariusCausemann/nbmorph@568772f43ad86551ec12c2af856b7cb6079a8b07 -
Branch / Tag:
refs/tags/v0.3 - Owner: https://github.com/MariusCausemann
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yaml@568772f43ad86551ec12c2af856b7cb6079a8b07 -
Trigger Event:
push
-
Statement type:
File details
Details for the file nbmorph-0.3.0-py3-none-any.whl.
File metadata
- Download URL: nbmorph-0.3.0-py3-none-any.whl
- Upload date:
- Size: 16.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1bc863b05cedd75190403d32771126d31061d54f47736f153b51410acfa24ac0
|
|
| MD5 |
f776a6c9e4aec74af4b0da563702eaf1
|
|
| BLAKE2b-256 |
3cf9eec368905584e458da6c026d2c21590ad526b336ebd073a3f8d7f8950725
|
Provenance
The following attestation bundles were made for nbmorph-0.3.0-py3-none-any.whl:
Publisher:
release.yaml on MariusCausemann/nbmorph
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
nbmorph-0.3.0-py3-none-any.whl -
Subject digest:
1bc863b05cedd75190403d32771126d31061d54f47736f153b51410acfa24ac0 - Sigstore transparency entry: 1392739695
- Sigstore integration time:
-
Permalink:
MariusCausemann/nbmorph@568772f43ad86551ec12c2af856b7cb6079a8b07 -
Branch / Tag:
refs/tags/v0.3 - Owner: https://github.com/MariusCausemann
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yaml@568772f43ad86551ec12c2af856b7cb6079a8b07 -
Trigger Event:
push
-
Statement type:













