Generate metro-map-style SVG diagrams from Mermaid graph definitions
Project description
nf-metro
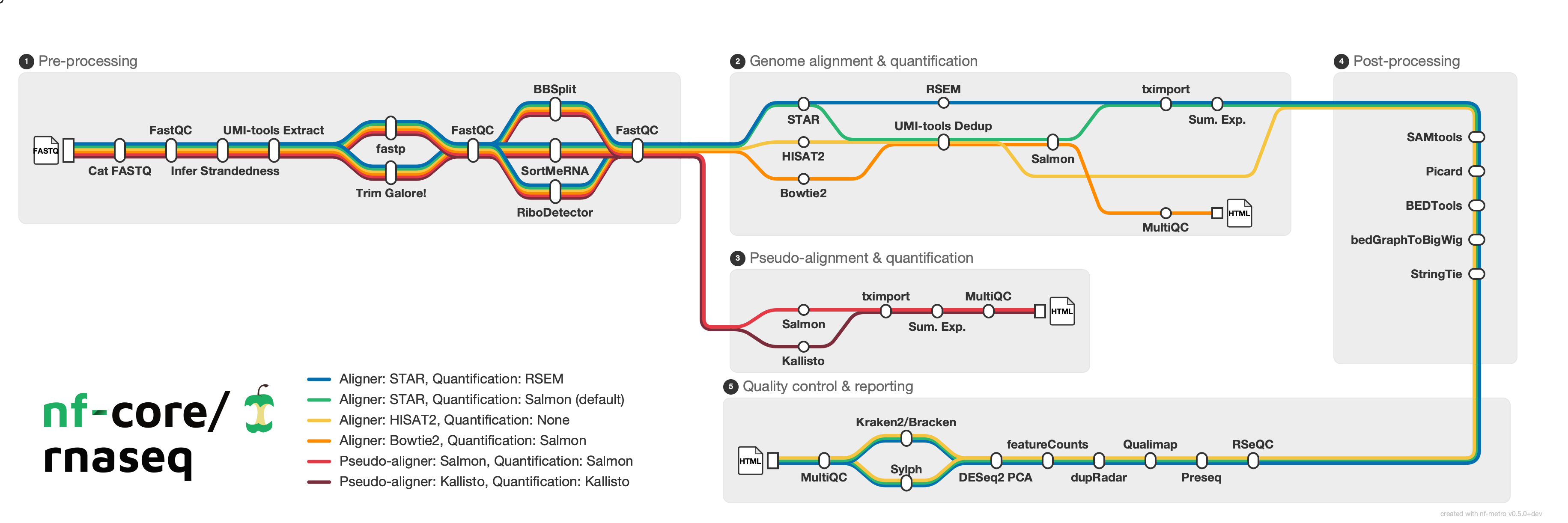
Generate metro-map-style SVG diagrams from Mermaid graph definitions with %%metro directives. Designed for visualizing bioinformatics pipeline workflows (e.g., nf-core pipelines) as transit-style maps where each analysis route is a colored "metro line."
Installation
pip install nf-metro
For development:
pip install -e ".[dev]"
Requires Python 3.10+.
Quick start
Render a metro map from a .mmd file:
nf-metro render examples/simple_pipeline.mmd -o pipeline.svg
Validate your input without rendering:
nf-metro validate examples/simple_pipeline.mmd
Inspect structure (sections, lines, stations):
nf-metro info examples/simple_pipeline.mmd
CLI reference
nf-metro render
Render a Mermaid metro map definition to SVG.
nf-metro render [OPTIONS] INPUT_FILE
| Option | Default | Description |
|---|---|---|
-o, --output PATH |
<input>.svg |
Output SVG file path |
--theme [nfcore|light] |
nfcore |
Visual theme |
--width INTEGER |
auto | SVG width in pixels |
--height INTEGER |
auto | SVG height in pixels |
--x-spacing FLOAT |
60 |
Horizontal spacing between layers |
--y-spacing FLOAT |
40 |
Vertical spacing between tracks |
--max-layers-per-row INTEGER |
auto | Max layers before folding to next row |
--animate / --no-animate |
off | Add animated balls traveling along lines |
--debug / --no-debug |
off | Show debug overlay (ports, hidden stations, edge waypoints) |
--logo PATH |
none | Logo image path (overrides %%metro logo: directive) |
The --logo flag lets you use the same .mmd file with different logos for dark/light themes:
nf-metro render pipeline.mmd -o pipeline_dark.svg --theme nfcore --logo logo_dark.png
nf-metro render pipeline.mmd -o pipeline_light.svg --theme light --logo logo_light.png
nf-metro validate
Check a .mmd file for errors without producing output.
nf-metro validate INPUT_FILE
nf-metro info
Print a summary of the parsed map: sections, lines, stations, and edges.
nf-metro info INPUT_FILE
Examples
The examples/ directory contains ready-to-render .mmd files:
| Example | Description |
|---|---|
simple_pipeline.mmd |
Minimal two-line pipeline with no sections |
rnaseq_auto.mmd |
nf-core/rnaseq with fully auto-inferred layout |
rnaseq_sections.mmd |
nf-core/rnaseq with manual grid overrides |
Topology gallery
The examples/topologies/ directory has 15 examples covering a range of layout patterns. See the topology README for descriptions and rendered previews.
A few highlights:
| Wide Fan-Out | Section Diamond | Variant Calling |
 |
 |
 |
| Fold Serpentine | Multi-Line Bundle | RNA-seq Lite |
 |
 |
 |
Input format
Input files use a subset of Mermaid graph LR syntax extended with %%metro directives. The format has three layers: global directives that configure the overall map, section directives inside subgraph blocks that control section layout, and edges that define connections between stations.
Walkthrough: nf-core/rnaseq
The full example is at examples/rnaseq_sections.mmd. Here's how each part works.
Global directives
%%metro title: nf-core/rnaseq
%%metro logo: examples/nf-core-rnaseq_logo_dark.png
%%metro style: dark
title:sets the map title (shown top-left unless a logo is provided)logo:embeds a PNG image in place of the text titlestyle:selects a theme (darkorlight)
Lines (routes)
Each metro line represents a distinct path through the pipeline. Lines are defined with an ID, display name, and color:
%%metro line: star_rsem | Aligner: STAR, Quantification: RSEM | #0570b0
%%metro line: star_salmon | Aligner: STAR, Quantification: Salmon (default) | #2db572
%%metro line: hisat2 | Aligner: HISAT2, Quantification: None | #f5c542
%%metro line: pseudo_salmon | Pseudo-aligner: Salmon, Quantification: Salmon | #e63946
%%metro line: pseudo_kallisto | Pseudo-aligner: Kallisto, Quantification: Kallisto | #7b2d3b
In the rnaseq pipeline, each line corresponds to a parameter-driven analysis route. All five lines share the preprocessing section, then diverge based on aligner choice.
Grid placement
Sections are placed on a grid automatically via topological sort, but explicit positions can be set:
%%metro grid: postprocessing | 2,0,2
%%metro grid: qc_report | 1,2,1,2
The format is section_id | col,row[,rowspan[,colspan]]. In this example:
postprocessingis pinned to column 2, row 0, spanning 2 rows verticallyqc_reportis pinned to column 1, row 2, spanning 2 columns horizontally
Legend
%%metro legend: bl
Position the legend: tl, tr, bl, br (corners), bottom, right, or none.
Sections
Sections are Mermaid subgraph blocks. Each section is laid out independently, then placed on the grid:
graph LR
subgraph preprocessing [Pre-processing]
%%metro exit: right | star_salmon, star_rsem, hisat2
%%metro exit: bottom | pseudo_salmon, pseudo_kallisto
cat_fastq[cat fastq]
fastqc_raw[FastQC]
...
end
Section directives:
%%metro entry: <side> | <line_ids>- declares which lines enter this section and from which side (left,right,top,bottom)%%metro exit: <side> | <line_ids>- declares which lines exit and to which side%%metro direction: <dir>- section flow direction:LR(default),RL(right-to-left), orTB(top-to-bottom)
Entry/exit hints control port placement on section boundaries. A section can have exit hints on multiple sides (e.g., preprocessing exits right for aligners and bottom for pseudo-aligners), but all lines from a section leave through a single exit port. If all exit hints point to one side, that side is used; otherwise it defaults to right.
Section directions
Most sections flow left-to-right (LR, the default). Two other directions are useful for layout:
Top-to-bottom (TB) - used for the Post-processing section, which acts as a vertical connector carrying lines downward:
subgraph postprocessing [Post-processing]
%%metro direction: TB
%%metro entry: left | star_salmon, star_rsem, hisat2
%%metro exit: bottom | star_salmon, star_rsem, hisat2
samtools[SAMtools]
picard[Picard]
...
end
Right-to-left (RL) - used for the QC section, which flows backward to create a serpentine layout:
subgraph qc_report [Quality control & reporting]
%%metro direction: RL
%%metro entry: top | star_salmon, star_rsem, hisat2
rseqc[RSeQC]
preseq[Preseq]
...
end
Stations and edges
Stations use Mermaid node syntax. Edges carry comma-separated line IDs to indicate which routes use that connection:
cat_fastq[cat fastq]
fastqc_raw[FastQC]
cat_fastq -->|star_salmon,star_rsem,hisat2,pseudo_salmon,pseudo_kallisto| fastqc_raw
All five lines pass through this edge. Later, lines diverge:
star -->|star_rsem| rsem
star -->|star_salmon| umi_tools_dedup
hisat2_align -->|hisat2| umi_tools_dedup
Here different lines take different paths through the section, creating the visual fork in the metro map.
Inter-section edges
Edges between stations in different sections go outside all subgraph/end blocks:
%% Inter-section edges
sortmerna -->|star_salmon,star_rsem| star
sortmerna -->|hisat2| hisat2_align
sortmerna -->|pseudo_salmon| salmon_pseudo
sortmerna -->|pseudo_kallisto| kallisto
stringtie -->|star_salmon,star_rsem,hisat2| rseqc
These are automatically rewritten into port-to-port connections with junction stations at fan-out points. You just specify the source and target stations directly.
Directive reference
| Directive | Scope | Description |
|---|---|---|
%%metro title: <text> |
Global | Map title |
%%metro logo: <path> |
Global | Logo image (replaces title text) |
%%metro style: <name> |
Global | Theme: dark, light |
%%metro line: <id> | <name> | <color> |
Global | Define a metro line |
%%metro grid: <section> | <col>,<row>[,<rowspan>[,<colspan>]] |
Global | Pin section to grid position |
%%metro legend: <position> |
Global | Legend position: tl, tr, bl, br, bottom, right, none |
%%metro file: <station> | <label> |
Global | Mark a station as a file terminus with a document icon |
%%metro entry: <side> | <lines> |
Section | Entry port hint |
%%metro exit: <side> | <lines> |
Section | Exit port hint |
%%metro direction: <dir> |
Section | Flow direction: LR, RL, TB |
License
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file nf_metro-0.2.0.tar.gz.
File metadata
- Download URL: nf_metro-0.2.0.tar.gz
- Upload date:
- Size: 1.7 MB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
6ed31c3b03d709e8b7ac6435c96c105f1f2ef4b0ce8de2630bc7c93fe45cd052
|
|
| MD5 |
a585771b163646cb2eb34926a445b0df
|
|
| BLAKE2b-256 |
ad9dd1727610f108b2b62690d040384afe2b2cca919471ea4284408cf9215a4e
|
Provenance
The following attestation bundles were made for nf_metro-0.2.0.tar.gz:
Publisher:
publish.yml on pinin4fjords/nf-metro
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
nf_metro-0.2.0.tar.gz -
Subject digest:
6ed31c3b03d709e8b7ac6435c96c105f1f2ef4b0ce8de2630bc7c93fe45cd052 - Sigstore transparency entry: 959548731
- Sigstore integration time:
-
Permalink:
pinin4fjords/nf-metro@ab50c6c20253c779dd738a84096b5be0bd4d56f3 -
Branch / Tag:
refs/tags/0.2.0 - Owner: https://github.com/pinin4fjords
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@ab50c6c20253c779dd738a84096b5be0bd4d56f3 -
Trigger Event:
release
-
Statement type:
File details
Details for the file nf_metro-0.2.0-py3-none-any.whl.
File metadata
- Download URL: nf_metro-0.2.0-py3-none-any.whl
- Upload date:
- Size: 73.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
55d8ce3ebb82783fb8254f983138e201d78f6a617f310fc8e07237e23a60c2db
|
|
| MD5 |
e31bd253b802fdb0e7812c0b6835a21d
|
|
| BLAKE2b-256 |
0bd65a5c8c9e5bbabc4d769b7bb18b86e2caf1a5cd3dd90d334c640844c2be0d
|
Provenance
The following attestation bundles were made for nf_metro-0.2.0-py3-none-any.whl:
Publisher:
publish.yml on pinin4fjords/nf-metro
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
nf_metro-0.2.0-py3-none-any.whl -
Subject digest:
55d8ce3ebb82783fb8254f983138e201d78f6a617f310fc8e07237e23a60c2db - Sigstore transparency entry: 959548797
- Sigstore integration time:
-
Permalink:
pinin4fjords/nf-metro@ab50c6c20253c779dd738a84096b5be0bd4d56f3 -
Branch / Tag:
refs/tags/0.2.0 - Owner: https://github.com/pinin4fjords
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@ab50c6c20253c779dd738a84096b5be0bd4d56f3 -
Trigger Event:
release
-
Statement type:











