Graph-based foundation model for spatial transcriptomics data
Project description

💫 Graph-based foundation model for spatial transcriptomics data
Novae is a deep learning model for spatial domain assignments of spatial transcriptomics data (at both single-cell or spot resolution). It works across multiple gene panels, tissues, and technologies. Novae offers several additional features, including: (i) native batch-effect correction, (ii) analysis of spatially variable genes and pathways, and (iii) architecture analysis of tissue slides.
[!NOTE] Novae was developed by the authors of
sopaand is part of thescverseecosystem. Read our article here.
Documentation
Check Novae's documentation to get started. It contains installation explanations, API details, and tutorials.
Overview
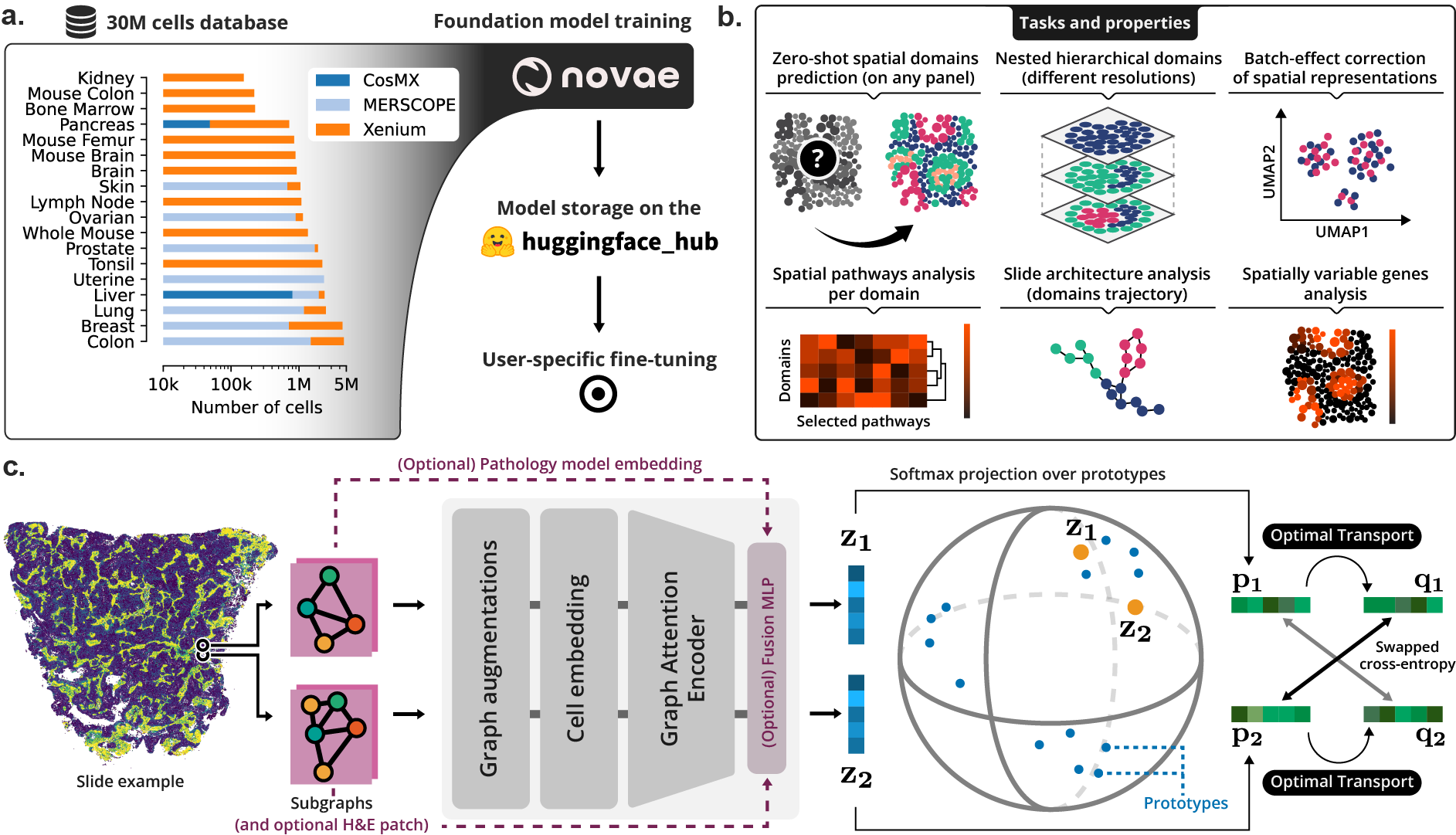
(a) Novae was trained on a large dataset, and is shared on Hugging Face Hub. (b) Illustration of the main tasks and properties of Novae. (c) Illustration of the method behind Novae (self-supervision on graphs, adapted from SwAV).
Installation
novae can be installed from PyPI on all OS, for any Python version >=3.11 and <3.14 (3.14 support coming soon).
pip install novae
[!NOTE] See this installation section for more details about extras and other installations modes.
Usage
Here is a minimal usage example. For more details, refer to the documentation.
import novae
# compute cell neighbors
novae.spatial_neighbors(adata)
# load a pre-trained model
model = novae.Novae.from_pretrained("MICS-Lab/novae-human-0")
# compute spatial domains
model.compute_representations(adata, zero_shot=True)
model.assign_domains(adata)
Cite us
Our article is published in Nature Methods. You can cite Novae as below:
Blampey, Q., Benkirane, H., Bercovici, N. et al. Novae: a graph-based foundation model for spatial transcriptomics data.
Nat Methods (2025). https://doi.org/10.1038/s41592-025-02899-6
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file novae-1.0.2.tar.gz.
File metadata
- Download URL: novae-1.0.2.tar.gz
- Upload date:
- Size: 52.4 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.5.20
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1c54a3ba3c0d7d7875d4464882569d662a303e9275077f676142ff8d9bb26a02
|
|
| MD5 |
b5378e842b5c7cc19bc9b66cb3f0ca5c
|
|
| BLAKE2b-256 |
4accd1f014a9b031b421a80be8f243cc3128dbd1322c183a385d324e3a2a4b84
|
File details
Details for the file novae-1.0.2-py3-none-any.whl.
File metadata
- Download URL: novae-1.0.2-py3-none-any.whl
- Upload date:
- Size: 67.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.5.20
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9f8672627533b26eab70101b27ae7eecef398ed8a4102dec6fe9eb22b34c6e9e
|
|
| MD5 |
7358639b8d4519b4e1694dec96f73de9
|
|
| BLAKE2b-256 |
6989d7cfda9a9873e56f0525136bc70548c5c3ed0b892e07c6d00de7f9baa9f6
|



















