Interface for NumPy structured arrays
Project description
np-struct
np-struct is a user friendly interface to NumPy structured arrays, with added support for transferring arrays across interfaces.
The Struct type is designed to mirror the struct typedef in C, but can be used for any complicated data structure. They behave similar to the standard ndarray, but support mixed data types, bitfields, labeling, and variable length arrays. Arrays are easily written or loaded from disk in the standard .npy binary format.
Installation
pip install np-struct
Usage
import np_struct
from np_struct import Struct
import numpy as np
Structures
Create a C-style structure with Numpy types.
class example(Struct):
data1 = np.uint32()
data2 = np.complex128([0]*3)
Structures can be initialized with arbitrary shapes by using the shape kwarg:
ex = example(shape = (3,), byte_order='>')
ex[0].data2 = 1 + 2j
>>> ex
Struct example: (3,)
[
data1: uint32[0]
data2: complex128[1.+2.j 1.+2.j 1.+2.j]
]
...
[
data1: uint32[0]
data2: complex128[0.+0.j 0.+0.j 0.+0.j]
]
Members can also be initialized by passing in their name to the constructor with an initial value.
>>> example(data2 = np.zeros(shape=(3,2)))
Struct example:
data1: uint32[0]
data2: complex128[[0.+0.j 0.+0.j]
[0.+0.j 0.+0.j]
[0.+0.j 0.+0.j]]
The structure inherits from np.ndarray and supports all math functions that a normal structured array does. To cast as a standard numpy structured array,
>>> ex.view(np.ndarray)
array([([0], [0.+0.j, 0.+0.j, 0.+0.j]), ([0], [0.+0.j, 0.+0.j, 0.+0.j]),
([0], [0.+0.j, 0.+0.j, 0.+0.j])],
dtype=[('data1', '<u4', (1,)), ('data2', '<c16', (3,))])
Nested structures are also supported,
class nested(Struct):
field1 = example()
field2 = example()
n = nested()
n.field1.data2 += 1j*np.pi
>>> n
Struct nested:
field1: Struct example:
data1: uint32[0]
data2: complex128[0.+3.14159265j 0.+3.14159265j 0.+3.14159265j]
field2: Struct example:
data1: uint32[0]
data2: complex128[0.+0.j 0.+0.j 0.+0.j]
To save to disk,
n = nested(shape=2)
n[1].field2.data1 = 3
np.save("test.npy", n)
n_disk = nested(np.load("test.npy"))
>>> n_disk.field2.data1
array([[[0]],
[[3]]], dtype=uint32)
Labeled Arrays
ldarray supports indexing with coordinates, interpolation, and can be written to disk in the standard .npy
binary format,
Math operations that change the coordinates or array shape (i.e. sum or transpose) silently revert the labeled array
to a standard numpy array without coordinates. np-struct leaves it up to the user to re-cast the array as an
ldarray with the appropriate coordinates.
>>> from np_struct import ldarray
>>> import numpy as np
>>> coords = dict(a=['data1', 'data2'], b=np.arange(0, 20, 0.2))
>>> data = np.arange(200).reshape(2, 100)
>>> ld = ldarray(data, coords=coords)
>>> ld
ldarray([[ 0, 1, ... 98, 99],
[100, 101, ... 198, 199]])
Coordinates: (2, 100)
a: ['data1' 'data2']
b: [ 0. 0.2 ... 19.6 19.8]
Coordinate indexing can be done with the .sel method or by using a dictionary in the indexing brackets [...].
Indexing with slices is inclusive on the endpoint:
>>> ld.sel(b=slice(15, 16), a="data1")
ldarray([75, 76, 77, 78, 79, 80])
Coordinates: (6,)
b: [15. 15.2 ... 15.8 16. ]
Values can be indexed with lists or arrays:
>>> ld.sel(b = [0.2, 19.6])
ldarray([[ 1, 98],
[101, 198]])
Coordinates: (2, 2)
a: ['data1' 'data2']
b: [ 0.2 19.6]
To use a dictionary to index,
>>> ld[dict(b = 19.8)] = 77
>>> ld
ldarray([[ 0, 1, 2, ... 98, 77],
[100, 101, 102, ... 198, 77]])
Coordinates: (2, 100)
a: ['data1' 'data2']
b: [ 0. 0.2 0.4 ... 19.6 19.8]
Arrays can be interpolated with the .interpolate() method. This works the same as the .sel() method
but allows coordinates to be outside the indexing tolerance.
>>> ang = np.linspace(0, 2 * np.pi, 21)
>>> ang_int = np.linspace(0, 2 * np.pi, 61)
>>> ld = ldarray(
... np.array([np.exp(1j * t), np.exp(0.5 * 1j * t)]),
... coords = dict(a=["exp(t)", "exp(0.5t)"], ang=ang)
)
# the interpolation by default is order=3, but reduces to linear near the endpoints.
# the dtype is not required if the output is the same dtype as the input, which is true in this case
# but is shown for completeness.
>>> ld_int = ld.interpolate(ang=ang_int, a="exp(t)", dtype=np.complex128).squeeze()
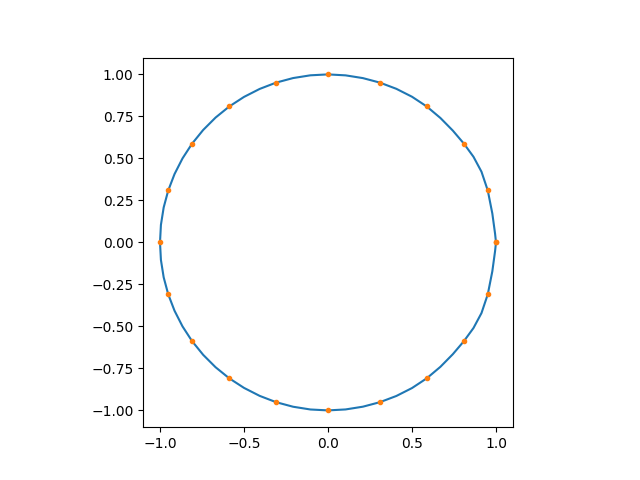
# plot interpolation results
import matplotlib.pyplot as plt
plt.plot(ld_int.real, ld_int.imag)
plt.plot(ld[0].real, ld[0].imag, marker=".", linestyle="")
plt.gca().set_aspect("equal")
Real or complex-valued arrays can be written to disk using the normal numpy methods if the coordinates are not needed.
To keep the coords, use ldarray.save() and load(). Array is stored as a structured array in the usual
.npy binary format.
ld.save("ld_file.npy")
ldarray.load("ld_file.npy")
Examples
Struct example
Labeled Array example
License
np-struct is licensed under the MIT License.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file np_struct-0.0.6.tar.gz.
File metadata
- Download URL: np_struct-0.0.6.tar.gz
- Upload date:
- Size: 23.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d3135054cbf5f2b2151923c322cc98a7fa1d2e2109b32bc4a244791dd3f80ff1
|
|
| MD5 |
db9b8bdc6502df86b088753f4f4b80d7
|
|
| BLAKE2b-256 |
2765602f660fe3bbb70796a6355baff75cad996af7f582f1712005f1c8886fa1
|
File details
Details for the file np_struct-0.0.6-py3-none-any.whl.
File metadata
- Download URL: np_struct-0.0.6-py3-none-any.whl
- Upload date:
- Size: 19.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
739efcf0a8d72eb06f41fad36b318f876b1dfeebd713441ce0a2cc003cae978d
|
|
| MD5 |
1ee18ffd6e4410f5dd2a81a696eb14d0
|
|
| BLAKE2b-256 |
f42c491e2739aa0280b432c475ad32829c873d199c17f4ddbbaf945bee2bf5d3
|