Python based open source text-only project preparation framework for Nanoscribe's two-photon 3D lithography system Quantum X align.
Project description

npxpy
npxpy is a versatile open source Python package that enables you to build projects (NANO files) for the 3D direct laser lithography system Nanoscribe Quantum X align (QXa) via CLI/Scripts. It is designed such that it adheres to the same workflow logic as Nanoscribe's GUI software nanoPrintX, making the application additionally user-friendly to experienced users of the QXa.
Table of Contents
Installation
You can install npxpy via pip together with all features (recommended) :
pip install npxpy[all]
It is recommended to install npxpy in a virtual environment to prevent dependency issues.
A more selective installation with respect to features like the 3D-viewport or GDS-parsing is possible as well by
exchanging [all] with [viewport] and [gds], respectively. If you are interested in a light-weight
installation you are able to install only the core features of npxpy via:
pip install npxpy
Beware that the light-weight installation lacks the other aforementioned features entirely.
Features and Usage
Straightforward print project preparation with usual workflow logic is embeded in the Python-software ecosystem with extras like GDS-parsing for high workflow integration.
>>> import npxpy
>>> # Initialize the presets and resources that you want to use in this project.
>>> # You can either load presets directly from a .toml...
>>> preset_from_file = npxpy.Preset.load_single(file_path="preset_from_file.toml")
>>>
>>> # ... or initialize it inside of your script.
>>> edit_presets = {
... "writing_speed": 220000.0,
... "writing_power": 50.0,
... "slicing_spacing": 0.8,
... "hatching_spacing": 0.3,
... "hatching_angle": 0.0,
... "hatching_angle_increment": 0.0,
... "hatching_offset": 0.0,
... "hatching_offset_increment": 0.0,
... "hatching_back_n_forth": True,
... "mesh_z_offset": 0.0,
... }
>>>
>>> preset_from_args = npxpy.Preset(name="preset_from_args", **edit_presets)
>>>
>>> # Load your resources simply via path to their directories.
>>> stl_mesh = npxpy.Mesh(file_path="./example_mesh.stl", name="stl_structure")
>>> marker = npxpy.Image(file_path="./example_marker.png", name="marker_image")
>>>
>>> # Initialize your project and load your presets and resources into it.
>>> project = npxpy.Project(objective="25x", resin="IP-n162", substrate="FuSi")
>>> project.load_presets(preset_from_file, preset_from_args)
>>> project.load_resources(stl_mesh, marker)
>>>
>>> # Prepare the nodes of your project as usual.
>>> # Setup alignment nodes
>>> coarse_aligner = npxpy.CoarseAligner(residual_threshold=8)
>>> marker_aligner = npxpy.MarkerAligner(
... name="Marker Aligner", image=marker, marker_size=[10, 10]
... )
>>>
>>> # Set anchors manually...
>>> ca_positions = [
... [200.0, 200.0, 0.0],
... [200.0, -200.0, 0.0],
... [-200.0, -200.0, 0.0],
... [-200.0, 200.0, 0.0],
... ]
>>> ma_positions = [
... [0, 200, 0.33],
... [200, 0, 0.33],
... [0, -200, 0.33],
... [-200, 0, 0.33],
... ]
>>>
>>> coarse_aligner.set_coarse_anchors_at(ca_positions)
>>> marker_aligner.set_markers_at(ma_positions)
>>>
>>> # ... or incorporate them in a GDS-design and read them in.
>>> import npxpy.gds
>>>
>>> gds = npxpy.gds.GDSParser("gds_file.gds")
>>>
>>> interface_aligner = gds.get_custom_interface_aligner(
... cell_name="cell_with_print_scene",
... interface_layer=(1, 0),
... signal_type="reflection",
... detector_type="confocal",
... area_measurement=True,
... measure_tilt=True,
... )
>>>
>>> # Initialize printing scene
>>> scene = npxpy.Scene(name="scene", writing_direction_upward=True)
>>>
>>> # Initialize structure with desired preset and mesh defined above.
>>> structure = npxpy.Structure(
... name="structure", preset=preset_from_file, mesh=stl_mesh
... )
>>>
>>> # Arrange hierarchy of all nodes as desired either with .add_child()...
>>> coarse_aligner.add_child(scene.add_child(interface_aligner))
>>>
>>> # ...or more conveniently by using .append_node() to append
>>> # consecutively to the lowest node.
>>> scene.append_node(marker_aligner, structure)
>>>
>>> # Eventually, add all highest-order nodes of interest
>>> # (here only coarse_aligner) to project.
>>> project.add_child(coarse_aligner)
>>>
>>> # After allocating your nodes, you can copy, manipulate and add additional
>>> # instances as you like.
>>> scene_1 = scene.deepcopy_node(copy_children=True)
>>> scene_1.name = "scene_1"
>>> scene_1.translate([254.0, 300.0, 0.0])
>>>
>>> # You can access descendants/ancestors as you go via semantically ordered lists.
>>> structure_1 = scene_1.all_descendants[-1]
>>> structure_1.preset = preset_from_args
>>> structure_1.name = "structure_1"
>>>
>>> coarse_aligner.add_child(scene_1)
>>>
>>> # Checking the node order can be done as well
>>> project.tree()
Project (project)
└──Coarse aligner (coarse_alignment)
├──scene (scene)
│ └──cell_with_print_scene(1, 0) (interface_alignment)
│ └──Marker Aligner (marker_alignment)
│ └──structure (structure)
└──scene_1 (scene)
└──cell_with_print_scene(1, 0) (interface_alignment)
└──Marker Aligner (marker_alignment)
└──structure_1 (structure)
>>> # Export your project to a .nano-file.
>>> project.nano(project_name="my_project")
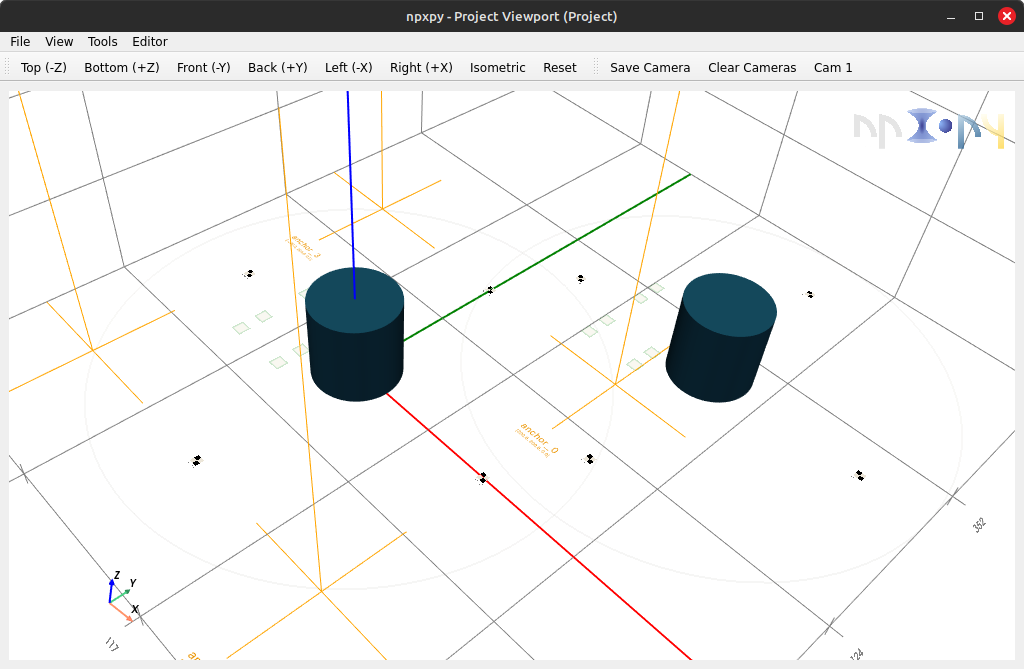
Features like a viewport based on pyvistaqt are also available for keeping track of your project visually as well.
>>> viewport = project.viewport()
Documentation
To view more functionalities and use case examples of npxpy, refer to the the provided documentation.
How to Cite
If npxpy contributes to your research, software, or project, we kindly request that you cite it in your publications using the provided DOI above.
License
This project is licensed under the MIT License - see the LICENSE
file for details.
TL;DR: You may use, modify, and distribute this software freely, provided the license and copyright notice are included.
What This Means for Users and Contributors
-
Freedom to Use: You are free to use this software in any project (commercial, personal, or otherwise) without restrictions, as long as the MIT License terms are met.
-
Modifications and Derivatives: You may modify the code, create derivative works, and distribute them under any license of your choice. The only requirements are:
- Include the original MIT License and copyright notice with your distribution.
- Clearly state any significant changes made to the original code.
-
Linking and Distribution: You may link this software with proprietary code or other open-source projects without restrictions. No obligations apply to the proprietary components of your project.
-
Contribution: By contributing to this project, you agree that your contributions will be licensed under the MIT License. This ensures your changes remain freely usable by others under the same terms.
For more details on your rights and responsibilities under this license, please review the LICENSE file.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file npxpy-0.4.0.tar.gz.
File metadata
- Download URL: npxpy-0.4.0.tar.gz
- Upload date:
- Size: 1.3 MB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8870c3da004aa59155f5279b0bbf0c14a8c00da95db3e641992fa7cc8044b8c2
|
|
| MD5 |
12e22c61c320e18bc8aaee6f9194e4c6
|
|
| BLAKE2b-256 |
ae769525f50da233920b2211c74f6fd1654bd673e36ef080dba9cba1c82a5ffb
|
Provenance
The following attestation bundles were made for npxpy-0.4.0.tar.gz:
Publisher:
release-and-publish-to-pypi.yml on cuenlueer/npxpy
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
npxpy-0.4.0.tar.gz -
Subject digest:
8870c3da004aa59155f5279b0bbf0c14a8c00da95db3e641992fa7cc8044b8c2 - Sigstore transparency entry: 763594821
- Sigstore integration time:
-
Permalink:
cuenlueer/npxpy@21e0cbafb4b17782a7a83cf013cc0bdc05096f1c -
Branch / Tag:
refs/heads/main - Owner: https://github.com/cuenlueer
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release-and-publish-to-pypi.yml@21e0cbafb4b17782a7a83cf013cc0bdc05096f1c -
Trigger Event:
push
-
Statement type:
File details
Details for the file npxpy-0.4.0-py3-none-any.whl.
File metadata
- Download URL: npxpy-0.4.0-py3-none-any.whl
- Upload date:
- Size: 395.9 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
89d7db891badd89a3752ed1d273838e3fb483a12658bc876778ed9e8fe2aa98b
|
|
| MD5 |
7fdd838983ab66a9a43df15e80b43059
|
|
| BLAKE2b-256 |
0dc6760faf354549b01e2a4f8e206e7520672361cfa2a8154715869cc2cbc60d
|
Provenance
The following attestation bundles were made for npxpy-0.4.0-py3-none-any.whl:
Publisher:
release-and-publish-to-pypi.yml on cuenlueer/npxpy
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
npxpy-0.4.0-py3-none-any.whl -
Subject digest:
89d7db891badd89a3752ed1d273838e3fb483a12658bc876778ed9e8fe2aa98b - Sigstore transparency entry: 763594825
- Sigstore integration time:
-
Permalink:
cuenlueer/npxpy@21e0cbafb4b17782a7a83cf013cc0bdc05096f1c -
Branch / Tag:
refs/heads/main - Owner: https://github.com/cuenlueer
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release-and-publish-to-pypi.yml@21e0cbafb4b17782a7a83cf013cc0bdc05096f1c -
Trigger Event:
push
-
Statement type:













