NTxPred2: A method for predicting the neurotoxic activity of peptides and proteins.
Project description
NTxPred2
A method for predicting the neurotoxic activity of peptides and proteins.
📌 Introduction
NTxPred2 is designed to assist researchers in therapeutic peptide and protein development by providing advanced methods for quantifying and classifying neurotoxic peptides and proteins that target the central nervous system.
It employs large language model word embeddings as features for predicting neurotoxic activity. The final model offers Prediction, Protein-Scanning, and Design modules, implemented using machine learning and protein language models.
🔗 Visit the web server for more information: NTxPred2 Web Server
📖 Please cite relevant content for complete details, including the algorithm behind the approach.
📚 Reference
Rathore et al. A large language model for predicting neurotoxic peptides and neurotoxins. #Journal Name#
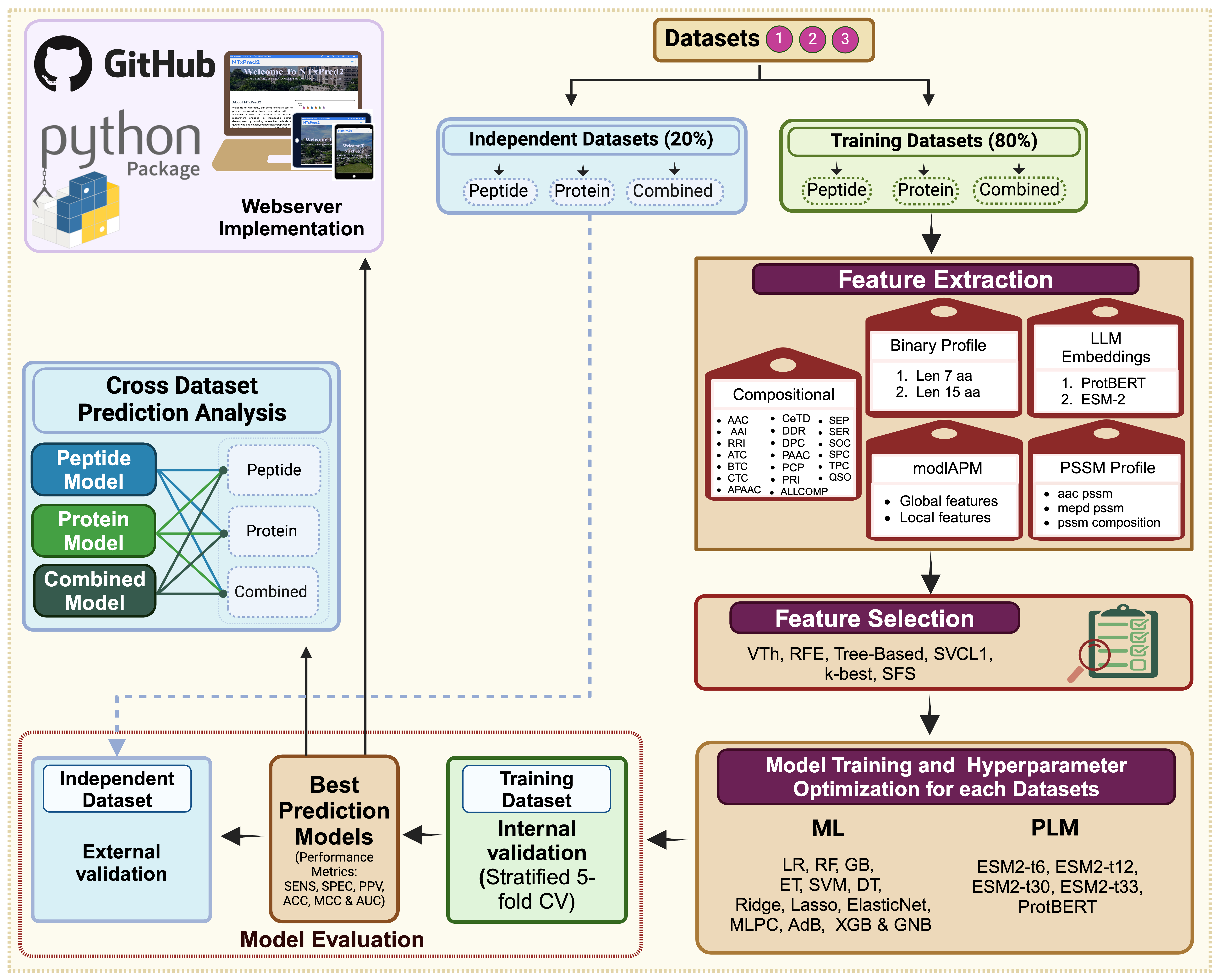
🖼️ NTxPred2 Workflow Representation
🛠️ Installation
🔹 PIP Installation
To install NTxPred2 via PIP, run:
pip install ntxpred2
To check available options, type:
ntxpred2 -h
🔹 Standalone Installation
NTxPred2 is written in Python 3 and requires the following dependencies:
✅ Required Libraries
python=3.10.7
pytorch
Additional required packages:
pip install scikit-learn==1.5.2
pip install pandas==1.5.3
pip install numpy==1.25.2
pip install torch==2.1.0
pip install transformers==4.34.0
pip install joblib==1.4.2
pip install onnxruntime==1.15.1
Bio (Biopython): 1.81
tqdm: 4.64.1
torch: 2.6.0
🔹 Installation using environment.yml
- Create a new Conda environment:
conda env create -f environment.yml
- Activate the environment:
conda activate NTxPred2
⚠️ Important Note
- Due to the large size of the model file, the model directory has been compressed and uploaded.
- Download the zip file from Download Page.
- Extract the file before using the code or model.
🔬 Classification
NTxPred2 classifies peptides and proteins as neurotoxic or non-neurotoxic based on their primary sequence.
🔹 Model Options
- ESM2-t30 (Peptide Model): For sequences 7-50 amino acids long.
- ET (Protein Model): For sequences ≥ 51 amino acids.
- ET (Combined Model): For sequences of mixed length.
- Default Model: ESM2-t30 (Peptide Model), selected for best performance and efficiency.
🚀 Usage
🔹 Minimum Usage
ntxpred2.py -h
To run an example:
ntxpred2.py -i example.fasta
🔹 Full Usage
usage: ntxpred2.py [-h]
[-i INPUT]
[-o OUTPUT]
[-t THRESHOLD]
[-j {1,2,3,4}]
[-m {1,2,3}]
[-d {1,2}]
[-wd WORKING DIRECTORY]
Required Arguments
| Argument | Description |
|---|---|
-i INPUT |
Input: Peptide or protein sequence (FASTA format or simple format) |
-o OUTPUT |
Output file (default: outfile.csv) |
-t THRESHOLD |
Threshold (0-1, default: 0.5) |
-j {1,2,3,4} |
Job type: 1-Prediction, 2-Protein Scanning, 3-Design, 4-Design all possible mutants |
-m {1,2,3} |
Model selection: 1-ESM2-t30 (Peptides), 2-ET (Proteins), 3-ET (Combined) |
-wd WORKING |
Working directory for saving results |
📂 Input & Output Files
✅ Input File Format
NTxPred2 supports two formats:
- FASTA Format: (Example:
example.fasta) - Simple Format: (Example:
example.seq, each sequence on a new line)
✅ Output File
- Results are saved in CSV format.
- If no output file is specified, results are stored in
outfile.csv.
🔍 Jobs & Features
🔹 Job Types
| Job | Description |
|---|---|
| 1️⃣ Prediction | Predicts whether input peptide/protein is neurotoxic or not. |
| 2️⃣ Protein Scanning | Identifies neurotoxic regions in a protein sequence. |
| 3️⃣ Design | Generates mutant peptides/proteins with a single amino acid/dipeptide at a specified position. |
| 4️⃣ Design All Possible Mutants | Generates and predicts all possible mutants. |
🔹 Additional Options
| Option | Description |
|---|---|
-p POSITION |
Position to insert mutation (1-indexed) |
-r RESIDUES |
Mutated residues (single/double letter amino acid codes) |
-w {8-20} |
Window length (Protein Scan mode only, default: 12) |
-d {1,2} |
Display: 1-Neurotoxic only, 2-All peptides (default) |
📑 Package Contents
| File | Description |
|---|---|
| INSTALLATION | Installation instructions |
| LICENSE | License information |
| README.md | This file |
| ntxpred2.py | Python program for classification |
| example.fasta | Example file (FASTA format) |
| example.seq | Example file (Simple format) |
📦 PIP Installation (Again for Reference)
pip install ntxpred2
Check options:
ntxpred2 -h
🚀 Start predicting neurotoxicity with NTxPred2 today!
🔗 Visit: NTxPred2 Web Server
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file ntxpred2-1.1.tar.gz.
File metadata
- Download URL: ntxpred2-1.1.tar.gz
- Upload date:
- Size: 27.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7cb35a939349f6cb671638c8054e686efc9282eba065caedd56198d3e179e90f
|
|
| MD5 |
07ae05d9ce7ca03c4e138238768bec00
|
|
| BLAKE2b-256 |
0cf3ebd16011d8578cc07190b5320b0ca309d23da43e589298783d17d00ab565
|
File details
Details for the file ntxpred2-1.1-py3-none-any.whl.
File metadata
- Download URL: ntxpred2-1.1-py3-none-any.whl
- Upload date:
- Size: 25.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
6557874429b28c6c8c98d22bccd161d20391e6d158cafe8fc9f7d5f60709f6b9
|
|
| MD5 |
6a569377c840ac6ac20ff87f0abdab95
|
|
| BLAKE2b-256 |
39232e2461c26f061e4b7908dbc1127a631e8c12b4b9045a605ffa23b5fc769b
|