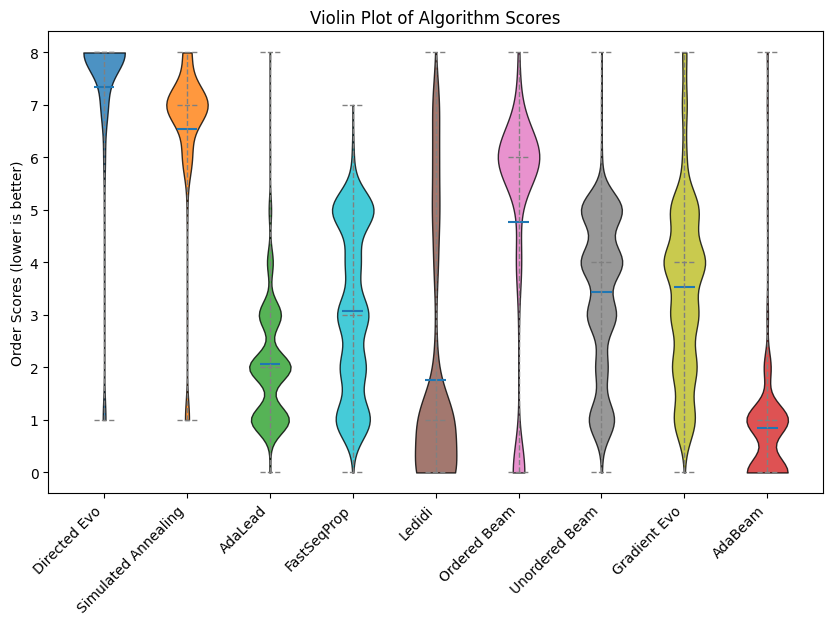
Optimizers from the nucleobench package.
Project description
NucleoBench
This is the initial repo for an upcoming paper, NucleoBench: A Large-Scale Benchmark of Neural Nucleic Acid Design Algorithms.
This repo is covered by the Apache-2.0 license.
This repo is intended to be used in a few ways:
- Reproducing the results from our paper.
- Running the NucleoBench sequence designers on custom problems.
- Using our new designer, AdaBeam, on a custom problem.
To do these, you can clone this repo, use the Docker image (for the benchmark), or use the PyPi package for our designers.
Results
Installation from PyPi
pip install nucleobench
Then you can use it in python:
from nucleobench import optimizations
opt = optimizations.get_optimization('beam_search_unordered') # Any optimizer name.
Installation & testing from GitHub
# Clone the repo.
git clone https://github.com/move37-labs/nucleobench.git
cd nucleobench
# Create and activate the conda environment.
conda env create -f environment.yml
conda activate nucleobench
# Run all the unittests.
pytest nucleobench/
You can also run the integration tests, which require an internet connection:
pytest docker_entrypoint_test.py
Running NucleoBench from PyPi or Docker
See the recipes/colab folder for examples of how to run the designers with PyPi.
See the recipes/docker folder for examples of how to run the designers with Docker.
See the recipes/python folder for examples of how to run the designers with the cloned github repo.
Building a Docker image
To help deploy NucleoBench to the cloud, we've created a docker container. To build it yourself, see the top of Dockerfile for instructions. One way of creating a docker file is:
docker build -t nucleobench -f Dockerfile .
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file nucleopt-0.1.6.tar.gz.
File metadata
- Download URL: nucleopt-0.1.6.tar.gz
- Upload date:
- Size: 46.4 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
27545190a1e08dbbd918d2009b719d82039e9b579c5b63f4eda932461087a13c
|
|
| MD5 |
e456c9533ee9fc56e70e2c2ad80dcc54
|
|
| BLAKE2b-256 |
a30026fd9a793ff5c2a502a5f3bbfc4e065d4784a87b752ab2f5f1482a56f3a1
|
File details
Details for the file nucleopt-0.1.6-py3-none-any.whl.
File metadata
- Download URL: nucleopt-0.1.6-py3-none-any.whl
- Upload date:
- Size: 65.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e67190a415010b32c1c9f1423ea42de397951d183fb323ccd569c86b1a9073fc
|
|
| MD5 |
a1f0828c04b3a2205c0c561b8b777fab
|
|
| BLAKE2b-256 |
b617ea56fdbc5620afb362b98b7b6fd70a152a381bf4c8e7aed9daa1f0a23704
|