Generalizable UMAP Implementation
Project description
NUMAP
This is the official PyTorch implementation of NUMAP, a new and generalizable UMAP implementation, from the paper "Generalizable Spectral Embedding with Applications to UMAP".
See our GitHub repository for more information and the latest updates.
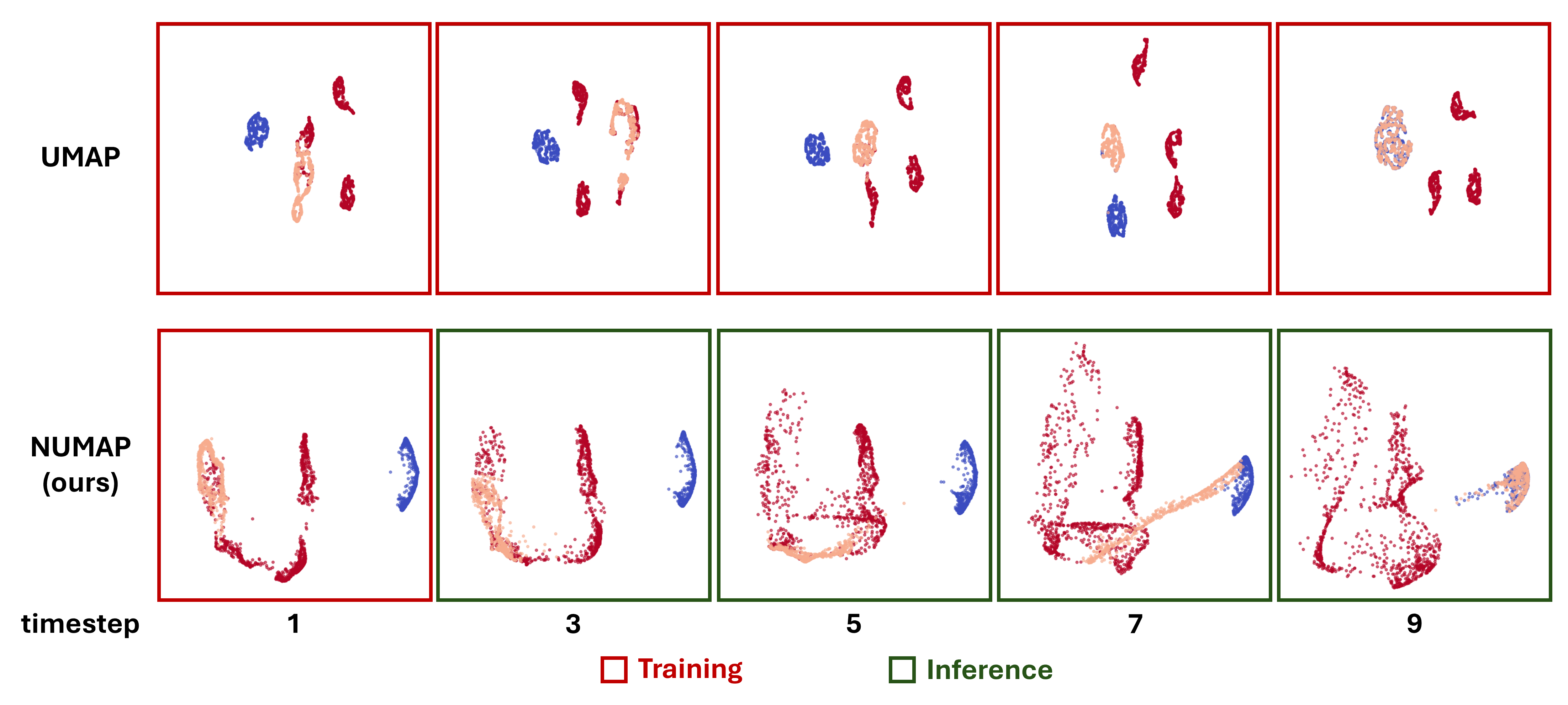
NUMAP can be used to visualize many types of data in a low-dimensional space, while enabling a simple out-of-sample extension. One application of NUMAP is to visualize time-series data, and help understand the process in a given system. For example, the following figure shows the transition of a set of points from one state to another, using NUMAP. In a biological point of view, this can be viewed as a simplified simulation of the cellular differentiation process.
The package is based on UMAP and GrEASE (Generalizable and Efficient Approximate Spectral Embedding). It is easy to use and can be used with any PyTorch dataset, on both CPU and GPU. The package also includes a test dataset and a test script to run the model on the 2 Circles dataset.
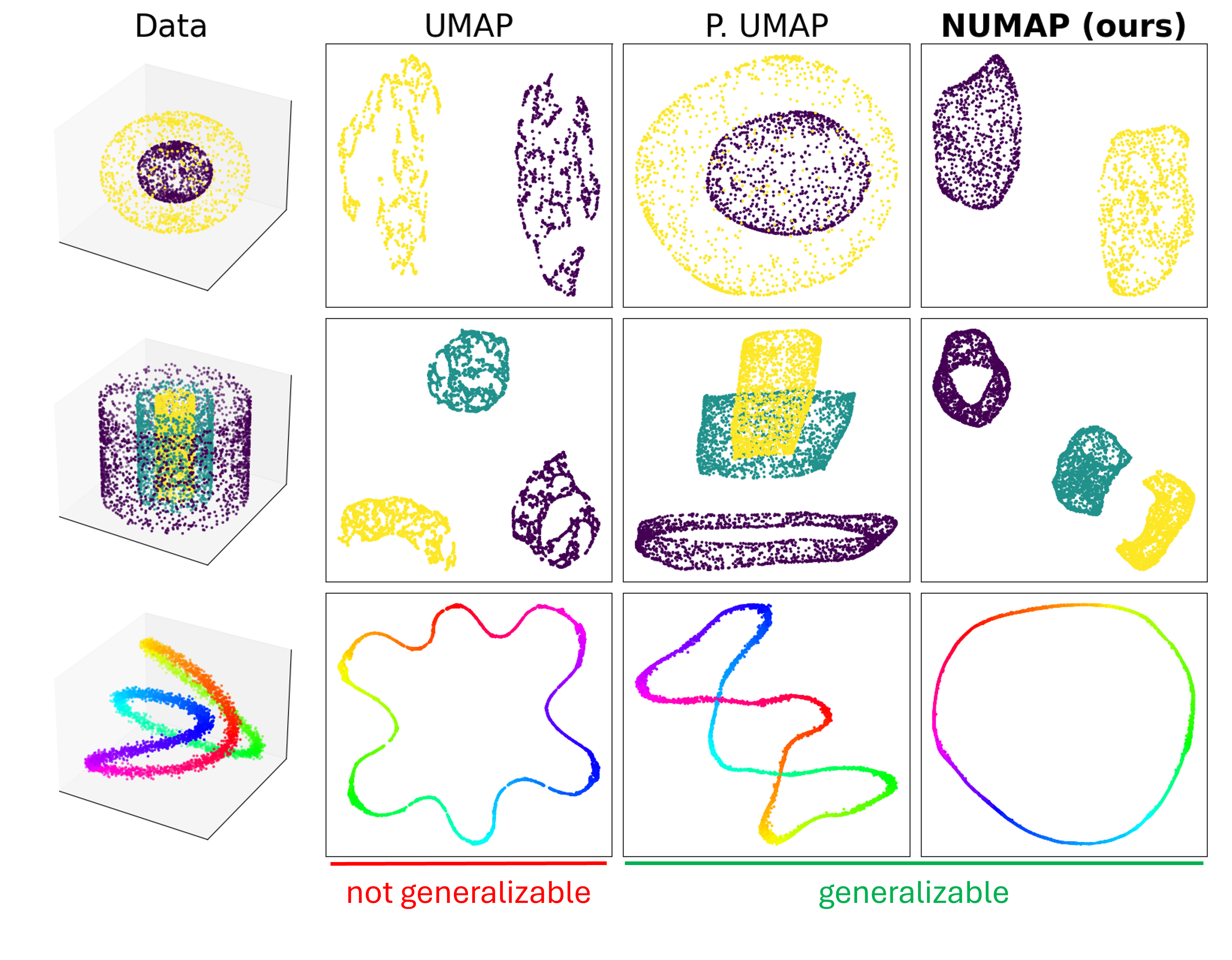
The incorporation of GrEASE enables preservation of both local and global structures of the data, as UMAP, with the new capability of out-of-sample extension.
Installation
To install the package, simply use the following command:
pip install numap
Usage
The basic functionality is quite intuitive and easy to use, e.g.,
from numap import NUMAP
numap = NUMAP(n_components=2) # n_components is the number of dimensions in the low-dimensional representation
numap.fit(X) # X is the dataset and it should be a torch.Tensor
X_reduced = numap.transfrom(X) # Get the low-dimensional representation of the dataset
Y_reduced = numap.transform(Y) # Get the low-dimensional representation of a test dataset
You can read the code docs for more information and functionalities.
Running examples
In order to run the model on the 2 Circles dataset, you can either run the file, or using the command-line command:
python tests/run_numap.py
This will run NUMAP and UMAP on the 2 Circles dataset and plot the results.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file numap-0.2.2.tar.gz.
File metadata
- Download URL: numap-0.2.2.tar.gz
- Upload date:
- Size: 15.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.12.6
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
bceb90f7a9fb499c94754f4c3670a6fa39c0b6ae4c99820264e687eef25a3a0a
|
|
| MD5 |
17371cee0574623cc86ca0bc8ebd8b22
|
|
| BLAKE2b-256 |
3e81a11fb5b7c16b0b8de5e660a933801676cc6763ef137542b9f64ba1f257ae
|
File details
Details for the file numap-0.2.2-py3-none-any.whl.
File metadata
- Download URL: numap-0.2.2-py3-none-any.whl
- Upload date:
- Size: 22.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.12.6
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9c250a4afa03229b8a1721d791153b9f3c6bc811ff784717cc112599f6b1a177
|
|
| MD5 |
fadcf1f988edc47fe8b434a42b9d4f73
|
|
| BLAKE2b-256 |
9045ff3c9a7d0a647626fc7c1cdb8fe734335939a3c389576834921dad4aba19
|