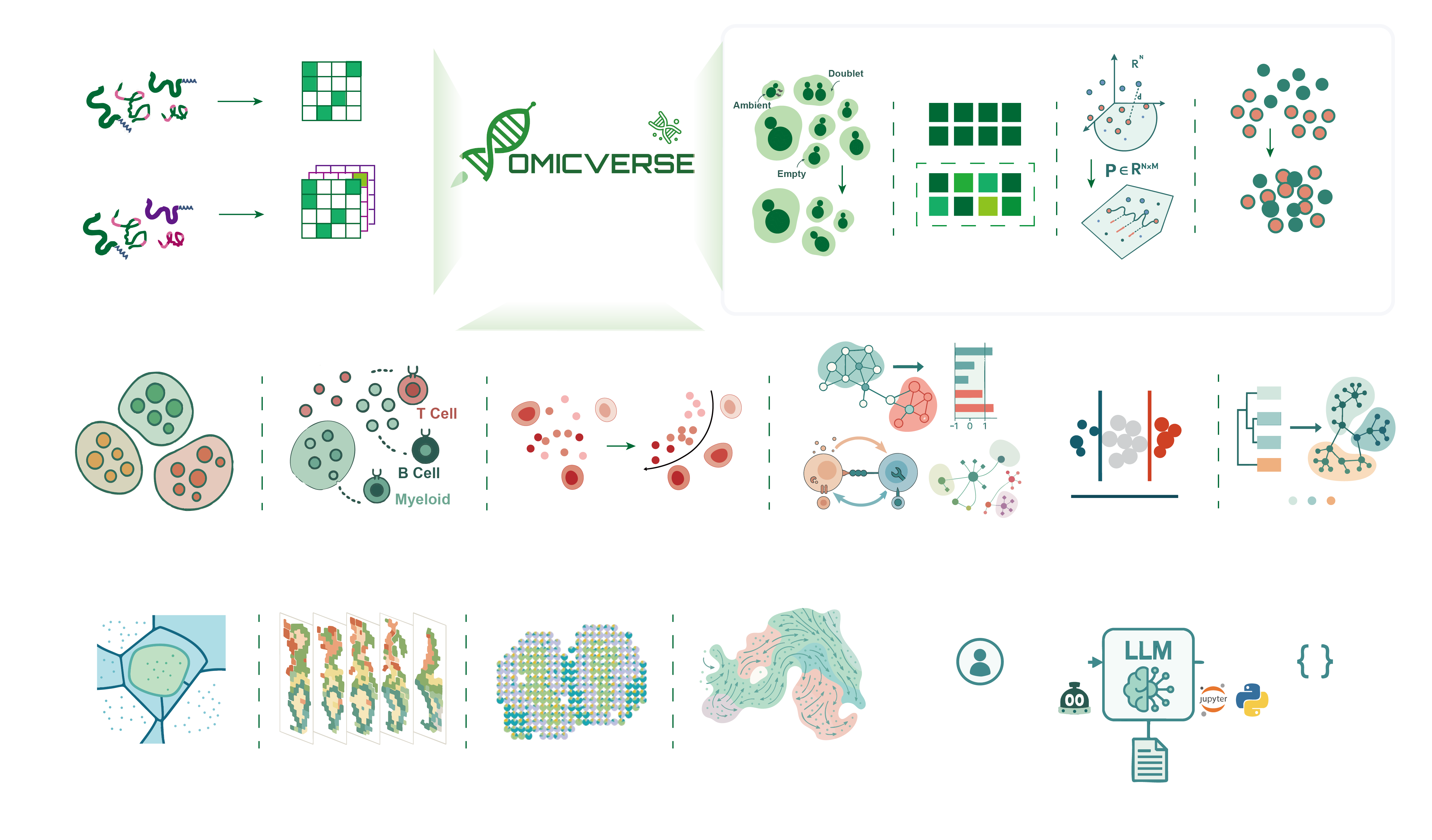
OmicVerse: A single pipeline for exploring the entire transcriptome universe
Project description

OmicVerse is the fundamental package for multi omics included bulk ,single cell and spatial RNA-seq analysis with Python. For more information, please read our paper: OmicVerse: a framework for bridging and deepening insights across bulk and single-cell sequencing
[!IMPORTANT]
Star Us, You will receive all release notifications from GitHub without any delay ~ ⭐️
If you like OmicVerse and want to support our mission, please consider making a 💗donation to support our efforts.
Star History

1 Introduction
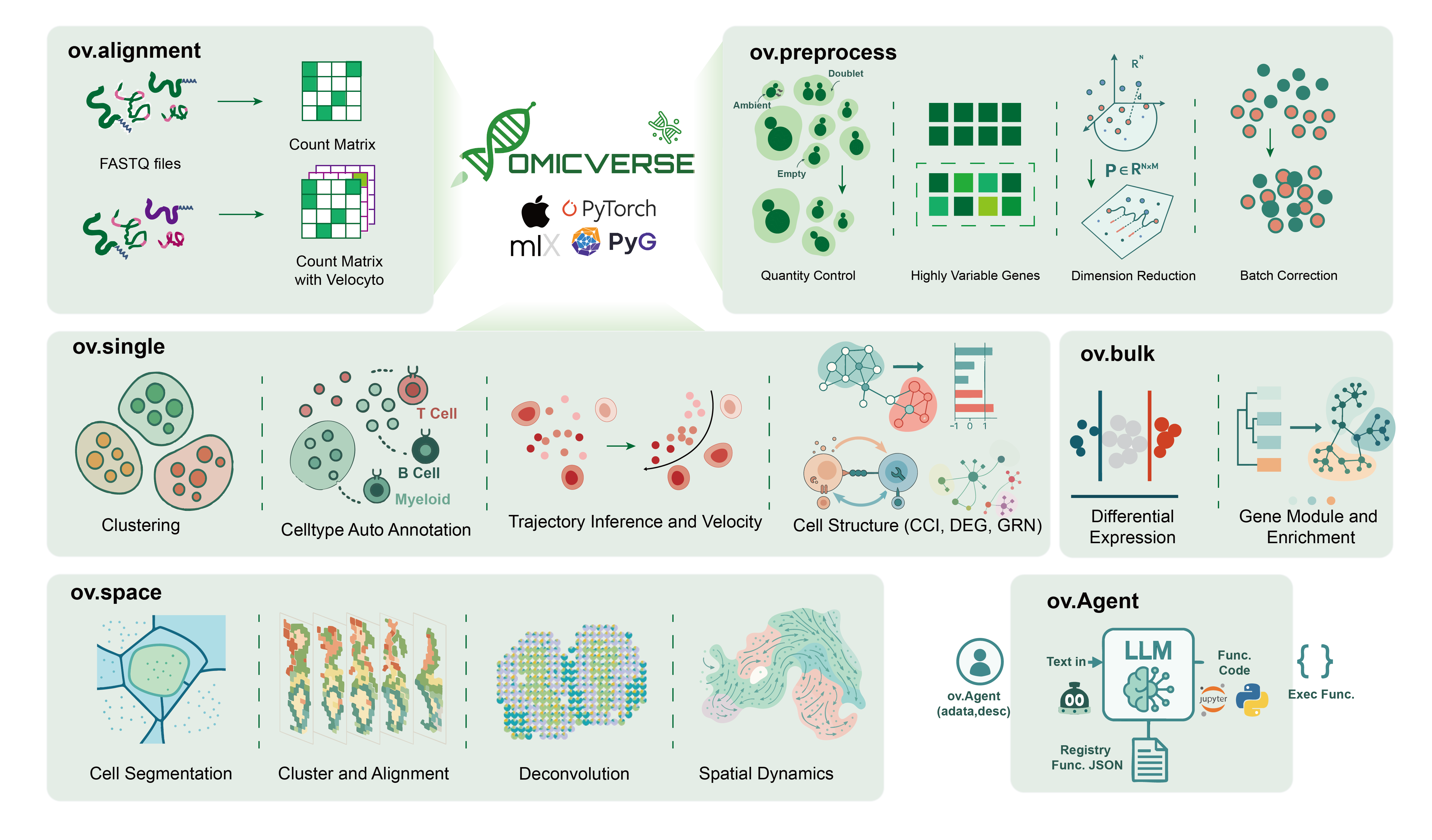
OmicVerse v2 is a unified Python project for modern transcriptomics and multi-omics analysis. It brings together bulk RNA-seq, single-cell, spatial transcriptomics, downstream visualization, model-based analysis, and AI-assisted workflows in one package and documentation system.
[!NOTE] OmicVerse v2 is organized as a broader analysis platform rather than a single-method package. In addition to core analysis modules, it now includes agent-style workflows through J.A.R.V.I.S., MCP-based tool serving for AI clients, and a growing documentation/tutorial system under
omicverse_guide.
2 Directory structure
.
├── omicverse # Main Python package
├── omicverse_guide # Documentation files
├── omicverse_web # Web Analysis Platform
├── sample # Some test data
├── LICENSE
└── README.md
3 General Getting Started
OmicVerse can be installed via conda or pypi and you need to install pytorch at first. Please refer to the installation tutorial for more detailed installation steps and adaptations for different platforms (Windows, Linux or Mac OS).
You can use conda install omicverse -c conda-forge or pip install -U omicverse for installation.
Please checkout the documentations and tutorials at omicverse page or omicverse.readthedocs.io.
4 J.A.R.V.I.S Getting Started
4.1 OpenClaw
OmicVerse provide an directly interact analysis with OpenClaw project. You can use
omicverse claw 'help me annotate the lung scrna-seq'
This module supported by ov.Agent function. And i think that will be convinent for you to analysis the anndata using OpenClaw.
The full tutorial could be found at here
4.2 MCP Server (Model Context Protocol)
OmicVerse provide an MCP server that exposes registered analysis tools to AI assistants (Claude Code, etc.) via the standard Model Context Protocol.
# Install with MCP dependencies
pip install -e "omicverse[mcp]"
# Start the server (stdio transport)
python -m omicverse.mcp # or: omicverse-mcp
python -m omicverse.mcp --phase P0 # core pipeline tools only
The full tutorial could be found at here
4.3 J.A.R.V.I.S Msg system
If you want to analysis the AnnData using Mobile Phone, maybe you can try
# Install with jarvis dependencies
pip install "omicverse[jarvis]"
# Start to chat with JARVIS using telegram
omicverse jarvis --channel telegram --token "$TELEGRAM_BOT_TOKEN"
The full tutorial could be found at here
4 Data Framework and Reference
The omicverse is implemented as an infrastructure based on the following four data structures.
The table contains the tools have been published
| Scanpy 📦 📖 |
dynamicTreeCut 📦 📖 |
scDrug 📦 📖 |
MOFA 📦 📖 |
COSG 📦 📖 |
CellphoneDB 📦 📖 |
| AUCell 📦 📖 |
Bulk2Space 📦 📖 |
SCSA 📦 📖 |
WGCNA 📦 📖 |
StaVIA 📦 📖 |
pyDEseq2 📦 📖 |
| NOCD 📦 📖 |
SIMBA 📦 📖 |
GLUE 📦 📖 |
MetaTiME 📦 📖 |
TOSICA 📦 📖 |
Harmony 📦 📖 |
| Scanorama 📦 📖 |
Combat 📦 📖 |
TAPE 📦 📖 |
SEACells 📦 📖 |
Palantir 📦 📖 |
STAGATE 📦 📖 |
| scVI 📦 📖 |
MIRA 📦 📖 |
Tangram 📦 📖 |
STAligner 📦 📖 |
CEFCON 📦 📖 |
PyComplexHeatmap 📦 📖 |
| STT 📦 📖 |
SLAT 📦 📖 |
GPTCelltype 📦 📖 |
PROST 📦 📖 |
CytoTrace2 📦 📖 |
GraphST 📦 📖 |
| COMPOSITE 📦 📖 |
mellon 📦 📖 |
starfysh 📦 📖 |
COMMOT 📦 📖 |
flowsig 📦 📖 |
pyWGCNA 📦 📖 |
| CAST 📦 📖 |
scMulan 📦 📖 |
cellANOVA 📦 📖 |
BINARY 📦 📖 |
GASTON 📦 📖 |
pertpy 📦 📖 |
| inmoose 📦 📖 |
memento 📦 📖 |
GSEApy 📦 📖 |
marsilea 📦 📖 |
scICE 📦 📖 |
sude 📦 📖 |
| GeneFromer 📦 📖 |
scGPT 📦 📖 |
scFoundation 📦 📖 |
UCE 📦 📖 |
CellPLM 📦 📖 |
kb_python 📦 📖 |
| Scaden 📦 📖 |
BayesPrime 📦 📦 📖 |
InstaPrime 📦 📖 |
Cellpytist 📦 📖 |
latentvelo 📦 📖 |
graphvelo 📦 📖 |
| scvelo 📦 📖 |
Dynamo 📦 📖 |
CONCORD 📦 📖 |
FlashDeconv 📦 📖 |
Hospot 📦 📖 |
Banksy 📦 📖 |
| STAR 📦 📖 |
fastp 📦 📖 |
featureCounts 📦 📖 |
edgeR 📦 📖 |
spaco 📦 📖 |
Included Package not published or preprint
- [1] Cellula is to provide a toolkit for the exploration of scRNA-seq. These tools perform common single-cell analysis tasks
- [2] pegasus is a tool for analyzing transcriptomes of millions of single cells. It is a command line tool, a python package and a base for Cloud-based analysis workflows.
- [3] cNMF is an analysis pipeline for inferring gene expression programs from single-cell RNA-Seq (scRNA-Seq) data.
5 Contact
- Zehua Zeng (starlitnightly@gmail.com or zehuazeng@xs.ustb.edu.cn)
- Lei Hu (hulei@westlake.edu.cn)
6 Developer Guild and Contributing
If you would like to contribute to omicverse, please refer to our developer documentation.
Running tests locally
Install the test dependencies and execute the suite with pytest:
pip install -e .[tests]
# or install the pinned latest requirements
pip install -r requirements-latest.txt
pytest
The optional tests extra and the requirements-latest.txt file now include pytest-asyncio>=0.23, which is required for the asynchronous streaming tests under tests/utils/.
 |
|---|
[!IMPORTANT]
We would like to thank the following WeChat Official Accounts for promoting Omicverse.
7 Citation
If you use omicverse in your work, please cite the omicverse publication as follows:
OmicVerse: a framework for bridging and deepening insights across bulk and single-cell sequencing
Zeng, Z., Ma, Y., Hu, L. et al.
Nature Communication 2024 Jul 16. doi: 10.1038/s41467-024-50194-3.
Here are some other related packages, feel free to reference them if you use them!
CellOntologyMapper: Consensus mapping of cell type annotation
Zeng, Z., Wang, X., Du, H. et al.
imetaomics 2025 Nov 06. doi: 10.1002/imo2.70064.
8 Other
If you would like to sponsor the development of our project, you can go to the afdian website (https://ifdian.net/a/starlitnightly) and sponsor us.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file omicverse-2.1.1.tar.gz.
File metadata
- Download URL: omicverse-2.1.1.tar.gz
- Upload date:
- Size: 12.0 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.9.10
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
4a17c450c6763741c5b29b04fc7234b6f9d0fb606f1a53050ad3546aef577a26
|
|
| MD5 |
81d45bb5555e88e103932926ab6e6d38
|
|
| BLAKE2b-256 |
576562a0e2461f6a98da138d1e44e46c1f243d3a2265cb0df36b2d83b2125ee2
|
File details
Details for the file omicverse-2.1.1-py3-none-any.whl.
File metadata
- Download URL: omicverse-2.1.1-py3-none-any.whl
- Upload date:
- Size: 12.8 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.9.10
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
b575889b00fcb383d2d10e583c4db698daaa2a69b1a03cdceba4e837f38a87b9
|
|
| MD5 |
8d1a86265e838f86f4c0ae05dd96d519
|
|
| BLAKE2b-256 |
b93f2aa5abaac5da3ef52954688a18b141e593051c5ba1ad40ab1004bc0bf997
|

























