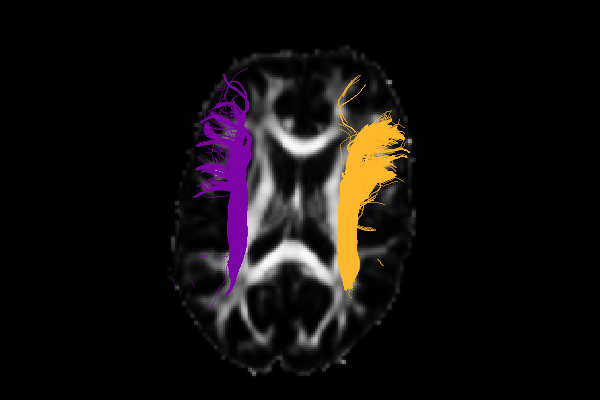
OpenDIVE (Open Diffusion Imaging Visualization for Everyone) is a command line interface tool for generating accessible, interpretable visualizations from diffusion MRI.
Project description
OpenDIVE
OpenDIVE (Open Diffusion Imaging Visualization for Everyone) is a command line interface tool for generating accessible, interpretable visualizations from diffusion MRI, initiated at BrainHack Vandy 2025.
Despite the prolific availability of software tools to visualize diffusion MRI data, there is no standardized visualization to summarize longitudinal changes. Similarly, the current standard of visualizations are not accessible to people with common forms of colorblindness. We propose a software package to both standardize and improve accessibility to representations of diffusion data.
Installation
You can install the package using Python (3.10+):
pip install open-dive
Usage
After installing the package, you should be able to use the open-dive command to produce images.
# Save a T1-weighted image with no interactive visualization
open-dive -n t1.nii.gz -s 50 -o coronal --save_path slice_50_coronal.png --headless
# Custom colorbar
open-dive -n dwi.nii.gz --size 800 600 --value_range 0 1500 --scalar_colorbar --volume_idx 1
# Overlay values on tractography
open-dive -n fa.nii.gz --tractography_path my_tractogram1.trk my_tractogram2.trk --tractography_values -0.5 0.7
# Overlay glass brain and tensor glyphs with custom viewing angles
open-dive -n dwi.nii.gz \
--volume_idx 0 \
--tensor_path tensor.nii.gz \
--glass_brain_path mask.nii.gz \
-o sagittal \
--az 45 \
--el 60 \
-scale 2
Please see the wiki for documentation on using the command.
Contributing
We welcome issues and pull requests! For details on contributing, please see CONTRIBUTING.md.
Aims
- We aim to generate a standardized display for anatomical images to overlay the diffusion models based on user input, including support for multiple track files, color maps, and illustration of bundle summary metrics (p-value, volume, effect size, etc.) per bundle.
- We aim to propose a colorblind-friendly colormap for diffusion MRI images.
For FAQs related to diffusion MRI, see our diffusion FAQ discussion.
Contributors
- Adam Saunders
- Elyssa McMaster
- Chris Rorden
- Johaan Kathilankal Jis
- Minyi Sun
- Adam Sadriddinov
- Lukas VanTilburg
- Kurt Schilling
License
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file open_dive-0.3.0.tar.gz.
File metadata
- Download URL: open_dive-0.3.0.tar.gz
- Upload date:
- Size: 11.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.7.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
4daa771f4720c79e12006a5cdc8bdc6fbcd243f472d5d3f6f5c21ae168b69f1a
|
|
| MD5 |
f36c3796dae15c470447681d0b808643
|
|
| BLAKE2b-256 |
dbd13a129276ffecc6c6d1c3927f3c1d751b22ca08ede378ac0c0978db69049e
|
File details
Details for the file open_dive-0.3.0-py3-none-any.whl.
File metadata
- Download URL: open_dive-0.3.0-py3-none-any.whl
- Upload date:
- Size: 11.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.7.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c4ddc321b530b0f00824e449c7b236c3778e713669352be71268ddf0b20a5b46
|
|
| MD5 |
97d30cb238d54a9da449abf50861e0e3
|
|
| BLAKE2b-256 |
2279c71242f84128de05a6c649ee5aaf1eee230f5e3bcd86acbf29c2990b1b7a
|