open-sourced Deep Visual Proteomics toolkit
Project description
OpenDVP
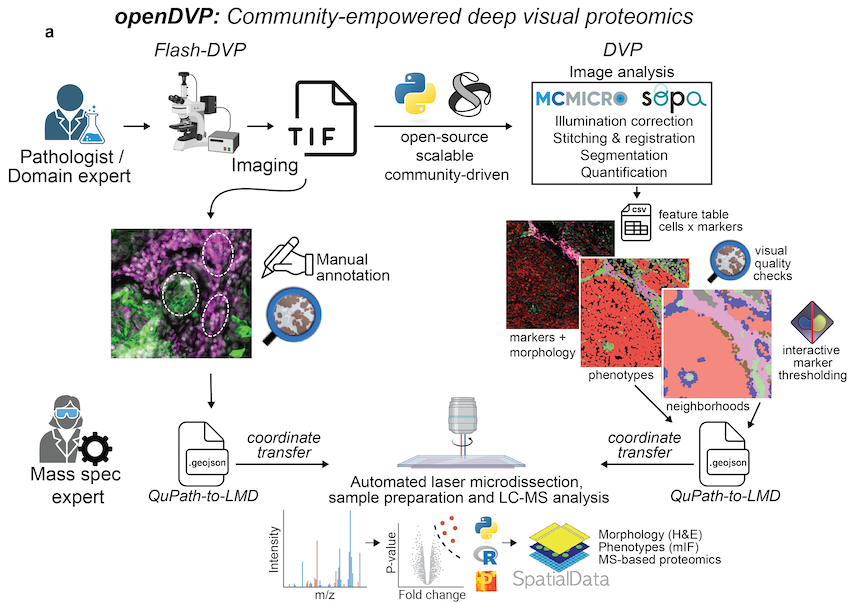
Overview
OpenDVP is an open-source framework designed to support Deep Visual Proteomics (DVP) across multiple modalities using community-supported tools. OpenDVP empowers researchers to perform Deep Visual Proteomics using open-source software. It integrates with community data standards such as AnnData and SpatialData to ensure interoperability with popular analysis tools like Scanpy, Squidpy, and Scimap.
Getting started
Please refer to the documentation, particularly the API documentation.
Installation
You will need Python 3.11 or 3.12 installed on your system. If you are new to creating Python environments, we suggest you use uv or pixi.
You can install openDVP via pip:
conda create --name opendvp -y python=3.12
pip install opendvp
To install the latest version:
pip install git+https://github.com/CosciaLab/openDVP.git@main
Tutorials
To understand what are the applications of openDVP, please check our
Tutorials.
Briefly, they introduce users to (1) Image analysis, (2) downstream proteomic analysis, and (3) Integration of imaging with proteomic data. Please download our Demo Dataset to best follow the tutorials :)
Community & Discussions
We are excited to hear from you and together we can improve spatial protemics.
We welcome questions, feedback, and community contributions!
Join the conversation in the GitHub Discussions tab.
Citation
Please cite the BioArxiv:
Nimo, J., Fritzsche, S., Valdes, D. S., Trinh, M., Pentimalli, T., Schallenberg, S., Klauschen, F., Herse, F., Florian, S., Rajewsky, N., & Coscia, F. (2025). OpenDVP: An experimental and computational framework for community-empowered deep visual proteomics [Preprint]. bioRxiv. https://doi.org/10.1101/2025.07.13.662099
Motivation
Deep Visual Proteomics (DVP) combines high-dimensional imaging, spatial analysis, and machine learning to extract complex biological insights from tissue samples. However, many current DVP tools are locked into proprietary formats, restricted software ecosystems, or closed-source pipelines that limit reproducibility, accessibility, and community collaboration.
- Work transparently across modalities and analysis environments
- Contribute improvements back to a growing ecosystem
- Avoid vendor lock-in for critical workflows
Qupath-to-LMD
Qupath to lmd is a tool we use to make it as easy as possible to go from QuPath annotations to LMD contours Check our Qupath-to-LMD Webapp, or watch our Youtube tutorial:
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file opendvp-0.7.3.tar.gz.
File metadata
- Download URL: opendvp-0.7.3.tar.gz
- Upload date:
- Size: 87.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
ee4f48b29591ddae7026b5f5ecf36c80c998cee8133e1801ab6d665358080d83
|
|
| MD5 |
383785c78fa1e40d2bc3d53af1ad043a
|
|
| BLAKE2b-256 |
72ccd720d612b1d5bb3d68096c3c44fb38384bcfae7eaf6ce46265213886a1dd
|
File details
Details for the file opendvp-0.7.3-py3-none-any.whl.
File metadata
- Download URL: opendvp-0.7.3-py3-none-any.whl
- Upload date:
- Size: 102.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
745c3511f41780804425af7046cf21d8e03145cceb019fd3e21d94e8d2f88c80
|
|
| MD5 |
fe9002e38af605a3fedde8035b0d57d9
|
|
| BLAKE2b-256 |
34ee156304416d107450d31f372ff4bb166f8e5b64e82f7002a00a7596deae67
|