Python bindings for OpenImpala — transport-property computation on 3-D voxel images
Project description
OpenImpala

OpenImpala is a high-performance computing framework for image-based modelling, built upon the AMReX library for massive parallelism using MPI. It tackles the challenge posed by large 3D imaging datasets (often billions of voxels) common in materials science and tomography.
OpenImpala directly solves physical equations, such as steady-state diffusion or conduction problems, on the voxel grid of the input image using finite differences. This approach bypasses the need for explicit mesh generation, working directly with the acquired image data. From the simulation results, it calculates effective homogenised transport properties (e.g., diffusivity, conductivity, tortuosity) characteristic of the microstructure.
These calculated coefficients can directly parameterize continuum-scale models, notably battery simulators like PyBamm and DandeLiion. This capability effectively bridges microstructural details obtained from imaging to device-level performance predictions. OpenImpala is designed for excellent scalability on distributed memory systems, making the analysis of large, high-resolution datasets feasible.
Table of Contents
- Features
- Continuous Integration and Delivery
- Getting Started (Recommended: Apptainer/Singularity)
- Python Installation (pip)
- Building from Source (for Developers)
- Native Installation (Advanced)
- Batch Processing (HPC)
- Example
inputsFile - Output
- Visualisation
- Applications & Related Publications
- Contributing
- Code Formatting
- Static Analysis
- Citation
- License
- Acknowledgements
- Contact & Support
Features
- Calculates effective transport properties (e.g., effective diffusivity, electrical/thermal conductivity) based on steady-state physics.
- Operates directly on segmented 3D image stacks (TIFF, HDF5, DAT support via internal readers).
- Massively parallel using MPI via the AMReX framework.
- Finite difference / finite volume method on the voxel grid.
- Output usable for parameterizing continuum models (e.g., PyBamm, DandeLiion).
- Includes tools for Volume Fraction calculation and basic image reading tests.
Continuous Integration and Delivery
This project uses GitHub Actions for automated building, testing, and releasing.
-
Build & Test CI (
build-test.yml): On every push and pull request, this workflow automatically builds the code and runs the full test suite inside a containerized environment to ensure correctness. Build and test logs are available as artifacts for debugging. -
Build & Release (
release.yml): When a new version is tagged for release, this workflow automatically builds the final Apptainer/Singularity container (.sif) and attaches it to a new GitHub Release. This is the official, recommended way to get the software.
Getting Started (Recommended: Apptainer/Singularity)
The easiest way to use OpenImpala is by downloading the pre-built container from the official GitHub Releases.
1. Download the Container
- Go to the GitHub Releases Page.
- Find the latest release and download the
.siffile (e.g.,openimpala-vX.Y.Z.sif) from the "Assets" section. - Ensure you have Apptainer or Singularity (version 3.x or later) installed on your system. See the Apptainer documentation.
2. Run the Application
The main application is Diffusion, configured via an inputs file. Use the -B (--bind) flag to mount your current working directory (containing your inputs file and data) into the container.
# Define the SIF file you downloaded
SIF_FILE="openimpala-vX.Y.Z.sif"
# --- Run Sequentially ---
# Mounts the current directory into /data inside the container
apptainer exec -B "$(pwd):/data" ${SIF_FILE} /usr/local/bin/Diffusion /data/inputs
# --- Run in Parallel with MPI ---
# Ensure OMP_NUM_THREADS=1 if using multiple MPI ranks
export OMP_NUM_THREADS=1
mpirun -np 4 apptainer exec -B "$(pwd):/data" ${SIF_FILE} /usr/local/bin/Diffusion /data/inputs
Python Installation (pip)
OpenImpala provides Python bindings that can be installed directly from PyPI. This gives you a high-level, NumPy-native API for computing transport properties on 3D voxel images without needing to write C++ code or manage input files.
Prerequisites
OpenImpala uses MPI for parallel computation. You must have a working MPI installation before installing:
# Ubuntu / Debian
sudo apt install libopenmpi-dev
# Fedora / RHEL / Rocky
sudo dnf install openmpi-devel
# then add to PATH: export PATH=/usr/lib64/openmpi/bin:$PATH
# macOS (Homebrew)
brew install open-mpi
# Conda (any platform)
conda install -c conda-forge openmpi
Note: The pre-built wheels are built against OpenMPI. Other MPI implementations (MPICH, Intel MPI) may work but are not tested. If you encounter MPI-related errors at runtime, ensure your MPI installation is ABI-compatible with OpenMPI.
Install
pip install openimpala
GPU acceleration is automatic. If you have an NVIDIA GPU and CuPy installed, OpenImpala detects it at runtime and offloads compute kernels to the GPU. No separate package is needed:
# Optional: install CuPy for automatic GPU acceleration
pip install cupy-cuda12x # match your CUDA toolkit version
If CuPy is not available, OpenImpala falls back to SciPy on the CPU.
Advanced / HPC: For clusters needing compiled C++ HYPRE solvers with native CUDA support:
pip install openimpala-cuda
The openimpala-cuda wheel requires a working NVIDIA CUDA 12 runtime (driver
- toolkit). On Colab, Kaggle, and most cluster nodes this is already present.
To install with optional dependencies:
# MPI-aware Python (recommended)
pip install openimpala[mpi]
# All optional dependencies
pip install openimpala[all]
Quick Start
import numpy as np
from openimpala import Session, volume_fraction, tortuosity
# All computation must happen inside a Session context manager,
# which initialises MPI and AMReX.
with Session():
# Create a simple 3D test image (32x32x32, phase 0 = pore, phase 1 = solid)
image = np.zeros((32, 32, 32), dtype=np.int32)
image[:, :, 16:] = 1 # half pore, half solid
vf = volume_fraction(image, phase=0)
print(f"Volume fraction: {vf.value:.4f}")
tau = tortuosity(image, phase=0, direction="x")
print(f"Tortuosity: {tau.value:.4f}")
CLI
OpenImpala also provides a command-line interface:
# Analyse a TIFF image stack
openimpala analyze my_sample.tif --phase 0 --direction x
# Volume fraction only
openimpala vf my_sample.tif --phase 0
Building from Source (for Developers)
If you are developing OpenImpala and need to compile your changes, you can replicate the CI build process locally. This workflow uses a two-stage container build.
-
Build the Dependency Container: First, build the container that holds all the compilers and libraries (MPI, Hypre, AMReX, etc.). This is based on the
Singularity.deps.defrecipe.# This command needs root privileges to install packages inside the container sudo apptainer build dependency_image.sif containers/Singularity.deps.def
-
Compile OpenImpala: Now, execute the
makecommand inside the dependency container you just built. This will compile your local source code using the tools from the container.# Mount your local project directory to /src inside the container and run make apptainer exec --bind "$(pwd):/src" dependency_image.sif bash -c "cd /src && make all -j"
The compiled executables will appear in the
build/directory on your local filesystem. -
Run Tests: To run the test suite, use the same container.
apptainer exec --bind "$(pwd):/src" dependency_image.sif bash -c "cd /src && make test"
Native Installation (Advanced)
Building natively requires manually installing all dependencies.
Dependencies
- A modern C++ compiler supporting C++17 (e.g., GCC >= 7, Clang >= 6).
- A Fortran compiler compatible with your C++ compiler and MPI library (e.g., gfortran).
- CMake (version 3.10 or later recommended).
- An MPI library implementation (e.g., OpenMPI, MPICH, Intel MPI).
- AMReX Library: Core dependency (https://github.com/AMReX-Codes/amrex). Ensure
AMREX_HOMEis set or AMReX is findable by CMake. - HYPRE Library: Required for the default linear solver (https://github.com/hypre-space/hypre). Ensure
HYPRE_HOMEis set or Hypre is findable. - LibTIFF Library: Required for reading TIFF input files (development package needed, e.g.,
libtiff-dev,libtiff-devel). - HDF5 Library: Required for reading HDF5 input datasets (C and C++ bindings, development package needed). Needed if building with HDF5 support.
- (Optional) Boost Filesystem: May be required depending on internal path handling.
Building from Source (CMake)
-
Clone the repository:
git clone [https://github.com/kramergroup/openImpala.git](https://github.com/kramergroup/openImpala.git) cd openImpala
-
Configure using CMake: Create a build directory.
mkdir build && cd build # Basic configuration: cmake .. -DCMAKE_INSTALL_PREFIX=../install -DCMAKE_BUILD_TYPE=Release # Or add paths to dependencies if not found automatically: # cmake .. -DAMReX_DIR=/path/to/amrex/lib/cmake/AMReX -DHYPRE_DIR=/path/to/hypre/lib/cmake/hypre
- Use
ccmake ..orcmake-gui ..for interactive configuration. - Set
CMAKE_CXX_COMPILER/CMAKE_Fortran_COMPILER(e.g., tompicxx,mpif90).
- Use
-
Compile the code:
make -j4 # Adjust job count
-
Run Tests (Optional):
make test
-
Install (Optional):
make installThe executable (e.g.,
Diffusion) will be inbuild/apps/.
Batch Processing (HPC)
The Diffusion application runs non-interactively, making it suitable for HPC batch jobs. An example SLURM script:
#!/bin/bash
#SBATCH --nodes=1
#SBATCH --ntasks-per-node=20
#SBATCH --time=01:00:00
#SBATCH --job-name=openimpala_diffusion
module load apptainer # Or singularity, depending on your HPC
export OMP_NUM_THREADS=1
# Navigate to your simulation directory
cd /path/to/your/simulation/directory/
# Define path to the downloaded SIF image
SIF_IMAGE=/path/on/hpc/to/openimpala-vX.Y.Z.sif
# Define path to the executable *inside* the container
APP=/usr/local/bin/Diffusion
# Define path to your inputs file
INPUT_FILE=./inputs
# Run the calculation, binding the current directory for I/O
mpirun -np $SLURM_NTASKS apptainer exec \
-B "$(pwd):$(pwd)" $SIF_IMAGE $APP $INPUT_FILE
echo "Job Finished"
Note: Adjust module load, paths (SIF_IMAGE, INPUT_FILE, cd), and mpirun flags according to your specific HPC environment.
Example inputs File
(This section moved here for better flow after installation)
The Diffusion application reads parameters from a text file (commonly named inputs). Create one based on this example (also found in apps/diffusion/inputs in the source tree):
# Example input file for Diffusion application
# --- Input File ---
filename = "data/SampleData_2Phase.tif" # Input file relative to execution or data_path
data_path = "." # Path prefix for filename
# Required for HDF5:
# hdf5_dataset = "exchange/data"
# Required for RAW/DAT:
# raw_width = 100; raw_height = 100; raw_depth = 100; raw_datatype = "UINT8"
# Required for TIFF sequence:
# tiff_stack_size = 100 # If filename is base pattern
# --- Analysis ---
phase_id = 1 # Phase to analyze
threshold_value = 1.0 # Threshold used by image readers (if applicable)
direction = "X" # Compute for X, Y, Z ("X", "Y", "Z", "All", or "X Z" etc.)
# --- Solver ---
solver_type = "FlexGMRES" # HYPRE solver (GMRES, FlexGMRES, Jacobi, PCG)
hypre_eps = 1e-9 # Solver relative tolerance
hypre_maxiter = 200 # Solver max iterations
# --- Grid ---
box_size = 32 # AMReX max grid size
# --- Output ---
results_dir = "./Diffusion_Results/" # Directory for output text/plot files ('~/' supported)
output_filename = "results.txt" # Name for summary output file
write_plotfile = 0 # Set to 1 to save AMReX plot files from solver
# --- Control ---
verbose = 1 # Output level (0=minimal)
Output
The Diffusion application produces results primarily through console messages and output files:
-
Console:
- Prints progress information during setup and solver execution (level controlled by the
verboseinput parameter). - Reports final calculated Volume Fraction (
VF) for the specifiedphase_id. - Reports final calculated Tortuosity (
Tau) for each computed direction. - May print solver diagnostics like number of iterations and final residual, especially if
verbose > 0. - Reports total wall-clock runtime for the simulation.
- Prints progress information during setup and solver execution (level controlled by the
-
Results File:
- A summary text file is written to the location specified by
results_dirandoutput_filenamein theinputsfile. - This file typically includes a record of the key input parameters used for the run (e.g., filename, phase, direction, solver settings) followed by the final calculated results (Volume Fraction, Tortuosity components).
- Example (
results.txt, content may vary slightly):# Diffusion Calculation Results # Input File: data/SampleData_2Phase.tif # Analysis Phase ID: 1 # Threshold Value: 1.0 # Solver: FlexGMRES # ... other key parameters ... # ----------------------------- # Note: Property values are typically non-dimensionalized or assume unit intrinsic properties. # Check units/scaling based on solver details. VolumeFraction: 0.512345678 Tortuosity_X: 1.987654321 Tortuosity_Y: 1.998765432 Tortuosity_Z: 1.976543210
- A summary text file is written to the location specified by
-
Plotfiles (Optional):
- If
write_plotfile = 1in theinputsfile, AMReX plotfiles may be generated in theresults_dir. - These typically contain the final computed field (e.g., the potential field, named perhaps
potential_X,potential_Y, orpotential_Zdepending on the run) and often include the thresholded phase map (phase_threshold) used for the calculation. - Plotfile names might include direction identifiers or timestamps.
- View these files using standard visualization tools like ParaView, VisIt, yt (see AMReX Visualization Docs).
- If
Visualisation
OpenImpala is built on the AMReX software framework. Output plotfiles (generated when write_plotfile = 1) can be visualised using several open-source visualisation packages, e.g. ParaView, VisIt, yt or AMRVis.
- AMReX Documentation: For further information on native AMReX plotfile formats and viewing options, see the AMReX Visualization Docs.
- Jupyter Notebooks: Alternatively, you can use Jupyter notebooks for analysis and visualisation. A guide demonstrating how to load and plot data from OpenImpala output is available here: https://github.com/jameslehoux/openimpala-jupyter.
As an example, the image below shows a calculated concentration gradient for steady-state diffusive flow (solved in the x-direction) within a 499^3 voxel Lithium Iron Phosphate (LFP) electrode microstructure (Source: [1]):
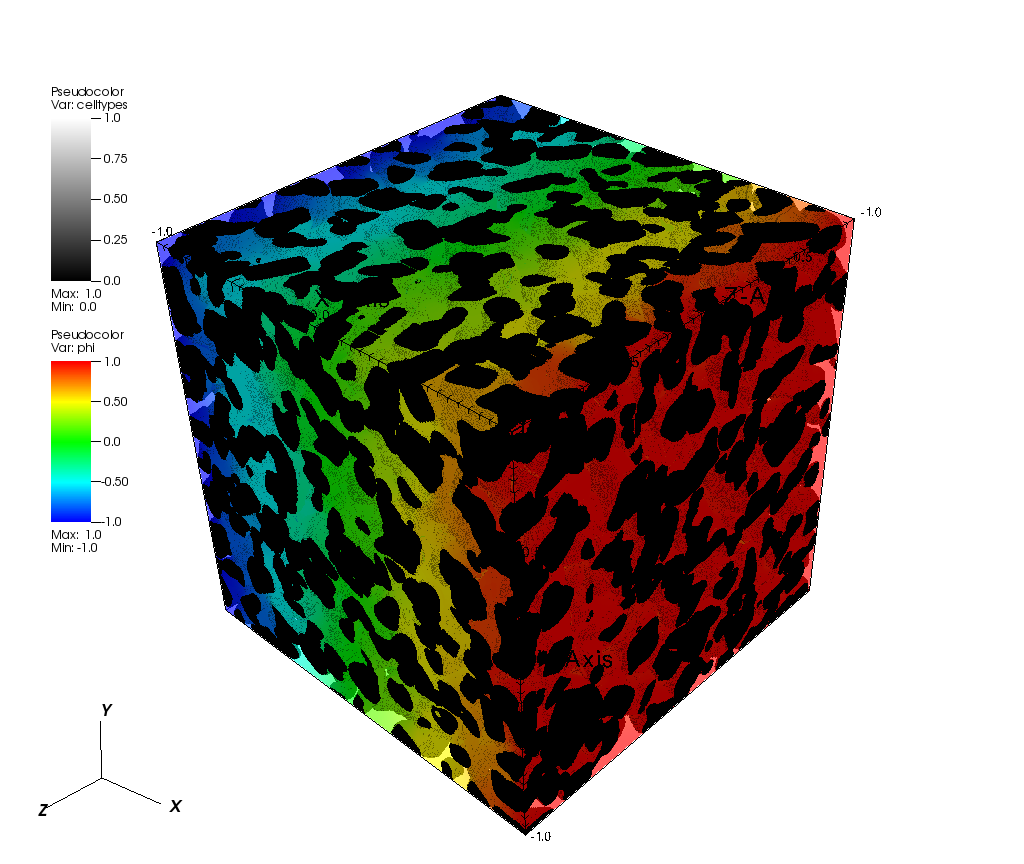
[1]: Le Houx, J., Osenberg, M., Neumann, M., Binder, J.R., Schmidt, V., Manke, I., Carraro, T. and Kramer, D., 2020. Effect of Tomography Resolution on Calculation of Microstructural Properties for Lithium Ion Porous Electrodes. ECS Transactions, 97(7), p.255.
Applications & Related Publications
OpenImpala is actively used in research. If you use OpenImpala in work leading to a publication, please cite the core software paper(s) (see Citation section) and consider letting the developers know or submitting a pull request to add your work here!
Below are some examples of publications using or discussing OpenImpala:
-
Le Houx, J., Melzack, N., James, A., Dehyle, H., Aslani, N., Pimblott, M., Ahmed, S., & Wills, R. G. A. (2024). The Aqueous Aluminium-Ion Battery: Optimising the Electrode Compression Ratio through Image-Based Modelling. ECS Meeting Abstracts, MA2024-01, 2579. https://doi.org/10.1149/MA2024-01462579mtgabs (Application: Aqueous Al-ion battery electrode compression analysis)
-
Le Houx, J., Ruiz, S., McKay Fletcher, D., Ahmed, S., & Roose, T. (2023). Statistical Effective Diffusivity Estimation in Porous Media Using an Integrated On-site Imaging Workflow for Synchrotron Users. Transport in Porous Media, 150, 71–88. https://doi.org/10.1007/s11242-023-01993-7 (Methodology: Effective diffusivity calculation & validation)
-
Fraser, E. J., Le Houx, J. P., Arenas, L. F., Ranga Dinesh, K. K. J., & Wills, R. G. A. (2022). The soluble lead flow battery: Image-based modelling of porous carbon electrodes. Journal of Energy Storage, 52, 104791. https://doi.org/10.1016/j.est.2022.104791 (Application: Soluble lead flow battery RVC electrodes)
-
Le Houx, J., & Kramer, D. (2021). OpenImpala: OPEN source IMage based PArallisable Linear Algebra solver. SoftwareX, 15, 100729. https://doi.org/10.1016/j.softx.2021.100729 (Core Software Paper)
-
Le Houx, J., Osenberg, M., Neumann, M., Binder, J. R., Schmidt, V., Manke, I., Carraro, T., & Kramer, D. (2020). Effect of Tomography Resolution on Calculation of Microstructural Properties for Lithium Ion Porous Electrodes. ECS Transactions, 97(7), 255. https://doi.org/10.1149/09707.0255ecst (Application: LFP electrode resolution effects)
Contributing
Contributions to OpenImpala are welcome! Whether it's reporting bugs, suggesting features, improving documentation, or submitting code, your input is valuable.
- Bug Reports & Feature Requests: Please use the GitHub Issues tracker to report problems or propose new features. Provide as much detail as possible, including steps to reproduce for bugs.
- Code Contributions:
- If you plan to make significant changes, please open an issue first to discuss your ideas.
- For code contributions (bug fixes, enhancements, new features, tests), please follow this general workflow:
- Fork the repository (https://github.com/BASE-Laboratory/OpenImpala).
- Create a new branch for your feature or fix (
git checkout -b feature/my-new-feature). - Make your changes and commit them with clear messages.
- Push your branch to your fork (
git push origin feature/my-new-feature). - Submit a Pull Request to the main repository.
- Documentation: Improvements to the README, code comments, or other documentation are always appreciated. You can submit these via Pull Requests.
Code Formatting
This project uses clang-format to enforce a consistent C++ code style. A .clang-format configuration file is provided in the repository root.
CI enforces formatting — pull requests with unformatted C++ code will fail the format check.
Running Locally
To format all C++ files in-place:
find src/ -type f \( -name "*.cpp" -o -name "*.H" -o -name "*.h" -o -name "*.hpp" \) \
-exec clang-format -i {} +
To check for formatting issues without modifying files:
find src/ -type f \( -name "*.cpp" -o -name "*.H" -o -name "*.h" -o -name "*.hpp" \) \
-exec clang-format --dry-run --Werror {} +
Note: Fortran files (
.F90,.f90) are not covered by clang-format and should not be passed to it.
Static Analysis
This project uses clang-tidy for static analysis. A .clang-tidy configuration file in the repository root enables conservative checks from the bugprone-*, performance-*, and modernize-use-nullptr categories.
CI enforces static analysis — pull requests that introduce new clang-tidy warnings will fail.
Running Locally
If you are building inside the dependency container, run the Makefile target:
apptainer exec --bind "$(pwd):/src" dependency_image.sif bash -c "cd /src && make tidy"
Or run clang-tidy on individual files with the project's include paths:
clang-tidy src/props/MyFile.cpp -- -std=c++17 -DOMPI_SKIP_MPICXX \
-Isrc -Isrc/props \
-I${AMREX_HOME}/include -I${HYPRE_HOME}/include \
-I${HDF5_HOME}/include -I${TIFF_HOME}/include
Citation
If you use OpenImpala in your research or publications, we kindly ask that you cite the relevant paper(s).
-
General Use & Software Framework: Please cite this paper when using OpenImpala for simulations or analysis based on its core functionality.
@article{LeHoux2021OpenImpala, title = {{{OpenImpala}}: {{OPEN}} source {{IMage}} based {{PArallisable}} {{Linear}} {{Algebra}} solver}, author = {Le Houx, James and Kramer, Denis}, year = {2021}, journal = {SoftwareX}, volume = {15}, pages = {100729}, doi = {10.1016/j.softx.2021.100729}, issn = {2352-7110} }
Le Houx, J., & Kramer, D. (2021). OpenImpala: OPEN source IMage based PArallisable Linear Algebra solver. SoftwareX, 15, 100729.
-
Specific Methods (e.g., Effective Diffusivity via Homogenization): Please consider citing this paper in addition to the primary software paper if you are using or comparing against the specific homogenization methods or on-site workflow described therein.
@article{le2023statistical, title={Statistical Effective Diffusivity Estimation in Porous Media Using an Integrated On-site Imaging Workflow for Synchrotron Users}, author={Le Houx, James and Ruiz, Siul and McKay Fletcher, Daniel and Ahmed, Sharif and Roose, Tiina}, journal={Transport in Porous Media}, volume={150}, number={1}, pages={71--88}, year={2023}, publisher={Springer}, doi={10.1007/s11242-023-01993-7} }
Le Houx, J., Ruiz, S., McKay Fletcher, D., Ahmed, S., & Roose, T. (2023). Statistical Effective Diffusivity Estimation in Porous Media Using an Integrated On-site Imaging Workflow for Synchrotron Users. Transport in Porous Media, 150, 71–88.
License
OpenImpala Copyright (c) 2020-2025, University of Southampton and contributors. All rights reserved.
The software is licensed under the BSD 3-Clause "New" or "Revised" License. The full license text can be found in the LICENSE file.
Acknowledgements
This work was financially supported by the EPSRC Centre for Doctoral Training (CDT) in Energy Storage and its Applications [grant ref: EP/R021295/1], the Ada Lovelace Centre (ALC) STFC project, CANVAS-NXtomo, ContAiNerised Voxel-bAsed Simulation of Neutron and X-ray Tomography data, the EPSRC prosperity partnership with Imperial College, INFUSE, Interface with the Future - Underpinning Science to Support the Energy transition EP/V038044/1, the Rutherford Appleton Laboratory and The Faraday Institution through James Le Houx's Emerging Leader Fellowship (Grant No. FIELF001), and from Research England’s ‘Expanding Excellence in England’ grant at the University of Greenwich via the “Multi-scale Multi-disciplinary Modelling for Impact” (M34Impact) programme.
The authors acknowledge the use of the IRIDIS High Performance Computing Facility, Diamond Light Source's Wilson HPC cluster, STFC's SCARF cluster and the University of Greenwich's M34Impact HPC Cluster in the completion of this work.
We thank the developers of AMReX, HYPRE, libtiff, and HDF5 upon which OpenImpala relies.
Contact & Support
For questions, bug reports, or feature requests, please use the GitHub Issues tracker for this repository.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distributions
Built Distributions
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file openimpala-4.2.9-cp312-cp312-manylinux_2_28_x86_64.whl.
File metadata
- Download URL: openimpala-4.2.9-cp312-cp312-manylinux_2_28_x86_64.whl
- Upload date:
- Size: 6.2 MB
- Tags: CPython 3.12, manylinux: glibc 2.28+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
79281c3144d5cfb59995902779f67b654b7631b02f63c1582a0000417156e185
|
|
| MD5 |
e5d93f5f5a7f61fcf868b652ba82dd8d
|
|
| BLAKE2b-256 |
7c852e593db3bc62a51b638052dacb6667ce0a2b61e73eb35e984068e526b374
|
Provenance
The following attestation bundles were made for openimpala-4.2.9-cp312-cp312-manylinux_2_28_x86_64.whl:
Publisher:
pypi-wheels-cpu.yml on BASE-Laboratory/OpenImpala
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
openimpala-4.2.9-cp312-cp312-manylinux_2_28_x86_64.whl -
Subject digest:
79281c3144d5cfb59995902779f67b654b7631b02f63c1582a0000417156e185 - Sigstore transparency entry: 1409190579
- Sigstore integration time:
-
Permalink:
BASE-Laboratory/OpenImpala@bcf82b24dab5484f757c1085d389952b73a9f150 -
Branch / Tag:
refs/tags/v4.2.9 - Owner: https://github.com/BASE-Laboratory
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
pypi-wheels-cpu.yml@bcf82b24dab5484f757c1085d389952b73a9f150 -
Trigger Event:
release
-
Statement type:
File details
Details for the file openimpala-4.2.9-cp311-cp311-manylinux_2_28_x86_64.whl.
File metadata
- Download URL: openimpala-4.2.9-cp311-cp311-manylinux_2_28_x86_64.whl
- Upload date:
- Size: 6.2 MB
- Tags: CPython 3.11, manylinux: glibc 2.28+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
13974bad6a9f608db7d4f5e1646d5ab514e0ee0a78d8e782476795c88318e1e4
|
|
| MD5 |
3e45627912f32ae99f6fac20a6e950ff
|
|
| BLAKE2b-256 |
1c6e35d7c4b7af7a5e3465f233180eb7d3a5855122c9d386393cc2fae1ae058a
|
Provenance
The following attestation bundles were made for openimpala-4.2.9-cp311-cp311-manylinux_2_28_x86_64.whl:
Publisher:
pypi-wheels-cpu.yml on BASE-Laboratory/OpenImpala
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
openimpala-4.2.9-cp311-cp311-manylinux_2_28_x86_64.whl -
Subject digest:
13974bad6a9f608db7d4f5e1646d5ab514e0ee0a78d8e782476795c88318e1e4 - Sigstore transparency entry: 1409190615
- Sigstore integration time:
-
Permalink:
BASE-Laboratory/OpenImpala@bcf82b24dab5484f757c1085d389952b73a9f150 -
Branch / Tag:
refs/tags/v4.2.9 - Owner: https://github.com/BASE-Laboratory
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
pypi-wheels-cpu.yml@bcf82b24dab5484f757c1085d389952b73a9f150 -
Trigger Event:
release
-
Statement type:
File details
Details for the file openimpala-4.2.9-cp310-cp310-manylinux_2_28_x86_64.whl.
File metadata
- Download URL: openimpala-4.2.9-cp310-cp310-manylinux_2_28_x86_64.whl
- Upload date:
- Size: 6.2 MB
- Tags: CPython 3.10, manylinux: glibc 2.28+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5913690d1aff8cd2d736f5041419bc60fbd81e38b2300cb38c65bdbffad2899e
|
|
| MD5 |
0be1464642b7cac08be9db88488bcb07
|
|
| BLAKE2b-256 |
a4eeae4a8631bf9290e126ae9126a4c080f0ff5a0e174c10684e89bf3463b1b9
|
Provenance
The following attestation bundles were made for openimpala-4.2.9-cp310-cp310-manylinux_2_28_x86_64.whl:
Publisher:
pypi-wheels-cpu.yml on BASE-Laboratory/OpenImpala
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
openimpala-4.2.9-cp310-cp310-manylinux_2_28_x86_64.whl -
Subject digest:
5913690d1aff8cd2d736f5041419bc60fbd81e38b2300cb38c65bdbffad2899e - Sigstore transparency entry: 1409190604
- Sigstore integration time:
-
Permalink:
BASE-Laboratory/OpenImpala@bcf82b24dab5484f757c1085d389952b73a9f150 -
Branch / Tag:
refs/tags/v4.2.9 - Owner: https://github.com/BASE-Laboratory
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
pypi-wheels-cpu.yml@bcf82b24dab5484f757c1085d389952b73a9f150 -
Trigger Event:
release
-
Statement type:
File details
Details for the file openimpala-4.2.9-cp39-cp39-manylinux_2_28_x86_64.whl.
File metadata
- Download URL: openimpala-4.2.9-cp39-cp39-manylinux_2_28_x86_64.whl
- Upload date:
- Size: 6.2 MB
- Tags: CPython 3.9, manylinux: glibc 2.28+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
45680e854752a54eb9312d0c42d18e12cf1c2927a3537b2b723c9e6ecdf2af01
|
|
| MD5 |
5527b4d5b4013fde7aa0598c6cf56c52
|
|
| BLAKE2b-256 |
8e3481f2cb1ce4e61f901d56780216b9de65762b02ed65fe920dd18fe402a2ba
|
Provenance
The following attestation bundles were made for openimpala-4.2.9-cp39-cp39-manylinux_2_28_x86_64.whl:
Publisher:
pypi-wheels-cpu.yml on BASE-Laboratory/OpenImpala
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
openimpala-4.2.9-cp39-cp39-manylinux_2_28_x86_64.whl -
Subject digest:
45680e854752a54eb9312d0c42d18e12cf1c2927a3537b2b723c9e6ecdf2af01 - Sigstore transparency entry: 1409190594
- Sigstore integration time:
-
Permalink:
BASE-Laboratory/OpenImpala@bcf82b24dab5484f757c1085d389952b73a9f150 -
Branch / Tag:
refs/tags/v4.2.9 - Owner: https://github.com/BASE-Laboratory
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
pypi-wheels-cpu.yml@bcf82b24dab5484f757c1085d389952b73a9f150 -
Trigger Event:
release
-
Statement type:




















