GPU/CUDA optimized implementation of 3D optical flow algorithms such as Farneback two frame motion estimation and Lucas Kanade dense optical flow algorithms
Project description
opticalflow3d
GPU/CUDA optimized implementation of 3D optical flow algorithms such as Farneback two frame motion estimation and Lucas Kanade dense optical flow algorithms.
Please see the related projects section for the other components of this pipeline
Overview
Being able to efficiently calculate the displacements between two imaging volumes will allow us to query the dynamics captured in the images. This would include 3D biological samples and 3D force micrsocopy. However, a CPU based implementation would be too time consuming. Thus, this repository was created to address this problem by providing GPU accelerated implementation of various optical flow algorithms. Speed is key here, and tricks such as separable convolutions are used.
Currently, two optical flow methods are implemented:
- Pyramidal Lucas Kanade dense optical flow algorithm
- Farneback two frame motion estimation algorithm
The following methods are also provided to help in the assessment of the vectors.
- forward mapping of the first image using the calculated vectors
Validation of the 3D Farneback algorithm
Accuracy of the algorithm on synthetic 3D fluorescent bead images
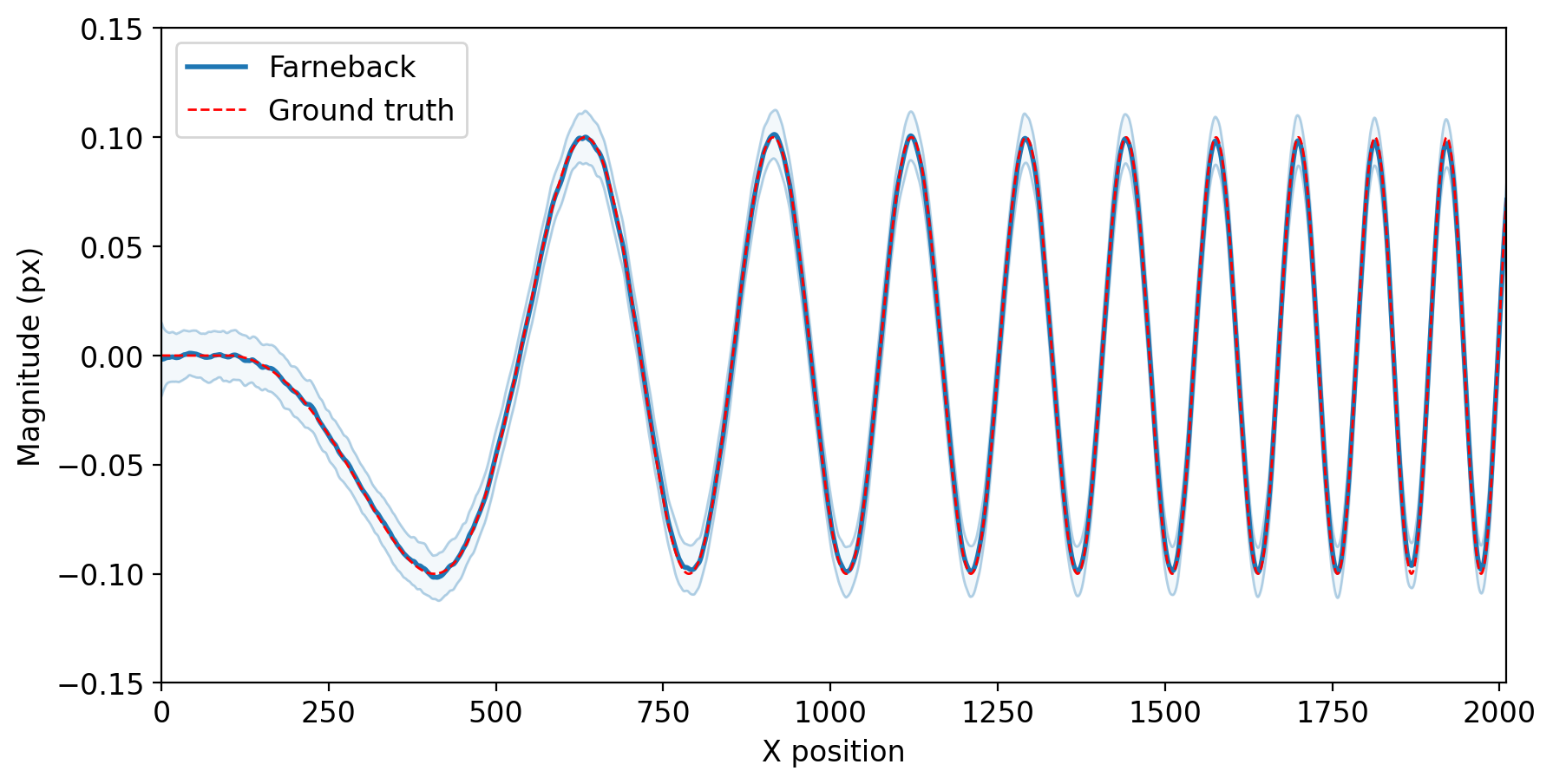
The Farneback two frame motion estimation algorithm was validated against the L5 SEM Challenge Sample 14 dataset [1]. The RMSE for the x component of the displacement field was 0.0112. The figure below shows the accuracy of the x component. The solid blue line represents the mean magnitude of the x component for each x position while the shaded region/thin blue shows the standard deviation of the x component magnitudes.
Execution time
The time taken to compute the displacements for a 2048×192×192 voxel image was 2.41s ± 21.4ms. The computation was done on a server running a Quadro RTX 6000 GPU and dual Intel(R) Xeon(R) Gold 5217 CPUs.
For comparison, the FFT-based DVC algorithm took 1980.04s while ALDVC took 9335.2s [1]. A few caveats have to be mentioned as both of these algorithms were running on the CPU only and ALDVC does calculate other parameters as well.
Usage
Required packages
The following packages are required. Please ensure that they are installed using either pip or conda.
- numpy
- numba
- scikit-image
- scipy
- cupy
- tqdm
Installation
This package is available via pip and can be installed using:
pip install opticalflow3d
Examples
Examples can be found in the examples folder
How to cite
Huang, C. K., Yong, X., She, D. T., & Lim, C. T. (2022). Surface curvature and basal hydraulic stress induce spatial bias in cell extrusion. bioRxiv.
Related projects
References
[1] Yang, J., Hazlett, L., Landauer, A. K., & Franck, C. (2020). Augmented Lagrangian Digital Volume Correlation (ALDVC). Experimental Mechanics, 60(9), 1205-1223.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file opticalflow3d-0.3.2.tar.gz.
File metadata
- Download URL: opticalflow3d-0.3.2.tar.gz
- Upload date:
- Size: 18.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.4.1 importlib_metadata/4.0.1 pkginfo/1.7.0 requests/2.25.1 requests-toolbelt/0.9.1 tqdm/4.57.0 CPython/3.8.13
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9d969dd9d11275be1106f4e9c42e429410c347d10f82a0f06f96efaa4eebea05
|
|
| MD5 |
1f61ebeaaef23f8a897b982f4277fa38
|
|
| BLAKE2b-256 |
35190abca65d2f33a51b819bb62cde93ff63f86aab9b57357a3a112aabb322c4
|
File details
Details for the file opticalflow3d-0.3.2-py3-none-any.whl.
File metadata
- Download URL: opticalflow3d-0.3.2-py3-none-any.whl
- Upload date:
- Size: 18.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.4.1 importlib_metadata/4.0.1 pkginfo/1.7.0 requests/2.25.1 requests-toolbelt/0.9.1 tqdm/4.57.0 CPython/3.8.13
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7ea6a099352e2699afd97bc99ae9ce0ffbe3225ad12d781549ccdfda3a577855
|
|
| MD5 |
6ee3468ad424753cec04db75a5292805
|
|
| BLAKE2b-256 |
7ade19d50bd5d614cb0e90c70c4ca7f808af2bd914b325fd7a689d67aad93cff
|











