Python ⇄ Julia bridge for the OptimalGIV package
Project description
optimalgiv
A minimal Python wrapper for OptimalGIV.jl
This interface enables Python users to call Granular Instrumental Variables (GIV) estimators directly on pandas DataFrames using JuliaCall. Julia is automatically installed and all dependencies are resolved without manual setup.
Note: This README provides full usage details for Python users.
For more technical and Julia-specific documentation, please see here
Model Specification
The Granular Instrumental Variables (GIV) model estimated by this package follows the specification:
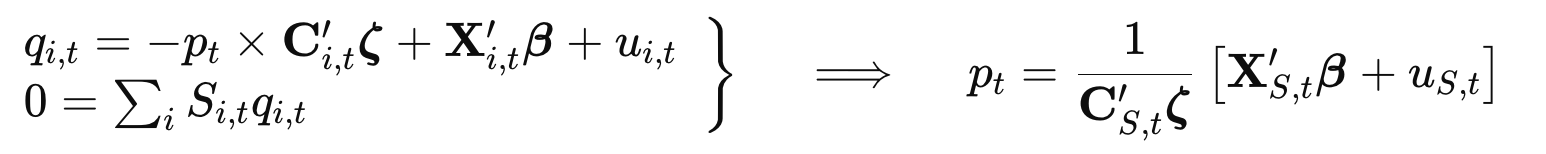
where:
- $q_{i,t}$ and $p_t$ are endogenous,
- $\mathbf{C}_{i,t}$ is a vector of controls for slopes,
- $\mathbf{X}_{i,t}$ is a vector of controls,
- $\boldsymbol{\zeta}$, $\boldsymbol{\beta}$ are coefficient vectors,
- $u_{i,t}$ is the idiosyncratic shock, and
- $S_{i,t}$ is the weighting variable.
The equilibrium price $p_t$ is derived by imposing the market clearing condition and the model is estimated using the moment condition:
$$ \mathbb{E}[u_{i,t} u_{j,t}] = 0 $$
for all $i \neq j$. This implies orthogonality across sectors' residuals.
Panel Data and Coverage
The GIV model supports unbalanced panel data. However, some estimation algorithms (e.g. "scalar_search" and "debiased_ols") require complete coverage, meaning:
$$ \sum_i S_{i,t} q_{i,t} = 0 $$
must hold exactly within the sample. This ensures internal consistency of the equilibrium condition.
If the adding-up constraint is not satisfied, the model will adjust accordingly, but the interpretation of estimated coefficients should be made with caution, as residual market imbalances may bias elasticities and standard errors. (See the complete_coverage argument below for details.)
Installation
pip install optimalgiv
On first importing, optimalgiv will automatically:
- Install Julia (if not already available)
- Install
OptimalGIV.jland supporting packages - Precompile and create a self-contained Julia environment
Usage
Basic Example
import pandas as pd
import numpy as np
from optimalgiv import giv
# Use a properly simulated dataset with 5 sectors ('id' column) and multiple periods ('t')
df = pd.read_csv("./simdata1.csv")
df['id'] = df['id'].astype('category') # ensure id interactions map to distinct groups
# Define the model formula
formula = "q + id & endog(p) ~ 0 + fe(id) + fe(id) & (η1 + η2)"
# Provide an initial guess (a good guess is critical)
guess = np.ones(5)
# Estimate the model
model = giv(
df = df,
formula = "q + id & endog(p) ~ 0 + fe(id) + fe(id) & (η1 + η2)",
id = "id",
t = "t",
weight = "absS",
algorithm = "iv",
guess = guess,
save = 'all', # saves both fixed‐effects (model.fe) and residuals (model.residual_df)
)
# View the result
model.summary()
## GIVModel (Aggregate coef: 2.13)
## ─────────────────────────────────────────────────────────────────────────
## Estimate Std. Error t-stat Pr(>|t|) Lower 95% Upper 95%
## ─────────────────────────────────────────────────────────────────────────
## id: 1 & p 1.00723 1.30407 0.772377 0.4405 -1.55923 3.57369
## id: 2 & p 1.77335 0.475171 3.73204 0.0002 0.8382 2.70851
## id: 3 & p 1.36863 0.382177 3.58114 0.0004 0.616491 2.12077
## id: 4 & p 3.3846 0.382352 8.85207 <1e-16 2.63212 4.13709
## id: 5 & p 0.619882 0.161687 3.83385 0.0002 0.301676 0.938087
Formula Specification
The model formula follows the convention:
q + interactions & endog(p) ~ exog_controls
Where:
-
q: Response variable (e.g., quantity). -
endog(p): Endogenous variable (e.g., price). Must appear on the left-hand side.Note: A positive estimated coefficient implies a negative response of
qtop(i.e., a downward-sloping demand curve). -
interactions: Exogenous variables used to parameterize heterogeneous elasticities, such as entity identifiers or group characteristics. -
exog_controls: Exogenous control variables. Supports fixed effects (e.g.,fe(id)) using the same syntax asFixedEffectModels.jl.
Examples of formulas:
# 1. Homogeneous elasticity with no intercept and two controls
formula = "q + endog(p) ~ 0 + n1 + n2"
# 2. Homogeneous elasticity, with fixed effects absorbed by id
formula ="q + endog(p) ~ n1 + n2 + fe(id)"
# 3. Heterogeneous elasticity by id, no controls
formula ="q + id & endog(p) ~ 1"
# 4. Heterogeneous elasticity by id, with one control
formula ="q + id & endog(p) ~ n1"
# 5. Fully saturated: elasticity by id, controls and intercepts vary by id (absorbed by fixed effect), no global intercept
formula ="q + id & endog(p) ~ 0 + fe(id) + fe(id) & (n1 + n2)"
Key Function: giv()
giv(df, formula: str, id: str, t: str, weight: str, **kwargs) -> GIVModel
Required Arguments
df:pandas.DataFramecontaining panel data. Must be balanced for some algorithms (e.g.,scalar_search).formula: A string representing the model (Julia-style formula syntax). See examples above.id: Name of the column identifying entities (e.g.,"firm_id").t: Name of the time variable column.weight: Name of the weight/size column (e.g., market sharesS_i,t).
Keyword Arguments (Optional)
-
algorithm: One of"iv"(default),"iv_twopass","debiased_ols", or"scalar_search". -
guess: Initial guess for ζ coefficients. (See below for usage details) -
exclude_pairs: Dictionary excluding pairs from moment conditions. Example:{1: [2, 3], 4: [5]}excludes (1,2), (1,3), and (4,5). -
quiet: SetTrueto suppress warnings and info messages. -
save:"none"(default),"residuals","fe", or"all"— controls what is stored on the returned model:"none": neither residuals nor fixed-effects are saved"residuals": saves residuals inmodel.residual_df"fe": saves fixed-effects inmodel.fe"all": saves bothmodel.residual_dfandmodel.fe
-
save_df: IfTrue, the full estimation dataframe (with residuals, coefficients, fixed effects) is stored inmodel.df. -
complete_coverage: Whether the dataset covers the full market in each time period, meaning $\sum_i S_{i,t} q_{i,t} = 0$ holds exactly within the sample.- Default is
None, which triggers auto-detection: the model checks this condition period-by-period and sets the flag toTrueorFalseaccordingly. - If the condition does not hold (
False), you can still force estimation by settingquiet=True, but results may be biased. Use with caution. - Required for
"scalar_search"and"debiased_ols"algorithms.
- Default is
-
return_vcov: Whether to compute and return the variance–covariance matrices. (default:True) -
tol: Convergence tolerance for the solver (:1e-6) -
iterations: Maximum number of solver iterations (:100)
Advanced keyword arguments (Optional; Use with caution)
-
contrasts(Dict[str, Union[str, Any]]) Specifies encoding schemes for categorical variables, following Julia'sStatsModels.jl.⚠️ Untested at all! — use at your own risk.
- Keys: column names (as strings).
- Values: either
- a string like
"HelmertCoding","TreatmentCoding"(converted automatically toStatsModels.<X>()), or - an actual Julia object like
jl.StatsModels.HelmertCoding()The bridge converts this to a JuliaDict(:id => HelmertCoding(), ...)for use in formula parsing.
- a string like
-
solver_options(Dict[str, Any]) Extra options passed to the nonlinear system solver fromNLsolve.jl. The Python dict is converted to a JuliaNamedTuplewith keyword-style arguments. Common options include:-
"method":"newton","anderson","trust_region", etc. -
"ftol": absolute residual tolerance -
"xtol": absolute solution tolerance -
"iterations": max iterations -
"show_trace": verbose output -
"linesearch": can be- a Julia object like
jl.LineSearches.HagerZhang(), or - a string like
"HagerZhang", which is expanded toLineSearches.HagerZhang()automatically
- a Julia object like
Example:
solver_opts = { "method": "newton", "ftol": 1e-8, "xtol": 1e-8, "iterations": 1000, "show_trace": True, "linesearch": "HagerZhang", # ← string is auto-converted } model = giv(df, formula, id="id", t="t", solver_options=solver_opts)
For the full list of options, see the NLsolve.jl documentation.
-
Algorithms
The package implements four algorithms for GIV estimation:
-
"iv"(Instrumental Variables)- Default, recommended
- Uses moment condition $$(\mathbb{E}[u_i,u_{S,-i}]=0)$$
- $$O(N)$$ implementation
- Supports
exclude_pairs(exclude certain pairs $E[u_i u_j] = 0$ from the moment conditions) - Supports flexible elasticity specs, unbalanced panels
-
"iv_twopass": Numerically identical toivbut uses a more straightforward O(N²) implementation with two passes over entity pairs. This is useful for:- Debugging purposes
- When the O(N) optimization in
ivmight cause numerical issues - When there are many pairs to be excluded, which will slow down the algorithm in
iv - Understanding the computational flow of the moment conditions
-
"debiased_ols"- Uses $$\mathbb{E}[u_iC_{it}p_{it}] = \sigma_i^2 / \zeta_{St}$$
- Requires complete market coverage
- More efficient but restrictive
-
"scalar_search"- Finds a single aggregate elasticity
- Requires balanced panel, constant weights, complete coverage
- Useful for diagnostics or initial-guess formation
Initial Guesses
A good guess is key to stable estimation. If omitted, OLS‐based defaults will typically fail. Examples:
import numpy as np
from optimalgiv import giv
# 1) Scalar guess (for homogeneous elasticity)
guess = 1.0
model1 = giv(
df,
"q + endog(p) ~ n1 + fe(id)",
id="id", t="t", weight="S",
guess=guess
)
# 2) Dict by group name (heterogeneous by id)
guess = {"id": [1.2, 0.8]}
model2 = giv(
df,
"q + id & endog(p) ~ 1",
id="id", t="t", weight="S",
guess=guess
)
# 3) Dict for multiple interactions
guess = {
"id": [1.0, 0.9],
"n1": [0.5, 0.3]
}
model3 = giv(
df,
"q + id & endog(p) + n1 & endog(p) ~ fe(id)",
id="id", t="t", weight="S",
guess=guess
)
# 4) Dict keyed by exact coefnames
names = model3.coefnames()
guess = {name: 0.1 for name in names}
model4 = giv(
df,
"q + id & endog(p) + n1 & endog(p) ~ fe(id)",
id="id", t="t", weight="S",
guess=guess
)
# 5) Scalar-search with heterogeneous formula
guess = {"Aggregate": 2.5}
model5 = giv(
df,
"q + id & endog(p) ~ 0 + fe(id) + fe(id)&(n1 + n2)",
id="id", t="t", weight="S",
algorithm="scalar_search",
guess=guess
)
# 6) Use estimated ζ from model5 as initial guess
guess = model5.endog_coef
model6 = giv(
df,
"q + id & endog(p) ~ 0 + fe(id) + fe(id)&(n1 + n2)",
id="id", t="t", weight="S",
guess=guess
)
Working with Results
# Methods
model.summary() # ▶ print full Julia-style summary
model.residuals() # ▶ numpy array of the residuals for each observation
model.confint(level=0.95) # ▶ (n×2) array of confidence intervals
model.coeftable(level=0.95)# ▶ pandas.DataFrame of estimates, SEs, t-stats, p-values
# Fields
model.endog_coef # ▶ numpy array of ζ coefficients
model.exog_coef # ▶ numpy array of β coefficients
model.agg_coef # ▶ float: aggregate elasticity
model.endog_vcov # ▶ VCOV of ζ coefficients
model.exog_vcov # ▶ VCOV of β coefficients
model.nobs # ▶ int: number of observations
model.dof_residual # ▶ int: residual degrees of freedom
model.formula # ▶ str: Julia-style formula
model.formula_schema # ▶ str: the internal schema of the Julia‐style formula after parsing
model.residual_variance # ▶ numpy array of the estimated variance of the residuals for each entity (ûᵢ’s variance)
model.N # ▶ int: the number of cross‐section entities in the panel
model.T # ▶ int: the number of time periods per entity in the panel
model.dof # ▶ int: the total number of estimated parameters (length of ζ plus length of β)
model.responsename # ▶ str: the name of the response variable(s)
model.converged # ▶ bool: solver convergence status
model.endog_coefnames # ▶ list[str]: ζ coefficient names
model.exog_coefnames # ▶ list[str]: β coefficient names
model.idvar # ▶ str: entity identifier column name
model.tvar # ▶ str: time identifier column name
model.weightvar # ▶ str or None: weight column name
model.exclude_pairs # ▶ dict: excluded moment-condition pairs
model.coefdf # ▶ pandas.DataFrame of entity-specific coefficients
model.fe # ▶ pandas.DataFrame of fixed-effects (if saved)
model.residual_df # ▶ pandas.DataFrame of residuals (if saved)
model.df # ▶ pandas.DataFrame of full estimation output (if save_df=True)
model.coef # ▶ numpy array of [ζ; β]
model.vcov # ▶ full (ζ+β) variance–covariance matrix
model.stderror # ▶ numpy array of standard errors
model.coefnames # ▶ list[str]: names of all coefficients (ζ then β)
References
Please cite:
- Gabaix, Xavier, and Ralph S.J. Koijen. Granular Instrumental Variables. Journal of Political Economy, 132(7), 2024, pp. 2274–2303.
- Chaudhary, Manav, Zhiyu Fu, and Haonan Zhou. Anatomy of the Treasury Market: Who Moves Yields? Available at SSRN: https://ssrn.com/abstract=5021055
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distributions
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file optimalgiv-0.1.10-py3-none-any.whl.
File metadata
- Download URL: optimalgiv-0.1.10-py3-none-any.whl
- Upload date:
- Size: 16.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.9.6
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c5536791adb9ae7f6ae296d9aca3d7d7024290831934a2f1040e0311af25c326
|
|
| MD5 |
71f3f56cd87c80ba169f42fca8888676
|
|
| BLAKE2b-256 |
c9165ae0a22d5cbbb5352251c52ef0ec96d1ba999fcc785fa27d1a603fedddeb
|











